A Non-Negative Factorization approach to node pooling in Graph Convolutional Neural Networks
The paper discusses a pooling mechanism to induce subsampling in graph structured data and introduces it as a component of a graph convolutional neural network. The pooling mechanism builds on the Non-Negative Matrix Factorization (NMF) of a matrix r…
Authors: Davide Bacciu, Luigi Di Sotto
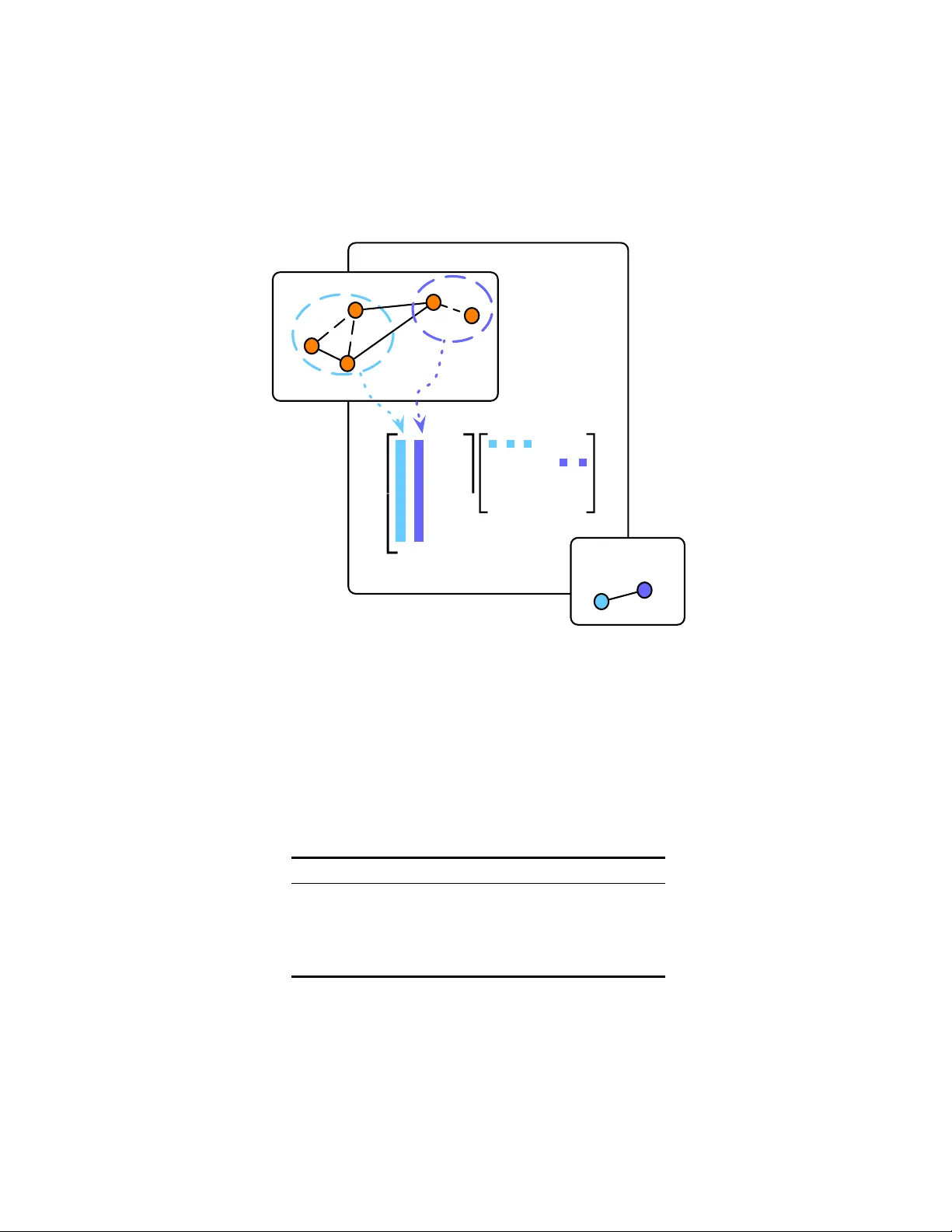
A Non-Negativ e F actorization approac h to no de p o oling in Graph Con v olutional Neural Net w orks Da vide Bacciu and Luigi Di Sotto Univ ersit` a di Pisa Dipartimen to di Informatica Largo B. P ontecorv o, 3 - Pisa (Italy) bacciu@di.unipi.it l.disotto@gmail.com D. Bacciu and L. Di Sotto, A Non-Negativ e F actorization approac h to node p ooling in Graph Con volutional Neural Netw orks T o app e ar in the Pr o c e e dings of the 18th International Confer enc e of the Italian Asso ciation for Artificial Intel ligenc e (AIIA 2019) , Springer, 2019 Abstract. The pap er discusses a po oling mec hanism to induce subsam- pling in graph structured data and introduces it as a comp onen t of a graph conv olutional neural net work. The p ooling mec hanism builds on the Non-Negative Matrix F actorization (NMF) of a matrix representing no de adjacency and no de similarit y as adaptiv ely obtained through the v ertices em b edding learned by the model. Suc h mec hanism is applied to obtain an incremen tally coarser graph where no des are adaptiv ely p ooled into comm unities based on the outcomes of the non-negative fac- torization. The empirical analysis on graph classification b enc hmarks sho ws ho w suc h coarsening pro cess yields significant improv ements in the predictive p erformance of the model with respect to its non-p o oled coun terpart. Keyw ords: Graph Con volutional Neural Net works · Differen tiable Graph P o oling · Non-Negative Matrix F actorization. 1 In tro duction No wada ys many real-w orld phenomena are mo deled as in teracting ob jects p ossi- bly living into high-dimensional manifolds with added top ological structure. Ex- amples can b e found in genomics with protein-protein interaction net works, fak e news discov ery in so cial net works, functional net works in neuroscience. Graphs are the natural mathematical mo del for such data with underlying non-Euclidean nature. Current Euclidean Con volutional Neural Netw orks hav e built their suc- cess lev eraging on the statistical prop erties of stationarity , lo calit y and comp o- sitionalit y of flat domains. Rendering con volutional neural netw orks able also to learn ov er non-Euclidean domains is not that straigh tforward in that is re- quired a re-designing of the computational mo del for adaptiv ely learning graph em b eddings. Ov er flat domains, i.e. grid-lik e structures, conv olutional filters are compactly supp orted b ecause of the grid regularit y and the a v ailabilit y of con- sisten t node ordering across different samples. This makes it p ossible to learn 2 D. Bacciu and L. Di Sotto filters of fixed size and indep enden t of the input signal dimension leveraging, to this end, weigh t sharing tec hniques. F urthermore, a set of symmetric functions is also applied for sub-sampling purp oses to fully exploit the m ulti-scale nature of the grids. The same does not apply to domains with highly v arying topolo- gies where learnt filters (non-T o eplitz operators) ma