Towards context in large scale biomedical knowledge graphs
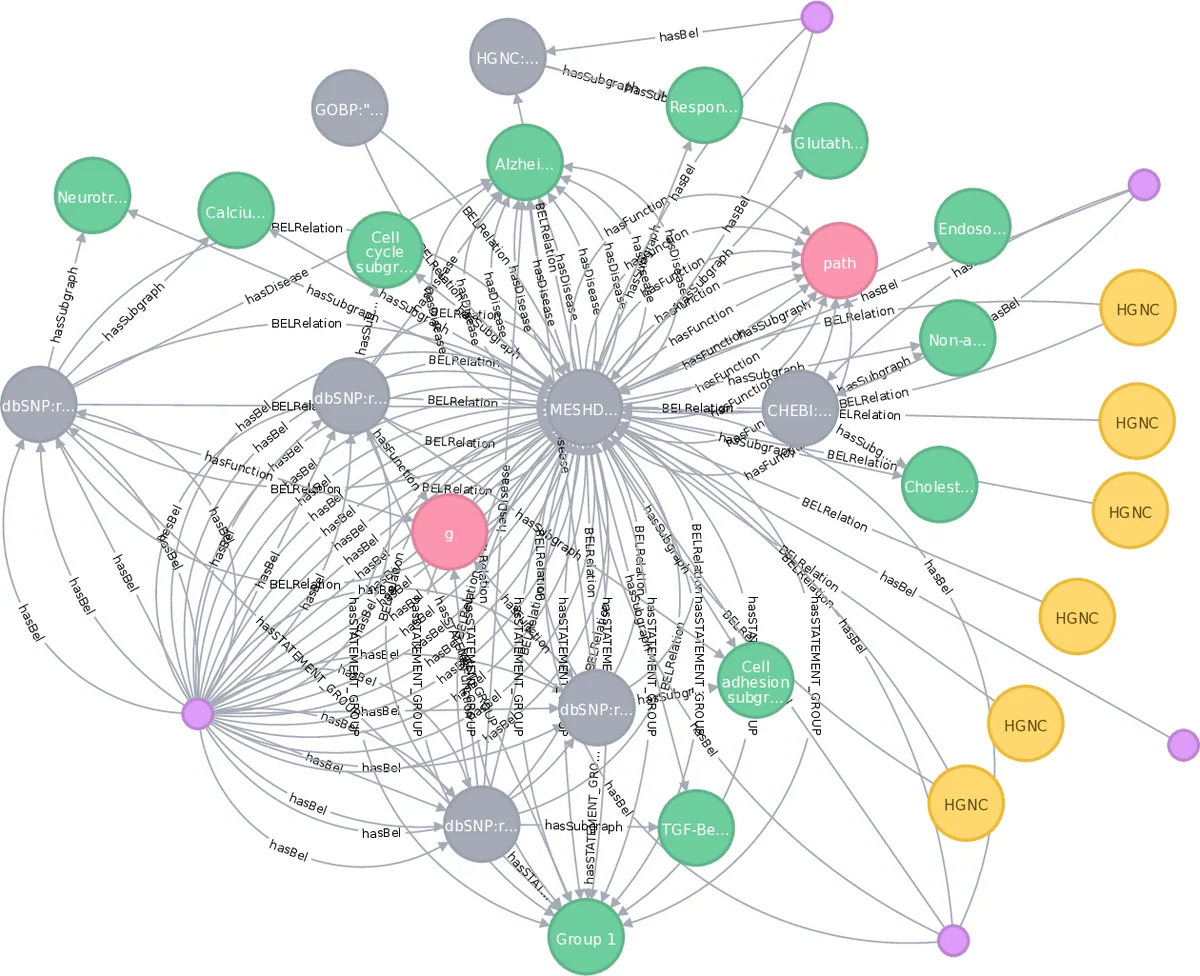
Contextual information is widely considered for NLP and knowledge discovery in life sciences since it highly influences the exact meaning of natural language. The scientific challenge is not only to extract such context data, but also to store this data for further query and discovery approaches. Here, we propose a multiple step knowledge graph approach using labeled property graphs based on polyglot persistence systems to utilize context data for context mining, graph queries, knowledge discovery and extraction. We introduce the graph-theoretic foundation for a general context concept within semantic networks and show a proof-of-concept based on biomedical literature and text mining. Our test system contains a knowledge graph derived from the entirety of PubMed and SCAIView data and is enriched with text mining data and domain specific language data using BEL. Here, context is a more general concept than annotations. This dense graph has more than 71M nodes and 850M relationships. We discuss the impact of this novel approach with 27 real world use cases represented by graph queries.
💡 Research Summary
This paper presents a novel framework for integrating and leveraging contextual information within large-scale biomedical knowledge graphs (KGs). Recognizing that context is crucial for disambiguating meaning in natural language and driving knowledge discovery, the authors address the dual challenge of extracting contextual data and storing it for efficient querying and analysis.
The core proposal is a multi-step KG construction and enrichment workflow based on labeled property graphs, moving beyond the limitations of traditional RDF triple stores. The process begins with a document graph built from PubMed metadata (authors, journals, MeSH terms). This initial graph is then iteratively enriched with contextual information derived from text mining (using tools like JProMiner for named entity recognition and relation extraction) and formalized biological knowledge expressed in the Biological Expression Language (BEL). This creates a virtuous cycle where newly extracted knowledge provides context for subsequent, more refined text mining analyses.
To manage the immense scale of the resulting graph (over 71 million nodes and 850 million relationships), the authors implement a polyglot persistence architecture, primarily utilizing the Neo4j graph database for its native graph storage and support for complex traversals.
The major theoretical contribution lies in the formal, graph-theoretic modeling of context. The authors define:
- Context (C) as a set assignable to any node or relationship in the KG via a mapping function
con(). - Extended Context Subgraph, which encompasses a set of entities and all their neighboring context nodes.
- Context Metagraph (M), a higher-level graph where each node represents a context from the original KG, and edges indicate connections between those contexts within the KG.
By linking the original KG (G) and the metagraph (M), a hypergraph structure (H_G|M) is formed. This structure allows for powerful, context-aware operations, such as filtering graph paths relevant to a specific demographic or experimental condition, or discovering hidden relationships between different contextual domains (e.g., linking molecular mechanisms to specific comorbidities).
The paper demonstrates the practical utility of this framework through 27 real-world use cases formulated as graph queries. These examples showcase the potential for context-aware KGs to answer complex biomedical questions, generate novel hypotheses, and enhance applications in natural language processing and machine learning by providing rich, structured, and semantically grounded training data. Ultimately, the work advocates for a shift from static knowledge repositories to dynamic, context-sensitive systems that actively support the process of scientific discovery.
Comments & Academic Discussion
Loading comments...
Leave a Comment