Combining Noise-to-Image and Image-to-Image GANs: Brain MR Image Augmentation for Tumor Detection
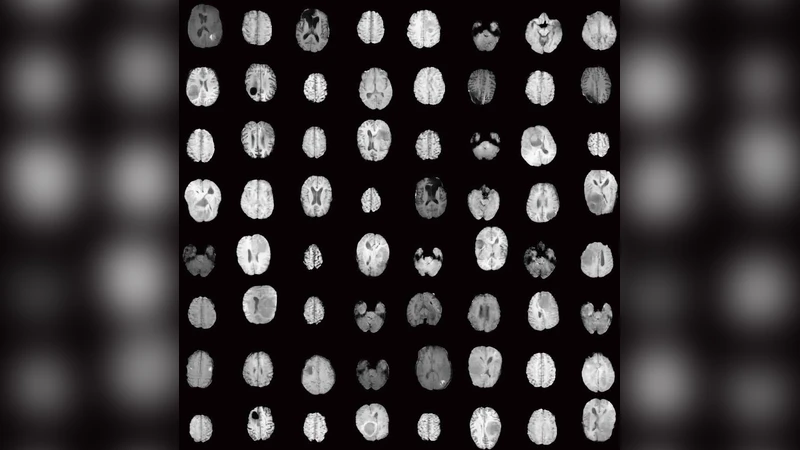
Convolutional Neural Networks (CNNs) achieve excellent computer-assisted diagnosis with sufficient annotated training data. However, most medical imaging datasets are small and fragmented. In this context, Generative Adversarial Networks (GANs) can synthesize realistic/diverse additional training images to fill the data lack in the real image distribution; researchers have improved classification by augmenting data with noise-to-image (e.g., random noise samples to diverse pathological images) or image-to-image GANs (e.g., a benign image to a malignant one). Yet, no research has reported results combining noise-to-image and image-to-image GANs for further performance boost. Therefore, to maximize the DA effect with the GAN combinations, we propose a two-step GAN-based DA that generates and refines brain Magnetic Resonance (MR) images with/without tumors separately: (i) Progressive Growing of GANs (PGGANs), multi-stage noise-to-image GAN for high-resolution MR image generation, first generates realistic/diverse 256 X 256 images; (ii) Multimodal UNsupervised Image-to-image Translation (MUNIT) that combines GANs/Variational AutoEncoders or SimGAN that uses a DA-focused GAN loss, further refines the texture/shape of the PGGAN-generated images similarly to the real ones. We thoroughly investigate CNN-based tumor classification results, also considering the influence of pre-training on ImageNet and discarding weird-looking GAN-generated images. The results show that, when combined with classic DA, our two-step GAN-based DA can significantly outperform the classic DA alone, in tumor detection (i.e., boosting sensitivity 93.67% to 97.48%) and also in other medical imaging tasks.
💡 Research Summary
This paper addresses the pervasive problem of limited and fragmented medical imaging datasets by proposing a two‑step generative adversarial network (GAN) based data augmentation (DA) pipeline specifically designed for brain magnetic resonance imaging (MRI) tumor detection. In the first step, the authors employ Progressive Growing of GANs (PGGAN) to synthesize high‑resolution (256 × 256) whole‑brain MR images from random noise. PGGAN starts training at a low resolution and progressively adds layers, thereby stabilizing training, reducing mode collapse, and enabling the generation of diverse tumor and non‑tumor images. The network is trained with a Wasserstein loss and gradient penalty (λ_gp = 10), using Adam (learning rate = 1 × 10⁻³, β₁ = 0, β₂ = 0.99) for 100 epochs and a batch size of 16. Separate PGGAN models are trained for each class to preserve class‑specific distributions.
The second step refines the texture and anatomical realism of the PGGAN‑generated images using an image‑to‑image translation GAN. Two alternatives are investigated: (1) MUNIT, which combines multimodal variational auto‑encoders with GANs, and (2) SimGAN, a refinement network that adds a self‑regularization term and a history‑based discriminator. For MUNIT, the authors adopt the UNIT loss augmented with a perceptual VGG loss, assigning weights (reconstruction = 10, adversarial = 1, cycle consistency = 10, KL‑divergence = 0.01, perceptual = 1). Training proceeds for 100 000 steps with a batch size of 1, learning rate = 1 × 10⁻⁴, and learning‑rate decay every 20 000 steps. SimGAN is trained with a combined realism loss and regularization term, also for 100 000 steps. Horizontal flipping is used as a simple augmentation during refinement.
After refinement, the synthetic images are combined with classic augmentation (rotations, flips, intensity scaling) and fed into a ResNet‑50 binary classifier for tumor vs. non‑tumor discrimination. The authors explore three important variables: (a) whether the ResNet‑50 backbone is pre‑trained on ImageNet, (b) the impact of discarding “weird‑looking” synthetic images identified by a visual Turing test performed by an expert radiologist, and (c) the effect of the refinement method (MUNIT vs. SimGAN). t‑SNE visualizations demonstrate that refined synthetic images occupy regions of feature space much closer to real images than raw PGGAN outputs, indicating a successful domain alignment.
Experimental results on the BraTS 2016 dataset (240 × 240 T1‑contrast‑enhanced axial slices, 220 high‑grade glioma patients) show that classic augmentation alone yields a sensitivity of 93.67 % for tumor detection. Adding the two‑step GAN‑augmented data raises sensitivity to 97.48 %, a statistically significant improvement. The gain is most pronounced when the ResNet‑50 backbone is ImageNet‑pre‑trained and when low‑quality synthetic images are removed, suggesting that both a strong prior and careful quality control are essential for maximizing benefit from GAN‑generated data.
The paper’s contributions are threefold: (1) Demonstrating that whole‑brain MRI can be generated at high resolution using PGGAN, moving beyond prior work that focused on small regions of interest; (2) Introducing a novel two‑step augmentation strategy that combines noise‑to‑image and image‑to‑image GANs, showing that texture refinement substantially boosts classification performance; (3) Providing a systematic analysis of how pre‑training and synthetic‑image quality affect the utility of GAN‑based augmentation in small, heterogeneous medical datasets.
Future directions proposed include extending the pipeline to multi‑modal imaging (e.g., combining MRI and CT), automating the detection and removal of low‑quality synthetic samples, integrating GAN‑generated labels for semi‑supervised learning, and evaluating the approach in prospective clinical settings to assess real‑world impact on diagnostic workflows.
Comments & Academic Discussion
Loading comments...
Leave a Comment