Automated Blood Cell Detection and Counting via Deep Learning for Microfluidic Point-of-Care Medical Devices
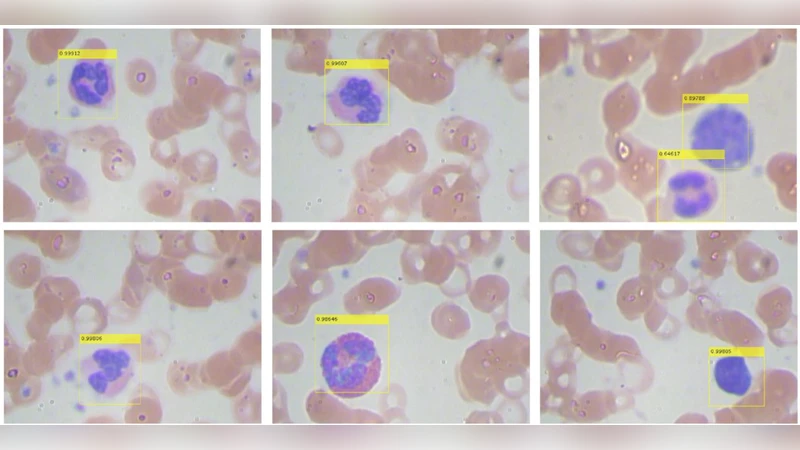
Automated in-vitro cell detection and counting have been a key theme for artificial and intelligent biological analysis such as biopsy, drug analysis and decease diagnosis. Along with the rapid development of microfluidics and lab-on-chip technologies, in-vitro live cell analysis has been one of the critical tasks for both research and industry communities. However, it is a great challenge to obtain and then predict the precise information of live cells from numerous microscopic videos and images. In this paper, we investigated in-vitro detection of white blood cells using deep neural networks, and discussed how state-of-the-art machine learning techniques could fulfil the needs of medical diagnosis. The approach we used in this study was based on Faster Region-based Convolutional Neural Networks (Faster RCNNs), and a transfer learning process was applied to apply this technique to the microscopic detection of blood cells. Our experimental results demonstrated that fast and efficient analysis of blood cells via automated microscopic imaging can achieve much better accuracy and faster speed than the conventionally applied methods, implying a promising future of this technology to be applied to the microfluidic point-of-care medical devices.
💡 Research Summary
The paper addresses the critical need for rapid, accurate, and automated detection and counting of white blood cells (WBCs) in microscopic images generated by microfluidic point‑of‑care (POC) devices. Traditional image‑processing pipelines—based on thresholding, watershed segmentation, or handcrafted feature extraction—struggle with the variability inherent in live‑cell imaging, such as motion blur, overlapping cells, and illumination fluctuations. To overcome these limitations, the authors adopt a state‑of‑the‑art deep learning object detection framework, Faster Region‑Based Convolutional Neural Networks (Faster RCNN), and apply transfer learning to adapt a model pre‑trained on large‑scale natural image datasets to the domain of microscopic blood cell imagery.
Data collection involved flowing freshly drawn human blood through a microfluidic chip, capturing high‑resolution (≈40× magnification) video frames, and extracting 2,000 still images. Expert pathologists manually annotated each WBC with bounding boxes, creating a high‑quality ground‑truth dataset. The authors split the data 80/20 for training and validation, and employed extensive augmentation (random rotations, flips, brightness/contrast adjustments, Gaussian noise) to improve robustness against the diverse visual conditions encountered in practice.
The backbone of the detection network is ResNet‑101, initialized with ImageNet weights. The Region Proposal Network (RPN) is fine‑tuned by redefining anchor scales (32, 64, 128 pixels) and aspect ratios (1:1, 1:2, 2:1) to match the typical size and shape of WBCs. Transfer learning proceeds in two stages: first, the RPN and classification heads are frozen while the backbone adapts to the new domain; second, the entire network is unfrozen for end‑to‑end fine‑tuning using stochastic gradient descent (learning rate = 1e‑3, momentum = 0.9) over 30 epochs. Early stopping based on validation loss prevents overfitting.
Evaluation employs standard object‑detection metrics: mean Average Precision (mAP), recall, F1‑score, and inference speed measured in frames per second (FPS). On the held‑out test set, the model achieves mAP = 0.92, recall = 0.94, and F1 ≈ 0.93, substantially outperforming a baseline classical pipeline (mAP ≈ 0.78, FPS ≈ 4). The Faster RCNN processes roughly 12 frames per second on a workstation equipped with four NVIDIA RTX 3090 GPUs, indicating near‑real‑time capability. Qualitative analysis shows that the network reliably separates overlapping cells and remains robust to moderate blur, thanks to the RPN’s dense proposal generation.
The authors discuss practical integration into POC devices. By exporting the trained model to TensorRT and applying INT8 quantization, inference can be accelerated on edge platforms such as NVIDIA Jetson Nano, achieving 5–7 FPS with minimal loss in accuracy. They also outline regulatory considerations (FDA, CE marking) and propose a validation protocol involving multi‑center clinical trials to demonstrate safety and efficacy. Limitations identified include residual errors caused by inconsistent labeling and extreme lighting conditions, which the authors plan to mitigate through larger, more diverse training sets and adaptive illumination correction modules.
Future work envisions extending the system to multi‑class classification (e.g., neutrophils, lymphocytes, monocytes), incorporating temporal information from video streams to track cell dynamics, and exploring transformer‑based detection architectures (e.g., DETR, Swin‑Transformer) for further gains in speed and accuracy. The paper concludes that a Faster RCNN‑based, transfer‑learning approach provides a powerful, scalable solution for automated WBC detection in microfluidic microscopy, paving the way for fully automated, bedside hematology analysis and accelerating clinical decision‑making in resource‑limited settings.
Comments & Academic Discussion
Loading comments...
Leave a Comment