y b e too representativ e of the considered domain, since they highly dep end on the eigen-basis of the filter op erator and they ma y thus fail to mo del sharp changes in the graph signal. State-of-the-art Graph Conv olutional Net works (GCNs) [11,17] try to ov ercome the ab o v e difficulties with conv olutions based on k -order Chebyshev p olynomials, in tro ducing the interesting dualit y of implicitly learning the graph sp ectrum by simply acting on the spatial representation. GCNs efficiently a void the compu- tational burden of p erforming a sp ectral decomp osition of the graph, yielding to learned filters that are indep enden t of the n umber of nodes in the graph. When considering graph classification tasks, we lac k a principled m ulti-resolution oper- ator pro viding coarser and more abstract represen tations of the input data as w e go deeper in the net w ork. Standard approac hes to graph p o oling emplo y sym- metric functions suc h as max, summation or av erage along features axes of the graph embeddings. In [29], it is giv en an account of the discriminative p ow er of these different coarse ning op erators. In the present work, we in tro duce a simple p ooling op erator for graphs that builds on the Non-Negativ e Matrix F actoriza- tion (NMF) metho ds to leverage on the communit y structure underlying graph structured data to induce subsampling, or equiv alently , a multiscale view of the input graph in order to capture long-range in teractions as we go deep er in Graph Con volutional Netw orks (GCNs). That would b e of practical in terest esp ecially in the context of graph classification or regression tasks where the whole graph is fed into do wnstream learning systems as a single signature vector. Suc h mech- anism is th us applied to incrementally obtain coarser graphs where nodes are p ooled in to comm unities based on the soft assignments output of the NMF of the graph adjacency matrix and Gram matrix of learned graph embeddings. Results on graph classification tasks sho w how jointly using suc h a coarsening operator with GCNs translate into improv ed predictive p erformances. 2 Bac kground In the follo wing w e in tro duce some basic notation used throughout the paper, then w e briefly introduce the necessary bac kground to understand state-of-the- art Graph Conv olutional Neural Netw orks (GCNs). W e mainly refer to sp ectral graph theory as introduced in [4,7,9]. 2.1 Basic notation A graph G is a tuple G = ( V , E ), where V is the set of vertices of the graph and E is the set of edges connecting v ertices, i.e. E ⊆ V × V . Let N ( i ) b e the set of neigh b ours of a no de i ∈ V . And let A ∈ I R n × n , with n = |V | , b e the adjacency A NMF approach to no de p ooling in GCNs 3 matrix such that A i,j = a i,j > 0 if ( i, j ) ∈ E 0 otherwise. Note that in the ab ov e form ulation we consider undirected graphs, i.e. suc h that ( i, j ) ∈ E and ( j, i ) ∈ E . Th us, matrix A is such that A = A T . In the presen t w ork, without loss of generality , w e generalize to undirected graphs W e also indicate with X ∈ I R n × d as the matrix of the n signals x i ∈ I R d asso ciated to each no de i ∈ V . 