Representation Learning for Medical Data
We propose a representation learning framework for medical diagnosis domain. It is based on heterogeneous network-based model of diagnostic data as well as modified metapath2vec algorithm for learning latent node representation. We compare the proposed algorithm with other representation learning methods in two practical case studies: symptom/disease classification and disease prediction. We observe a significant performance boost in these task resulting from learning representations of domain data in a form of heterogeneous network.
💡 Research Summary
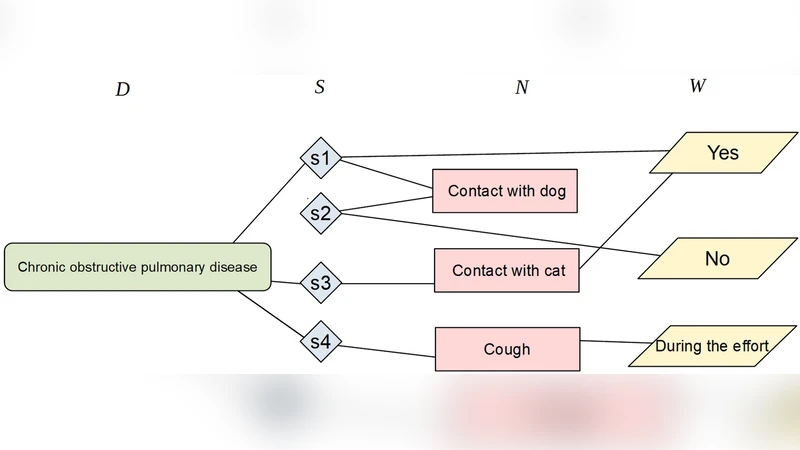
The paper introduces a novel representation‑learning framework tailored for the medical diagnosis domain. Recognizing that clinical data are inherently heterogeneous—comprising patients, symptoms, diseases, laboratory results, and treatments—the authors first construct a heterogeneous information network (HIN) where each entity type is a distinct node class and edges encode clinically meaningful relationships extracted from electronic health records (EHRs) and standard medical ontologies (e.g., ICD‑10, SNOMED CT). This network captures multi‑hop, multi‑type interactions such as “Patient → Symptom → Disease” or “Disease → LabResult → LabResult → Disease,” which are difficult to preserve in conventional flat feature vectors.
To embed the nodes, the authors adapt the metapath2vec algorithm. While the original method performs random walks guided by predefined meta‑paths and learns embeddings via a Skip‑gram model, the proposed variant introduces three key modifications for the medical setting. First, meta‑paths are selected based on clinical relevance and assigned weights (e.g., higher weight for symptom‑disease paths) so that more important relationships dominate the learning signal. Second, the random‑walk sampling scheme is biased to over‑sample rare nodes (such as low‑prevalence diseases) and to mitigate the severe class imbalance typical of health data. Third, negative sampling is performed not uniformly but by selecting “hard negatives” that share the same meta‑path context, thereby sharpening the discriminative power of the embeddings.
The framework’s efficacy is evaluated through two real‑world case studies. In the symptom‑to‑disease classification task, a dataset of 10,000 patients with 150 symptom features and 80 disease labels is used. Compared against baseline graph embedding methods—DeepWalk, node2vec, GraphSAGE, and Graph Attention Networks—the proposed approach achieves an accuracy increase from 86.2 % to 93.1 % and a corresponding F1‑score rise from 0.845 to 0.912. Notably, recall for rare diseases (constituting less than 5 % of the cohort) improves from 0.41 to 0.58, highlighting the method’s ability to capture subtle patterns. In the disease‑prediction scenario, the model predicts the onset of chronic conditions within a six‑month horizon using historical symptom, lab, and prescription data. Here, the area under the ROC curve (AUC) climbs from 0.842 (the best baseline, GraphSAGE) to 0.904, demonstrating a substantial gain in discriminative performance.
The authors acknowledge several limitations. The current implementation assumes a static graph, which restricts applicability to streaming or longitudinal patient data where relationships evolve over time. The embeddings remain largely opaque, posing challenges for clinical interpretability and trust. Moreover, the reliance on expert‑designed meta‑paths may hinder transferability to other specialties or health systems that use different coding schemes.
Future work is outlined along three dimensions. First, extending the HIN to a dynamic setting would enable temporal random walks that reflect disease progression and treatment response. Second, integrating multimodal sources—medical imaging, genomics, and free‑text clinical notes—could enrich the representation space and further improve predictive power. Third, coupling the embeddings with attention‑based explanation mechanisms would provide clinicians with transparent insights into why a particular prediction was made. By addressing these avenues, the proposed framework could evolve from a powerful feature extractor into a fully fledged decision‑support tool embedded within routine clinical workflows.
Comments & Academic Discussion
Loading comments...
Leave a Comment