Processing Columnar Collider Data with GPU-Accelerated Kernels
At high energy physics experiments, processing billions of records of structured numerical data from collider events to a few statistical summaries is a common task. The data processing is typically more complex than standard query languages allow, such that custom numerical codes are used. At present, these codes mostly operate on individual event records and are parallelized in multi-step data reduction workflows using batch jobs across CPU farms. Based on a simplified top quark pair analysis with CMS Open Data, we demonstrate that it is possible to carry out significant parts of a collider analysis at a rate of around a million events per second on a single multicore server with optional GPU acceleration. This is achieved by representing HEP event data as memory-mappable sparse arrays of columns, and by expressing common analysis operations as kernels that can be used to process the event data in parallel. We find that only a small number of relatively simple functional kernels are needed for a generic HEP analysis. The approach based on columnar processing of data could speed up and simplify the cycle for delivering physics results at HEP experiments. We release the \texttt{hepaccelerate} prototype library as a demonstrator of such methods.
💡 Research Summary
The paper presents a novel approach to processing the massive, structured numerical datasets generated by high‑energy‑physics (HEP) experiments, demonstrating that a large fraction of a typical collider analysis can be executed on a single multicore server at rates approaching one million events per second, with optional GPU acceleration. Traditional HEP workflows rely on ROOT trees, where each event is a complex C++ object containing variable‑length collections of particles. Analyses therefore involve repeated deserialization, decompression, and nested loops over particles, typically distributed across many batch jobs on CPU farms.
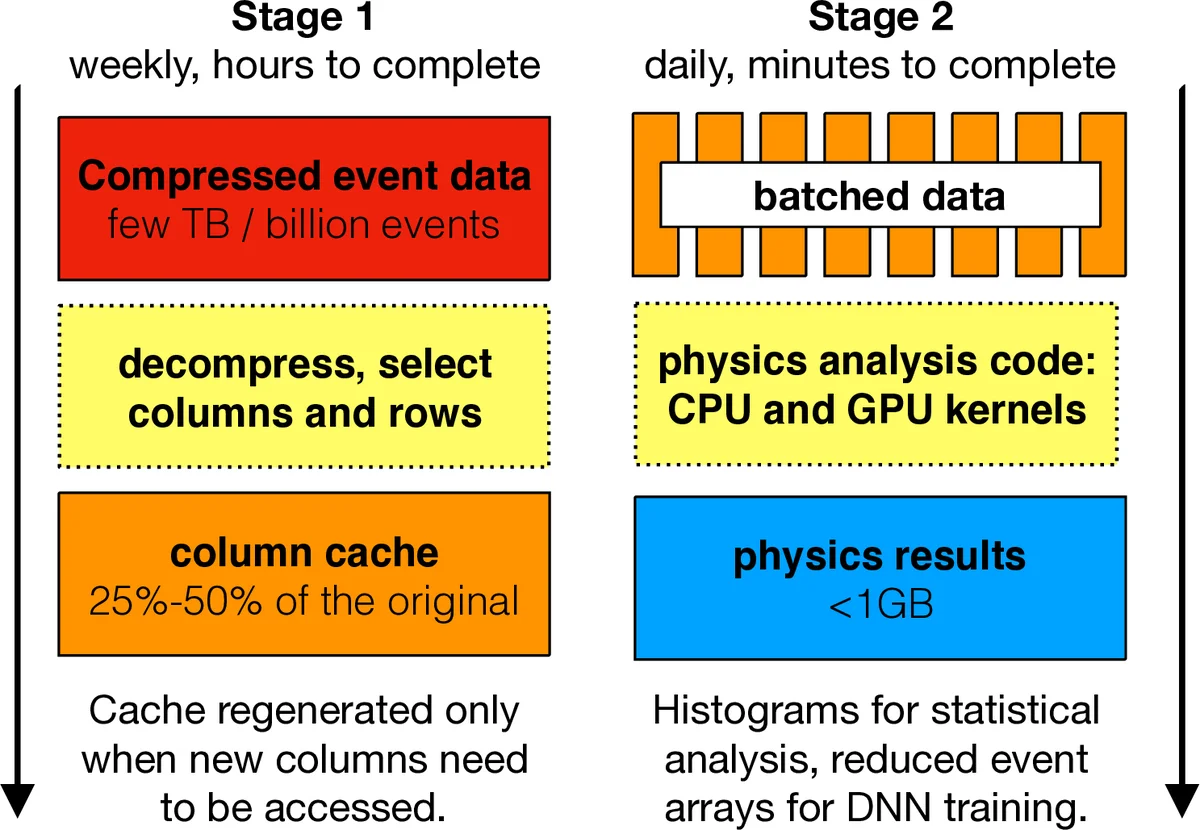
The authors propose to reshape the data into a column‑oriented, memory‑mapped sparse (jagged) format inspired by the uproot and awkward‑array libraries. Each physics variable (e.g., jet pT, η, φ) is stored as a one‑dimensional contiguous array; event boundaries are encoded in an offset array that marks where each event’s collection starts and ends. By selecting only the columns needed for a given analysis and caching them in an uncompressed, memory‑mapped file, the I/O overhead is dramatically reduced (the cache occupies roughly 20‑50 % of the original compressed size).
