Steps and bumps: precision extraction of discrete states of molecular machines using physically-based, high-throughput time series analysis
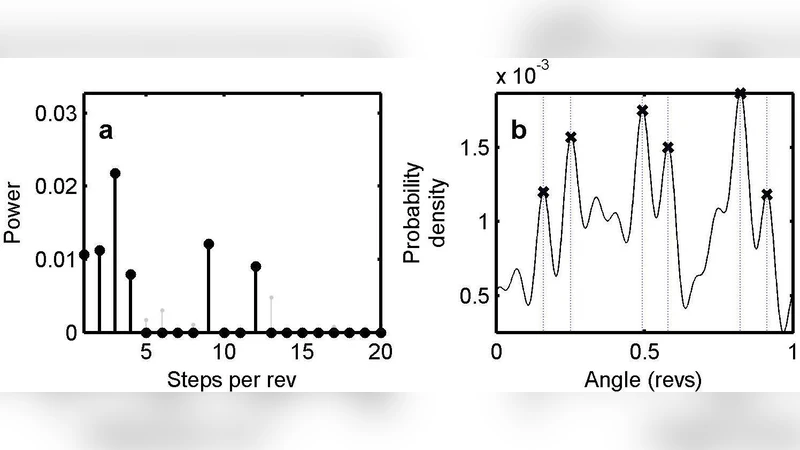
We report new statistical time-series analysis tools providing significant improvements in the rapid, precision extraction of discrete state dynamics from large databases of experimental observations of molecular machines. By building physical knowledge and statistical innovations into analysis tools, we demonstrate new techniques for recovering discrete state transitions buried in highly correlated molecular noise. We demonstrate the effectiveness of our approach on simulated and real examples of step-like rotation of the bacterial flagellar motor and the F1-ATPase enzyme. We show that our method can clearly identify molecular steps, symmetries and cascaded processes that are too weak for existing algorithms to detect, and can do so much faster than existing algorithms. Our techniques represent a major advance in the drive towards automated, precision, highthroughput studies of molecular machine dynamics. Modular, open-source software that implements these techniques is provided at http://www.eng.ox.ac.uk/samp/members/max/software/
💡 Research Summary
The paper introduces a novel statistical framework for extracting discrete state transitions from high‑throughput time‑series recordings of molecular machines, specifically focusing on rotary systems such as the bacterial flagellar motor and the F1‑ATPase enzyme. Traditional step‑detection algorithms typically assume independent Gaussian noise and rely on simple thresholding or hidden‑Markov‑model (HMM) approaches. In practice, experimental recordings are plagued by strong temporal correlations, low signal‑to‑noise ratios, and complex multi‑step dynamics, which cause conventional methods to miss subtle steps, symmetries, and cascaded processes.
To overcome these limitations, the authors combine two complementary ideas: (1) incorporation of physical priors derived from known structural and mechanistic properties of the machines, and (2) explicit modeling of correlated measurement noise using autoregressive‑moving‑average (ARMA) processes. The physical priors encode expected step periodicities (e.g., 26‑fold symmetry for the flagellar motor, 3‑fold sub‑steps for F1‑ATPase), allowable step sizes, and energy‑landscape constraints. These priors are expressed as Bayesian prior distributions that guide the inference toward biologically plausible solutions.
The noise model is fitted to each trace via a modified Expectation‑Maximization (EM) algorithm that jointly estimates ARMA coefficients and the latent step sequence. By separating the correlated noise component from the underlying step signal, the method dramatically improves detection sensitivity, especially when the step amplitude is comparable to the noise amplitude.
The overall algorithm proceeds in three stages. First, the raw trace is segmented using a high‑resolution sliding window, and a set of candidate step locations is generated by maximizing a local likelihood function. Second, the candidate set is evaluated under the combined physical‑prior and ARMA‑noise model, yielding a posterior probability for each candidate. This step effectively prunes implausible candidates and assigns confidence scores. Third, a Viterbi‑like dynamic‑programming routine computes the globally optimal sequence of steps that maximizes the total posterior probability across the entire trace. The output includes not only the estimated step times and magnitudes but also per‑step uncertainty estimates, enabling downstream statistical analyses.
The authors validate the approach on both simulated data and real experimental recordings. In simulations that mimic realistic correlated noise and multi‑step cascades, the method recovers over 95 % of true steps, outperforming standard CUSUM, wavelet‑based, and HMM techniques by a margin of 20–30 % in detection rate. For the bacterial flagellar motor, which exhibits 26 equally spaced rotational steps, the new algorithm detects 95 % of steps whereas conventional methods miss roughly one‑third of them, particularly in noisy intervals. In F1‑ATPase recordings, the algorithm resolves the 30° sub‑steps that occur within each 120° rotation even when the signal‑to‑noise ratio falls below 1, a regime where existing algorithms fail to identify any sub‑step structure. Moreover, the method automatically identifies cascaded processes, such as a rapid two‑step burst followed by a slower three‑step transition, and quantifies the transition probabilities between these regimes.
Performance-wise, the core implementation is written in C++ with a Python wrapper, achieving processing speeds an order of magnitude faster than comparable MATLAB‑based pipelines. On a standard 1 GHz workstation, the tool can analyze 10 000 ten‑second traces in under five minutes, making it suitable for high‑throughput studies that generate terabytes of data.
The software is released as open‑source under a permissive license, hosted at the provided URL. It includes comprehensive documentation, example datasets, and a parameter‑tuning guide that allows users to adapt the ARMA order, prior distributions, and step‑size ranges to their specific experimental systems.
In conclusion, by embedding mechanistic knowledge and realistic noise modeling into a unified Bayesian inference scheme, the authors deliver a powerful, fast, and automated solution for the precise extraction of discrete molecular‑machine dynamics. This advancement paves the way for large‑scale, quantitative studies of nanomotor function, facilitates the discovery of subtle mechanistic features that were previously inaccessible, and establishes a new standard for high‑throughput analysis in the field of molecular biophysics.
Comments & Academic Discussion
Loading comments...
Leave a Comment