2.2 Graph Con v olution via polynomial filters Sp ectral construction. A first approach to representation learning on graphs is to explicitly learn the graph sp ectrum. In matrix notation, we can express the generalized conv olution ov er graphs as follo ws [7] LX = U ΛU T X (1) where L is the combinatorial graph Laplacian, L = D − A , with D the degree matrix such that D ii = P j a ij , where U ∈ I R n × k is an orthonormal basis gen- eralizing the F ourier basis, and where Λ is a diagonal matrix b eing the sp ectral represen tation of the filter [4,9]. Matrices U and Λ are the solution to the gen- eralized eigen v alue problem LU = U Λ [4,9]. With such an approac h there are m ultiple problems: (a) the eigendecomp osition in (1), and its application (fil- tering), require non-trivial computational time; (b) the corresp onding filters are non-lo calized [11]; (c) filter size is O ( n ), hence introducing a direct link b et ween the parameters and the n no des in the graph (no weigh t sharing). Spatial construction. In [11], it is prop osed an alternative approach to explicit learning of the graph sp ectrum, b y sho wing how it can b e learned implicitly through a p olynomial expansion of the diagonal op erator Λ . F ormally , g θ ( Λ ) = K − 1 X k =0 θ k Λ k (2) where θ ∈ I R K is the vector of polynomial co efficien ts. In [11] is p oin ted out that sp ectral filters represented as K -order p olynomials are exactly K-lo calized and that w eight sharing is th us made possible, since filters ha v e size O ( K ). Graph CNN (GCNN), also known as ChebNet [11], exploited the previous observ ation b y emplo ying Cheb yshev polynomials for approximating filtering operation (1). Cheb yshev p olynomials are recursively defined using the recurrence relation T j ( λ ) = 2 λT j − 1 ( λ ) − T j − 2 ( λ ); T 0 ( λ ) = 1; T 1 ( λ ) = λ. (3) 4 D. Bacciu and L. Di Sotto Also, p olynomials recursiv ely generated b y (3) form an orthonormal basis in [ − 1 , 1] [7,11]. A filter can th us be represented as a polynomial of the form g θ ( ˆ L ) = K − 1 X k =0 θ k U T k ( ˆ Λ ) U T = K − 1 X k =0 θ k T k ( ˆ L ) , (4) where ˆ L = 2 Λ/λ max − I n and ˆ Λ = 2 Λ/λ max − I n indicate a rescaling of the Laplacian eigenv alues to [ − 1 , 1]. The filtering op eration in (1) can b e rewritten, for one-dimensional input graph signals, as ˆ x = g θ ( ˆ L ) x ∈ I R n , where the k -th p olynomial ˆ x k = T k ( ˆ L ) x can b e computed using the recurrence relation in (3) no w defined as ˆ x = 2 ˆ Lx k − 1 − ˆ x k − 2 with ˆ x 0 = x and ˆ x 1 = ˆ Lx . More generally , taking in to account multi-dimensionalit y of input data, we hav e a conv olutional la yer as follows ˆ X = σ K − 1 X k =0 T k ( ∆ ) X Θ k ! (5) with σ a non-linear activ ation, and Θ ∈ I R d in × d out the matrix of learnable pa- rameters, with d in n umber of input features and d out n umber of neurons. A widely used con volutional la yer ov er graphs are GCNs by [17] that are lay ers of the form of (5) with K = 2, namely ˆ X = ReLU ˆ AX Θ . (6) The Θ term, the matrix of p olynomial co efficien ts to b e learned, stems from (5) b y imp osing Θ 0 = − Θ 1 , and with ˆ A = A + I , and non-linearity b eing the ReLU function [17]. Th us, the main idea is to generate a represen tation for a no de i ∈ V b y aggregating its own features x i ∈ I R d and its neigh b ors features x j ∈ I R d , where j ∈ N ( i ). Note that, apart from the formulation meant to highligh t the symmetry with con volutions on image data, the GCN mo del is not substan tially different from the con textual approach to graph pro cessing put forw ard b y [22] a decade b efore GCN, and recently extended to a probabilistic form ulation [3] by leveraging an hidden tree Marko v mo del [1] with relaxed causalit y assumptions and a fingerprinting approach to structure embedding [2]. 