An efficient clustering algorithm from the measure of local Gaussian distribution
In this paper, I will introduce a fast and novel clustering algorithm based on Gaussian distribution and it can guarantee the separation of each cluster centroid as a given parameter, $d_s$. The worst run time complexity of this algorithm is approximately $\sim$O$(T\times N \times \log(N))$ where $T$ is the iteration steps and $N$ is the number of features.
💡 Research Summary
**
The paper proposes a novel clustering algorithm that combines a local multivariate Gaussian model with an R‑tree based indexing structure to achieve fast, automatic determination of cluster numbers while guaranteeing a minimum separation between cluster centroids. The method proceeds through five main stages.
-
R‑tree indexing – All data points are inserted into a K‑dimensional R‑tree, enabling average‑case (O(\log N)) range queries and providing the global minima and maxima for each feature.
-
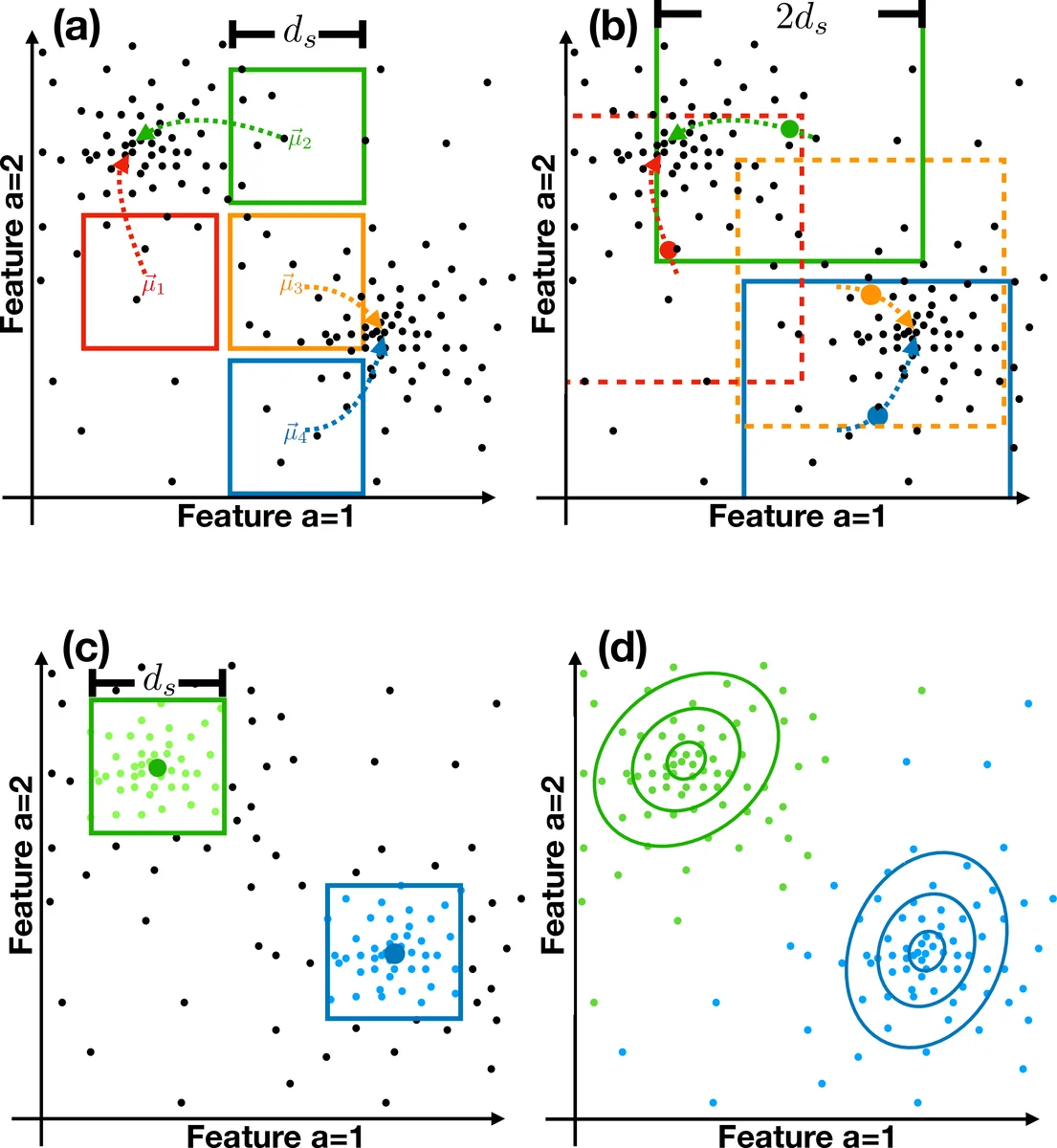
Seeding centroids – A user‑defined separation distance (d_s) defines a regular grid over the feature space. The grid cell centers become provisional centroids (\mu_c^{0}). For each provisional centroid the algorithm queries the R‑tree to collect all points within a hyper‑cube of side length (d_s) (volume (d_s^K)). The number of points (N_c) is recorded and any seed with (N_c < L) (a minimal count threshold) is discarded.
