Extracting temporal features into a spatial domain using autoencoders for sperm video analysis
In this paper, we present a two-step deep learning method that is used to predict sperm motility and morphology-based on video recordings of human spermatozoa. First, we use an autoencoder to extract temporal features from a given semen video and plot these into image-space, which we call feature-images. Second, these feature-images are used to perform transfer learning to predict the motility and morphology values of human sperm. The presented method shows it’s capability to extract temporal information into spatial domain feature-images which can be used with traditional convolutional neural networks. Furthermore, the accuracy of the predicted motility of a given semen sample shows that a deep learning-based model can capture the temporal information of microscopic recordings of human semen.
💡 Research Summary
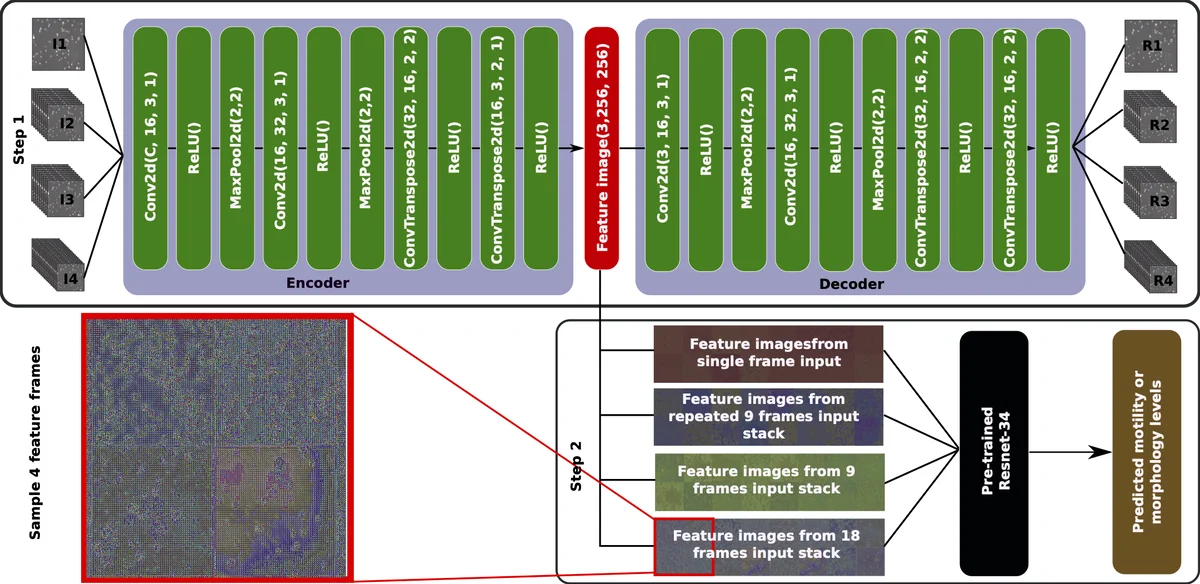
The paper presents a two‑stage deep learning pipeline designed to predict sperm motility and morphology from microscopic video recordings, as part of the 2019 MediaEval Medico task. In the first stage, an unsupervised autoencoder is trained to compress temporal information from video frames into a spatial representation called a “feature‑image”. Four different input configurations are explored: I1 – a single raw frame; I2 – the same frame duplicated across three channels; I3 – a stack of nine consecutive frames; and I4 – a stack of eighteen consecutive frames. The encoder maps each input to a 256 × 256 feature‑image, while the decoder reconstructs the original input, using mean‑squared‑error (MSE) loss. After 2,000 training epochs, only the encoder part is retained for the second stage.
In the second stage, the extracted feature‑images are fed into a pre‑trained ResNet‑34 model whose final fully‑connected layer is replaced to output three regression values, either the three motility percentages (progressive, non‑progressive, immotile) or the three morphology percentages (head, midpiece, tail defects). Transfer learning allows the model to benefit from ImageNet‑trained weights despite the modest dataset size (85 videos). A three‑fold cross‑validation, following the splits supplied by the organizers, is used to evaluate performance.
Results (Table 1) show that the baseline configurations (I1 and I2) achieve mean absolute errors (MAE) of roughly 13 for motility, indicating limited temporal information. When multiple frames are stacked, performance improves markedly: I3 (nine frames) reduces motility MAE to about 10.8, and I4 (eighteen frames) further lowers it to 9.46. This trend confirms that simultaneous observation of several frames captures the dynamic behavior of sperm, which is essential for accurate motility estimation. In contrast, morphology prediction exhibits similar MAE (≈5.6) across all input types, reflecting that morphological assessment can be reliably performed on a single static image.
The authors discuss several limitations and future directions. The current autoencoder optimises only reconstruction loss, which does not explicitly enforce temporal consistency. They suggest experimenting with variational autoencoders, generative adversarial networks, or custom loss terms that penalise temporal incoherence. Incorporating recurrent units (LSTM/GRU) either within the autoencoder or as a post‑processing step could further enhance the capture of motion dynamics. Comparative studies with conventional video features such as optical flow would also help quantify the added value of feature‑images.
In summary, the study introduces a novel method of converting video‑level temporal cues into spatial feature‑images that can be processed by standard CNNs. The approach demonstrates that, even with a limited number of annotated videos, transfer learning combined with unsupervised temporal feature extraction can achieve competitive motility prediction accuracy, while morphology prediction remains robust across input configurations. The work opens avenues for more sophisticated temporal representation learning in biomedical video analysis.
Comments & Academic Discussion
Loading comments...
Leave a Comment