Thyroid Cancer Malignancy Prediction From Whole Slide Cytopathology Images
We consider preoperative prediction of thyroid cancer based on ultra-high-resolution whole-slide cytopathology images. Inspired by how human experts perform diagnosis, our approach first identifies and classifies diagnostic image regions containing informative thyroid cells, which only comprise a tiny fraction of the entire image. These local estimates are then aggregated into a single prediction of thyroid malignancy. Several unique characteristics of thyroid cytopathology guide our deep-learning-based approach. While our method is closely related to multiple-instance learning, it deviates from these methods by using a supervised procedure to extract diagnostically relevant regions. Moreover, we propose to simultaneously predict thyroid malignancy, as well as a diagnostic score assigned by a human expert, which further allows us to devise an improved training strategy. Experimental results show that the proposed algorithm achieves performance comparable to human experts, and demonstrate the potential of using the algorithm for screening and as an assistive tool for the improved diagnosis of indeterminate cases.
💡 Research Summary
This paper addresses the challenging problem of pre‑operative thyroid cancer prediction from ultra‑high‑resolution whole‑slide cytopathology images (WSIs). The authors assembled a large, clinically realistic dataset of 908 fine‑needle aspiration (FNA) slides, each scanned at 40× magnification (≈150 k × 100 k pixels, 0.25 µm/pixel) and paired with a post‑operative histopathology diagnosis (ground‑truth malignancy) and the pre‑operative Bethesda System (TBS) category assigned by a cytopathologist. Because a single slide occupies tens of gigabytes, the authors down‑sampled the central focal plane by a factor of four and processed only that plane for the current study.
The proposed solution is a two‑stage cascade of convolutional neural networks (CNNs). Stage 1 is a VGG‑11‑based classifier trained to distinguish image patches that contain groups of follicular (thyroid) cells from background patches (blood, debris, empty glass). Supervision for this stage comes from a modest set of expert‑annotated positive patches: 5,461 regions across 142 slides. Negative examples are sampled randomly from the rest of the slides. This supervised detection step overcomes the limitations of classic multiple‑instance learning (MIL), where the tiny fraction of diagnostically relevant patches is often drowned out by overwhelming background.
Stage 2 aggregates the features of the patches flagged as follicular by Stage 1 to produce a slide‑level prediction. Uniquely, the network simultaneously predicts (i) the binary malignancy label Y∈{0,1} and (ii) the ordinal TBS score S∈{2,3,4,5,6}. The authors implement this as a single scalar output compared against a set of learned thresholds, i.e., a cumulative‑link ordinal regression model. Because the TBS categories are monotonic with respect to malignancy risk (higher TBS → higher malignancy probability), joint training acts as a regularizer: the model is encouraged to produce outputs that are consistent across both tasks, improving robustness especially for the indeterminate TBS 3‑5 cases.
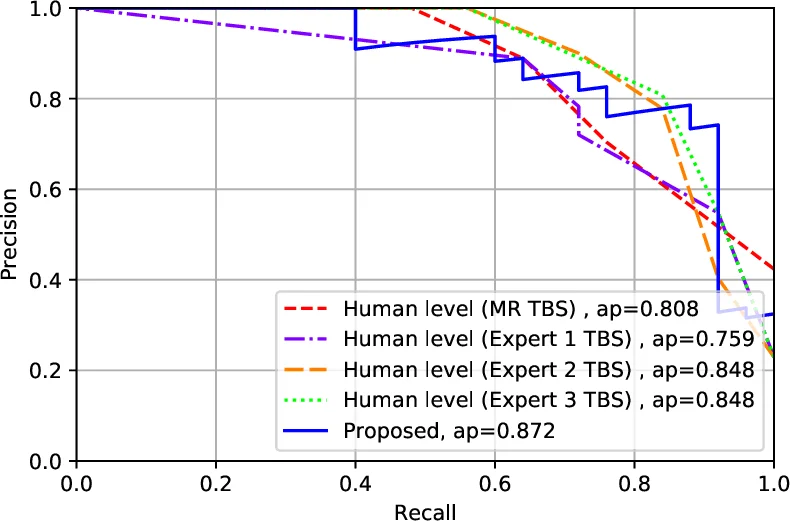
Training uses a combined loss that penalizes errors on both Y and S. The network is trained on the 799‑slide training set (the remainder of the 908 after reserving 109 for testing). The test set was selected chronologically, reviewed by three independent cytopathologists, and excluded slides that were out‑of‑focus or lacked sufficient follicular groups. The final evaluation on these 109 unseen slides shows that the algorithm achieves an area‑under‑the‑ROC curve comparable to three expert cytopathologists who examined the same digital slides. Notably, all slides for which the model predicted TBS 2 or TBS 6 were indeed benign or malignant, respectively, yielding 100 % accuracy on the two most decisive categories. Moreover, the model was able to re‑classify a subset of indeterminate (TBS 3‑5) cases into the definitive categories, suggesting a potential role in reducing unnecessary surgeries or follow‑up procedures.
Technical contributions include: (1) a supervised follicular‑cell detector that efficiently isolates the diagnostically relevant minority of image patches; (2) a novel joint malignancy‑and‑TBS prediction framework that leverages ordinal regression to enforce clinical monotonicity; (3) a practical pipeline that operates on down‑sampled, single‑plane WSIs while still achieving near‑human performance; and (4) the creation of a publicly relevant, 20 TB‑scale thyroid cytopathology dataset.
Clinically, the system could serve as a screening aid, flagging slides that merit closer human review and providing decision support for indeterminate cases. Future work is outlined to incorporate the full 9‑plane Z‑stack, explore deeper architectures or attention mechanisms for better feature aggregation, and extend the methodology to other cytopathology domains. The authors also suggest active‑learning strategies to reduce the annotation burden further.
In summary, the paper demonstrates that with careful region detection, joint ordinal‑regression training, and a realistic large‑scale dataset, deep learning can reach expert‑level accuracy in predicting thyroid cancer malignancy from whole‑slide cytopathology images, opening the door to practical computer‑assisted diagnosis in pathology laboratories.
Comments & Academic Discussion
Loading comments...
Leave a Comment