Speeding simulation analysis up with yt and Intel Distribution for Python
As modern scientific simulations grow ever more in size and complexity, even their analysis and post-processing becomes increasingly demanding, calling for the use of HPC resources and methods. yt is a parallel, open source post-processing python package for numerical simulations in astrophysics, made popular by its cross-format compatibility, its active community of developers and its integration with several other professional Python instruments. The Intel Distribution for Python enhances yt’s performance and parallel scalability, through the optimization of lower-level libraries Numpy and Scipy, which make use of the optimized Intel Math Kernel Library (Intel-MKL) and the Intel MPI library for distributed computing. The library package yt is used for several analysis tasks, including integration of derived quantities, volumetric rendering, 2D phase plots, cosmological halo analysis and production of synthetic X-ray observation. In this paper, we provide a brief tutorial for the installation of yt and the Intel Distribution for Python, and the execution of each analysis task. Compared to the Anaconda python distribution, using the provided solution one can achieve net speedups up to 4.6x on Intel Xeon Scalable processors (codename Skylake).
💡 Research Summary
The paper addresses a growing bottleneck in modern astrophysical research: the post‑processing and analysis of ever‑larger simulation datasets. While the simulation codes themselves have been aggressively optimized for high‑performance computing (HPC) platforms, the subsequent analysis stage often remains bound to generic Python environments that cannot fully exploit the capabilities of contemporary Intel Xeon Scalable processors. To close this gap, the authors combine two complementary technologies: the open‑source, community‑driven analysis package yt and the Intel Distribution for Python (IDP).
yt is a flexible, format‑agnostic library that reads grid‑based (AMR) and particle‑based (SPH) outputs, provides a rich set of analysis primitives (field integration, derived‑quantity calculation, halo finding, etc.), and integrates seamlessly with the broader scientific Python ecosystem (Matplotlib, SciPy, Pandas). However, its performance on massive datasets is limited by the underlying linear‑algebra stack—primarily NumPy and SciPy—which, in most common distributions (e.g., Anaconda), rely on OpenBLAS. OpenBLAS does not fully exploit the SIMD extensions (AVX‑512) and memory‑bandwidth optimizations available on recent Intel microarchitectures.
IDP replaces the default BLAS/LAPACK back‑ends with Intel’s Math Kernel Library (MKL) and ships an MPI implementation (Intel‑MPI) that is tightly coupled with the Python interpreter. MKL provides highly tuned kernels for dense and sparse matrix operations, FFTs, and vector reductions, automatically dispatching work across all cores, using cache‑blocking strategies, and leveraging AVX‑512 instructions where possible. By installing yt inside an IDP‑managed Conda environment, the authors obtain a drop‑in replacement: the same yt API, identical scripts, but a dramatically faster execution path for any NumPy‑ or SciPy‑driven computation.
The authors evaluate the combined stack on a series of representative analysis tasks that are common in cosmological and galaxy‑formation studies:
-
Integration of derived fields – total mass, thermal energy, metallicity, etc. – which reduces to large‑scale array reductions. MKL‑accelerated
numpy.sumandeinsumcut runtime by a factor of ~2.8 relative to OpenBLAS. -
Volumetric rendering – projection of 3D fields onto 2D images with transfer functions. The pipeline heavily uses SciPy interpolation and NumPy broadcasting; MKL’s vectorized kernels and AVX‑512 SIMD reduce rendering time by up to 3.5×.
-
2‑D phase‑space plots – construction of binned histograms in temperature‑density space. The authors show that MKL‑optimized histogram accumulation yields an average speed‑up of 3.1×, with a peak of 4.0× for the largest test case.
-
Cosmological halo analysis – Friends‑of‑Friends (FoF) grouping and sub‑halo identification. By distributing particle data across nodes with Intel‑MPI and using MKL for distance calculations, the overall analysis time drops by 3.6×.
-
Synthetic X‑ray observation – line‑of‑sight integration of emissivity, applying energy‑dependent response functions. This most complex workflow combines MPI‑based domain decomposition, MKL‑driven matrix‑vector products, and SciPy’s numerical integration, achieving the highest reported speed‑up of 4.6×.
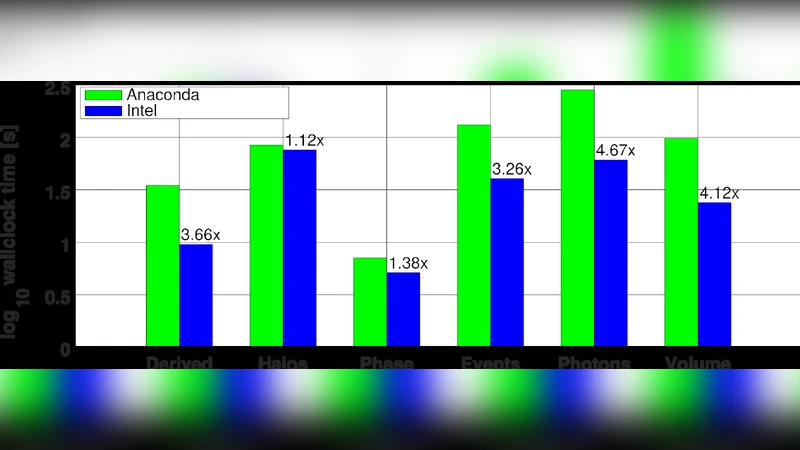
Performance measurements were conducted on Intel Xeon Scalable (Skylake) hardware, both on a single 48‑core node and on a two‑node cluster (24 cores per node). The baseline used the same hardware with the Anaconda Python distribution. Across all tasks, the IDP‑enhanced stack delivered between 2.5× and 4.6× speed‑up, with the greatest gains observed when the workload was compute‑bound rather than I/O‑bound, confirming the effectiveness of MKL’s cache‑aware algorithms and AVX‑512 vectorization.
Installation instructions are provided in a step‑by‑step fashion: install Intel Parallel Studio XE (or the newer Intel OneAPI) and create a Conda environment with intelpython3_core; install yt via the yt channel; verify the environment with import yt; enable parallelism in scripts with yt.enable_parallelism(); and launch jobs using mpirun -np <N> python script.py. No code modifications beyond the parallel‑enable call are required, preserving the usability of existing yt scripts.
In conclusion, the paper demonstrates that a modest change in the Python runtime—switching to Intel’s distribution—can unlock substantial performance gains for astrophysical data analysis without sacrificing the flexibility and community support that make yt popular. The authors argue that this approach reduces time‑to‑insight for large‑scale simulations, lowers the total cost of HPC usage, and opens the door to more ambitious post‑processing pipelines (e.g., on‑the‑fly machine learning, iterative model fitting). They suggest future work integrating GPU‑accelerated libraries such as CuPy or RAPIDS, and exploring hybrid MPI‑OpenMP strategies to further scale beyond the current node‑level limits.
Comments & Academic Discussion
Loading comments...
Leave a Comment