Computational MRI with Physics-based Constraints: Application to Multi-contrast and Quantitative Imaging
Compressed sensing takes advantage of low-dimensional signal structure to reduce sampling requirements far below the Nyquist rate. In magnetic resonance imaging (MRI), this often takes the form of sparsity through wavelet transform, finite differences, and low rank extensions. Though powerful, these image priors are phenomenological in nature and do not account for the mechanism behind the image formation. On the other hand, MRI signal dynamics are governed by physical laws, which can be explicitly modeled and used as priors for reconstruction. {1}These explicit and implicit signal priors can be synergistically combined in an inverse problem framework to recover sharp, multi-contrast images from highly accelerated scans. Furthermore, the physics-based constraints provide a recipe for recovering quantitative, bio-physical parameters from the data. This article introduces physics-based modeling constraints in MRI and shows how they can be used in conjunction with compressed sensing for image reconstruction and quantitative imaging. We describe model-based quantitative MRI, as well as its linear subspace approximation. We also discuss approaches to selecting user-controllable scan parameters given knowledge of the physical model. We present several MRI applications that take advantage of this framework for the purpose of multi-contrast imaging and quantitative mapping.
💡 Research Summary
This paper presents a unified framework that integrates physics‑based signal modeling with compressed sensing (CS) to enable highly accelerated, multi‑contrast magnetic resonance imaging (MRI) and direct quantitative mapping of tissue parameters. Traditional CS‑MRI relies on phenomenological image priors such as wavelet sparsity, total variation, or low‑rank structure to compensate for undersampling, but it ignores the underlying physical mechanisms that generate the MR signal. The authors argue that incorporating the Bloch equations—or their efficient surrogate, the extended phase graph (EPG) formalism—provides explicit constraints on the signal evolution, allowing the reconstruction algorithm to exploit both statistical and physical priors simultaneously.
Key components of the proposed approach are:
-
Physical Signal Model – The transverse magnetization at each voxel is expressed as ρ(r)·f(t;θ(r),u(t)), where ρ is proton density, θ denotes intrinsic tissue parameters (primarily T1 and T2), and u represents the sequence‑level control variables (RF flip angles, gradient timing, etc.). The function f is the solution of the Bloch differential equations, which can be simulated directly or approximated with EPG.
-
Linear Subspace Approximation – Because f is nonlinear, the authors pre‑compute a large set of signal evolutions for a grid of θ and u values, then perform singular‑value decomposition (SVD) to extract a low‑dimensional basis that captures the dominant dynamics. This subspace is incorporated into the CS reconstruction as a dictionary, dramatically reducing the number of unknowns while preserving the physics.
-
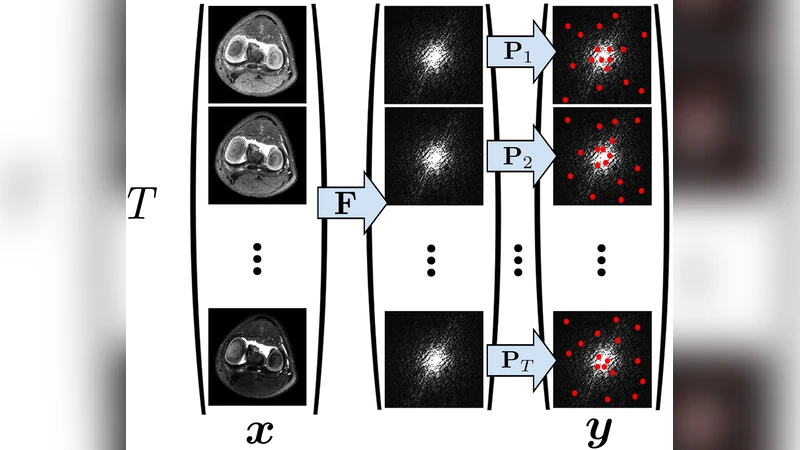
Model‑Based Reconstruction – Instead of first reconstructing a time‑series of images and then fitting T1/T2 maps, the method solves a joint non‑linear least‑squares problem that directly estimates (ρ,θ) from the raw k‑space data. The objective combines a data‑consistency term ‖E·x−y‖² (where E includes Fourier encoding and sampling masks for each echo) with regularization on the parameter maps (e.g., spatial TV). Optimization is performed with algorithms such as ADMM or variable‑splitting, leveraging the differentiability of the Bloch/EPG model.
-
Multi‑Contrast Imaging – By acquiring multiple echo times or varying sequence parameters within a single scan, the same dataset yields proton‑density‑weighted, T1‑weighted, and T2‑weighted images simultaneously. The physics‑based reconstruction automatically separates these contrasts, eliminating the need for separate scans.
-
Parameter‑Optimized Sampling – The framework uses the physical model to design optimal RF flip‑angle schedules, echo spacing, and k‑space sampling patterns. Techniques such as Cramér‑Rao bound analysis or simulation‑based optimization guide the selection, ensuring that the acquired data are maximally informative for both image quality and quantitative accuracy.
The authors validate the method on several anatomical sites (foot, knee, brain) using a conventional 12‑minute spin‑echo reference and a highly accelerated 48‑second fast spin‑echo (FSE) acquisition. When reconstructed with the physics‑constrained CS algorithm, the accelerated data exhibit markedly reduced blurring compared with standard CS reconstructions, and the resulting T1/T2 maps closely match the reference values. Moreover, the same acquisition provides high‑quality PD‑, T1‑, and T2‑weighted images, demonstrating the multi‑contrast capability.
The paper highlights several advantages: (i) reduced scan time without sacrificing spatial resolution, (ii) simultaneous acquisition of multiple contrasts, (iii) direct quantitative mapping from undersampled data, and (iv) a principled way to tailor scan parameters to the desired clinical question. It also acknowledges challenges, including the computational burden of non‑linear optimization, the need for accurate modeling of system imperfections (e.g., B1 inhomogeneity), and the extension of the approach to more complex contrasts such as diffusion or chemical exchange.
In summary, by marrying physics‑based signal evolution models with compressed sensing, the authors deliver a powerful, generalizable strategy for fast, artifact‑robust, and quantitatively accurate MRI. This work paves the way toward personalized, quantitative imaging protocols that can be integrated into routine clinical practice.
Comments & Academic Discussion
Loading comments...
Leave a Comment