-
Centroid convergence and collision removal – Remaining seeds are iteratively moved to the mean of their local point sets: (\mu_c^{t+1}= \frac{1}{N_c}\sum_{i\in c} x_i). Iteration stops when the Euclidean shift falls below a convergence tolerance (\epsilon). Simultaneously, any two centroids whose distance is less than the collision distance (d_c = 2 d_s) are considered colliding; the one with the smaller local count is removed. This process guarantees that final centroids are at least (d_s) apart, thereby implicitly determining the number of clusters.
-
Weighted covariance estimation – For each converged centroid, a local covariance matrix (\Sigma_c) is estimated. The naïve sample covariance (Eq. 4) is modified by weighting each point with a probability‑based factor (w_{i,c}= \frac{P(x_i|\mu_c)}{\sum_{j\in c} P(x_j|\mu_c)}) and a small regularization constant (\eta). The update rule (Eq. 5) becomes a mean‑field iteration: start with (\Sigma_c^{(0)}) from the unweighted formula, then repeatedly compute a new (\Sigma_c^{(t+1)}) using the weighted expression and a smoothed input (\Sigma^{\text{input}} = \frac{1}{2}(\Sigma^{(t)}+\Sigma^{(t-1)})) until the maximum elementwise change is below (\epsilon). This yields a robust estimate even though only a local subset of points is used.
-
Assignment and post‑processing – Once all (\mu_c) and (\Sigma_c) are fixed, each data point is assigned to the cluster that maximizes its Gaussian likelihood (P(x_i|\mu_c,\Sigma_c)). The authors further propose three optional filters to improve the quality of the output: (i) a probability cut‑off (L_p) that discards points with low likelihood, (ii) a percentage cut‑off (L%) that removes the lowest‑scoring fraction within each cluster, and (iii) a separation cut‑off (L_s) that eliminates points for which the ratio of the top two likelihoods falls below a threshold, thereby reducing ambiguous assignments.
Complexity analysis – The dominant costs are the R‑tree range queries ( (O(N\log N)) ), the centroid convergence loop ( (O(T_{\mu} N_c \log N)) ), and the covariance estimation ( (O(T_{\Sigma} N_c K^3)) ). Because the algorithm works on local neighborhoods, the number of points per centroid (N_c) is typically much smaller than the total (N). The authors argue that in practice the total number of outer iterations (T_{\max}) is below 20, leading to an overall worst‑case bound of (O(T_{\max} N \log N)). They contrast this with Gaussian Mixture Models ( (O(T N^2)) ) and HDBSCAN (roughly (O(N^2 \log N)) ), claiming a substantial speed advantage.
Experimental claim – A single‑threaded C++ implementation (using Boost for the R‑tree) was tested on synthetic data with 150 000 features, completing in under 20 seconds on a 2.5 GHz CPU. No real‑world benchmarks, scalability studies, or statistical evaluations are provided.
Strengths – The method automatically determines the number of clusters through the single parameter (d_s), avoids the quadratic cost of pairwise distance calculations by leveraging spatial indexing, and integrates a locally weighted Gaussian model that can capture cluster shape beyond spherical assumptions. The collision‑based pruning of redundant centroids is simple yet effective.
Weaknesses and open issues –
- The reliance on an R‑tree limits performance in high‑dimensional spaces where tree balance deteriorates (the “curse of dimensionality”).
- Parameter selection ( (d_s, L, \eta, \epsilon) ) is left to the practitioner; no systematic guidance or adaptive scheme is presented.
- The weighting scheme for covariance estimation is recursive and may suffer from numerical instability, especially when local point counts are small.
- No empirical comparison with baseline methods (K‑means, GMM, DBSCAN, HDBSCAN) is shown, making it difficult to assess accuracy, robustness to noise, or sensitivity to outliers.
- The algorithm assumes clusters are roughly Gaussian; performance on arbitrarily shaped or highly overlapping clusters is unclear.
Conclusion – The paper introduces an interesting hybrid approach that blends spatial indexing with a locally weighted Gaussian model to achieve near‑linear runtime clustering while automatically inferring the number of clusters. The conceptual contribution is solid, but the lack of thorough experimental validation, limited discussion of high‑dimensional behavior, and reliance on several heuristic parameters temper the practical impact. Future work should focus on extensive benchmarking, adaptive parameter tuning, and extensions to handle non‑Gaussian or manifold‑structured data.
Comments & Academic Discussion
Loading comments...
Leave a Comment