LabPipe: an extensible informatics platform to streamline management of metabolomics data and metadata
Summary: Data management in clinical metabolomics studies is often inadequate. To improve this situation we created LabPipe to provide a guided, customisable approach to study-specific sample collection. It is driven through a local client which manages the process and pushes local data to a remote server through an access controlled web API. The platform is able to support data management for different sampling approaches across multiple sites / studies and is now an essential study management component for supporting clinical metabolomics locally at the EPSRC/MRC funded East Midlands Breathomics Pathology Node. Availability and Implementation: LabPipe is freely available to download under a non-commercial open-source license (NPOSL 3.0) along with documentation and installation instructions at http://labpipe.org. Contact: rob.free@le.ac.uk
💡 Research Summary
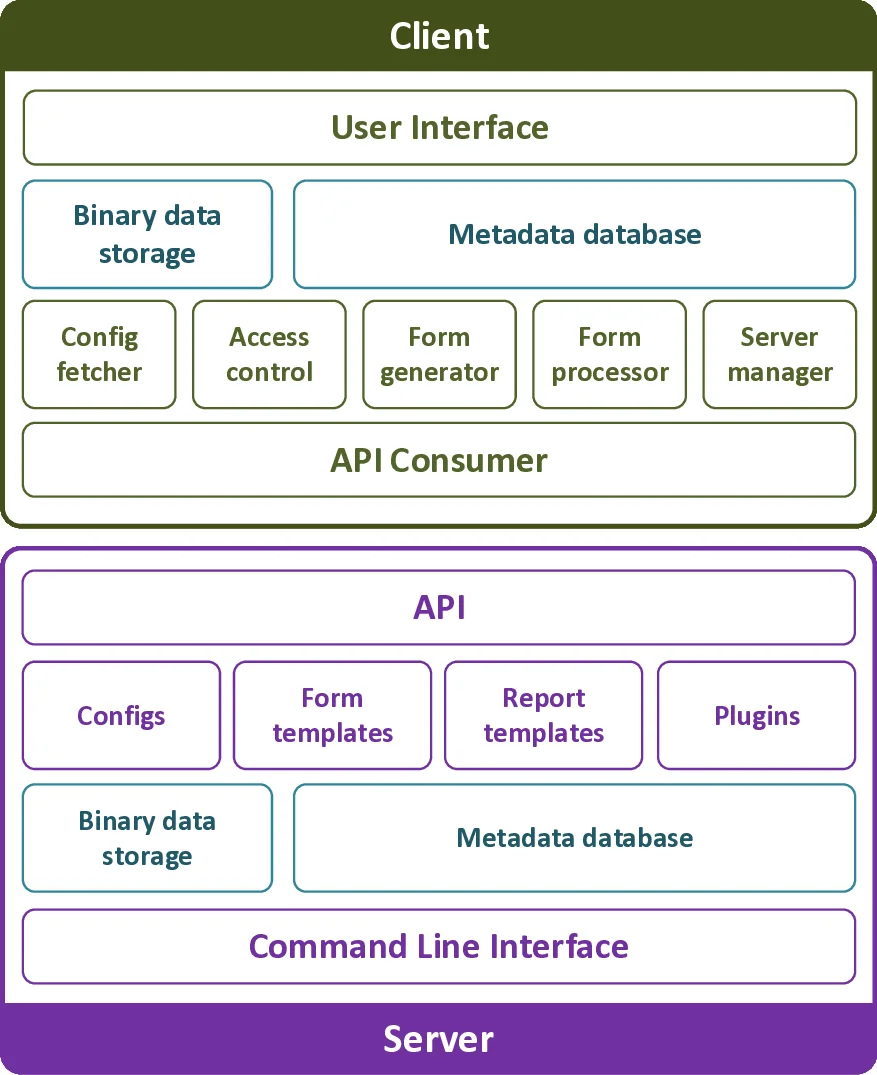
LabPipe is an extensible informatics platform designed to streamline the collection, annotation, and transfer of metabolomics data and associated metadata in multi‑site clinical studies. The system follows a modular client‑server architecture: a lightweight web‑service API (LPserver) provides role‑based access control and stores metadata in a NoSQL database, allowing flexible schema evolution and rapid loading of study‑specific configurations expressed in JSON. The client component (LPclient), built with the cross‑platform Electron framework, runs locally on Windows, macOS, or Linux machines and presents users with dynamically generated data‑entry forms based on server‑provided templates.
Key technical features include automatic generation of file identifiers from entered metadata, real‑time detection of new or modified data files, and a built‑in fallback queue that stores files locally when network connectivity is intermittent, automatically synchronising them once the connection is restored. A plugin architecture enables developers to add custom functionality such as email notifications, data validation rules, or integration with external analysis pipelines without modifying the core codebase.
The platform was piloted in the East Midlands Breathomics Pathology Node (EMBER) study, a large‑scale, multi‑disciplinary breathomics project funded by the EPSRC and MRC. EMBER involved two clinical sites, four distinct breath‑sampling approaches (both online real‑time analysis and offline collection), and nine analytical instruments. Prior to LabPipe, data handling relied on paper case‑report forms and manual USB transfers, leading to frequent transcription errors, data loss, and substantial staff overhead. After deployment, clinical staff used LPclient to enter sample metadata at the point of collection; the client automatically linked this metadata to the corresponding raw data files and pushed them to the central server. Researchers gained immediate, web‑based access to a unified repository of metabolomics data and metadata, facilitating rapid downstream statistical analysis and integration with clinical records.
Compared with existing solutions such as i2b2, tranSMART, REDCap, OpenSpecimen, and LabKey, LabPipe uniquely combines local file‑system integration, automated data‑file linking, and a highly configurable form‑generation engine. While those tools excel in data warehousing, electronic case‑report forms, or biobank management, none provide the seamless, semi‑automated workflow required for real‑time laboratory data capture across heterogeneous instruments.
The authors acknowledge current limitations, including the need for robust scaling strategies under heavy concurrent upload loads, deeper support for community‑adopted metadata standards (e.g., ISA‑Tab), and more granular audit‑logging for regulatory compliance. Future development plans aim to extend LabPipe beyond metabolomics to other laboratory modalities such as flow cytometry and real‑time PCR, incorporate ontology‑driven metadata annotation, and implement automatic conversion pipelines to FAIR‑compliant formats.
In conclusion, LabPipe delivers a user‑friendly, configurable, and open‑source solution that reduces manual data handling, minimizes errors, and accelerates data availability in clinical metabolomics research. Its successful integration into multiple ongoing studies at the University of Leicester demonstrates its practicality and potential for broader adoption across biomedical research domains. The software is freely available under the Non‑Commercial Open Source License (NPOSL 3.0) at http://labpipe.org, accompanied by full documentation and installation instructions.
Comments & Academic Discussion
Loading comments...
Leave a Comment