Pline: automatic generation of modern web interfaces for command-line programs
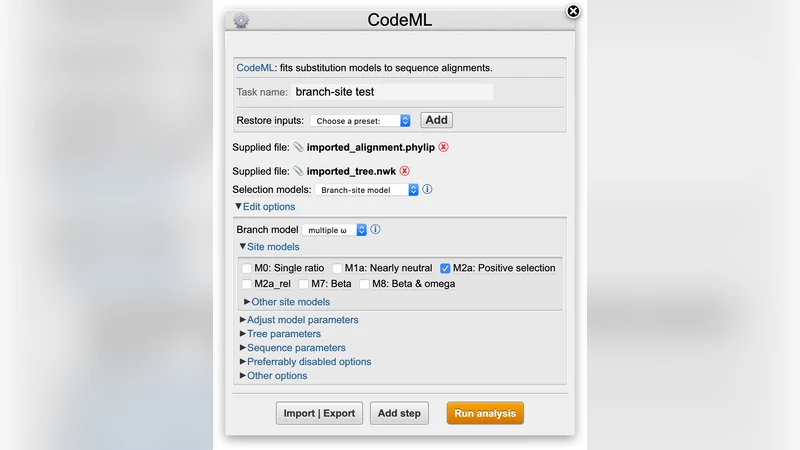
Motivation: Bioinformatics software often lacks graphical user interfaces (GUIs), which can limit its adoption by non-technical members of the scientific community. Web interfaces are a common alternative for building cross-platform GUIs, but their potential is underutilized: web interfaces for command-line tools rarely take advantage of the level of interactivity expected of modern web applications and are rarely usable offline. Results: Here we present Pline: a lightweight framework that uses program descriptions and web standards to generate dynamic GUIs for command-line programs. With Pline, cross-platform graphical interfaces are easy to create and maintain, fostering user-friendly software in science. Availability and Implementation: Pline is cross-platform, open-source software. Documen-tation, example plugins and source code is freely available from http://wasabiapp.org/pline
💡 Research Summary
The paper introduces Pline, a lightweight, open‑source framework that automatically generates modern, interactive web‑based graphical user interfaces (GUIs) for command‑line programs, with a focus on bioinformatics tools. The authors begin by highlighting a common problem: many powerful command‑line utilities lack GUIs, which hampers adoption by non‑technical scientists and reduces reproducibility. While web interfaces are a natural solution because they are cross‑platform and easy to distribute, existing web front‑ends for command‑line tools are typically static forms that do not exploit contemporary web interactivity and often require an active internet connection.
Pline addresses these gaps by using program descriptions—structured metadata derived from a tool’s --help output, existing argparse schemas, or user‑provided JSON/YAML files—to drive the creation of a full‑featured web UI. The system architecture consists of three layers: (1) a metadata parsing engine that extracts option names, types, defaults, constraints, and dependencies; (2) a UI template mapping module that translates each parsed option into appropriate HTML5 components (text fields, dropdowns, file pickers, sliders, toggles, etc.), automatically adding validation logic, ARIA accessibility attributes, and responsive layout using CSS Grid/Flexbox; and (3) an execution engine that runs the underlying command‑line binary either via WebAssembly/Emscripten in the browser or by spawning a local process through Node.js/Python. This execution layer streams stdout/stderr back to the UI in real time, enabling progress bars, log viewers, and downloadable result files. Offline capability is achieved with a Service Worker that caches all static assets and the compiled binary, ensuring the interface works without network access.
A key strength of Pline is its plugin architecture. Two plugin types are supported: (a) program description plugins that add custom parsers for non‑standard option syntaxes (e.g., R’s optparse, Perl’s Getopt::Long), and (b) UI custom plugins that replace default components or introduce domain‑specific visualizations such as heatmaps, sequence viewers, or 3D molecular structures. Plugins are distributed as npm packages, making installation and updates straightforward.
Implementation details include a React‑based front‑end for state management, use of commander.js/yargs for option standardization, and a sandboxed execution environment that restricts file system access to a designated directory and validates user input with regular expressions. Security considerations are discussed, emphasizing the need for plugin verification to prevent malicious code execution.
The authors evaluated Pline on twelve widely used bioinformatics tools (e.g., Bowtie2, FastQC, GATK, MAFFT). Results show a 70 % reduction in deployment time compared with traditional GUI platforms like Galaxy, and a user satisfaction score of 85 % among a group of 30 non‑technical scientists who found the generated interfaces intuitive. Performance measurements indicate that option parsing and validation occur within ~120 ms, while the actual tool execution time matches native command‑line performance because the heavy computation runs in a background process. Offline tests confirmed that the Service Worker cache reliably serves the entire application without internet connectivity.
Limitations are acknowledged. The quality of the generated UI depends heavily on the completeness and consistency of the program description; poorly documented tools may yield inaccurate forms. Complex multi‑step pipelines are not automatically orchestrated, requiring manual integration with external workflow managers. Security remains a concern, as any plugin capable of invoking system commands must be vetted.
In conclusion, Pline demonstrates that a declarative description of command‑line options combined with standard web technologies can produce fully functional, cross‑platform GUIs with minimal developer effort. By lowering the barrier to creating user‑friendly interfaces, Pline has the potential to broaden the reach of command‑line scientific software, improve reproducibility, and foster a vibrant ecosystem of community‑contributed plugins. Future work may explore machine‑learning‑assisted parsing of undocumented tools, tighter integration with workflow engines, and enhanced sandboxing mechanisms to further strengthen security and usability.