Computation is expressed as “kernels”: small functions that operate on these arrays. Using the Numba JIT compiler, the same Python code can be compiled to multithreaded CPU code (via prange) or to CUDA GPU kernels. The paper defines a minimal set of generic kernels—get_in_offsets, set_in_offsets, sum_in_offsets, max_in_offsets, min_in_offsets, fill_histogram, and get_bin_contents—that cover the majority of typical HEP operations such as per‑event reductions, histogram filling, and reweighting. Specialized kernels are also introduced for physics‑specific tasks that do not fit neatly into the generic set, such as ΔR‑based masking between two particle collections and selection of opposite‑sign muon pairs. All kernels accept boolean masks for events and particles, allowing selections to be propagated without copying data.
Performance is evaluated on a workstation equipped with a 24‑core Intel Xeon CPU, an NVIDIA GeForce Titan X GPU, and an Optane 900P cache. Using a pre‑loaded dataset of roughly 11 million events (tens of billions of particles), the authors benchmark each kernel across varying numbers of CPU threads and on the GPU. CPU scaling is sub‑linear, achieving up to an 8× speed‑up with 24 threads. GPU execution yields a 20‑30× speed‑up over a single CPU thread, with the most complex ΔR‑masking kernel reaching ~100 MHz (events per second) on the GPU versus 7 MHz on a CPU core.
To demonstrate end‑to‑end feasibility, the authors implement a simplified top‑quark‑pair analysis using CMS Open Data from 2012. The analysis chain includes trigger and MET selection, jet and lepton pT/η cuts, ΔR matching, pile‑up and lepton‑efficiency reweighting via histogram look‑ups, jet‑energy corrections (which increase computational load by ~40×), reconstruction of top‑quark candidates from jet triplets, and evaluation of a machine‑learning discriminator. The entire pipeline processes the 100 GB of selected data in a single pass, achieving an overall throughput of about one million events per second on the GPU‑enabled server—orders of magnitude faster than the conventional multi‑step batch workflow.
The key insights are: (1) columnar, offset‑based representations are memory‑efficient for variable‑length HEP data; (2) a small suite of generic reduction kernels suffices for most analysis needs, simplifying code maintenance; (3) Python + Numba provides a rapid‑prototyping environment while still delivering near‑native performance on both CPUs and GPUs; (4) GPU acceleration offers a practical path to dramatically shorten analysis turn‑around times without requiring large farm resources. The authors release the prototype library hepaccelerate to the community, inviting further development, such as multi‑GPU scaling, integration with modern file formats (e.g., Apache Arrow), and exploration of FPGA or tensor‑core accelerators. In summary, the work demonstrates that rethinking HEP data layout and leveraging modern heterogeneous computing can transform the speed and simplicity of collider data analysis.
Comments & Academic Discussion
Loading comments...
Leave a Comment