2.3 No de Pooling in Graph CNNs A first attempt to formalize graph p o oling can b e found in [9], a simple frame- w ork for multiresolution clustering of a graph is given based on a naive agglom- erativ e metho d. There are some recent works prop osing p ooling mechanisms for graph coarsening in Deep GCNs, in [10] a subset of the no des are dropp ed based on a learnable pro jection v ector where at eac h la yer only the top- k in- teresting nodes are retained. In [15], it is emplo yed a rough no de sampling and A NMF approach to no de p ooling in GCNs 5 a differentiable approac h through a LSTM model for learning aggregated node em b eddings, though it may render difficult satisfying inv ariance with resp ect to no de ordering. Interestingly , in [5] it is applied a simple and well known metho d from Graph Theory for no de decimation based on the largest eigenv ector u max of the graph Laplacian matrix. They further emplo y a more sophisticated pro- cedure to reduce Laplacian matrix using the sparsified Kron reduction. Another relev ant differen tiable approach is that put forward by DiffP o ol [31], where the mo del learns soft assignmen ts to p ool similar activ ating patterns into the same cluster, though the idea of learning hiearchical soft-clustering of graphs via ad- jacency matrix decomposition using a symmetric v ariant of NMF can b e dated bac k to [32]. In DiffP o ol, the learned soft assignment matrix is applied as a linear reduction operator on the adjacency matrix and the input signal matrix, and the coarsened graph is thus further con volv ed with GCNs. 3 NMFP o ol: no de p o oling by Non-Negative Matrix F actorization In the following section we introduce our model, NMFP o ol, a principled Pool- ing operator enabling deep graph CNNs develop multi-resolution representations of input graphs. NMFPool lev erages communit y structure underlying graphs to p ool similar nodes to progressiv ely gain coarser views of a graph. T o that end w e tak e inspiration from [32] in whic h latent communit y structure of graph data is made explicit via adjacency matrix decomp osition using Symmetric NMF (SNMF). NMFPool is grounded on that idea, building, instead, on a general non-symmetrical NMF of the adjacency matrix without constraining solutions to be sto c hastic. Before going further in to details of our approach, we first in- tro duce the formal definition of the NMF problem, then w e give an intuitiv e in terpretation of its solutions to clarify why NMF w ould help solve the graph p ooling problem on graphs. At the end we will show ho w to use pro duct factors of NMF as linear op erators to aggregate topology and con tent information as- so ciated to graphs. NMF is a p opular tec hnique for extracting salient features in data b y extracting a latent space representation of the original information. Throughout the pap er we refer to the original idea of NMF [19] though it has b een extensively studied in n umerical linear algebra in the last y ears b y man y authors and for a v ariety of applications. F ormally , the NMF problem can b e stated as follo ws: Definition 1. Given a non-ne gative matrix A ∈ I R n × m + , find non-ne gative ma- trix factors W ∈ I R n × k + and H ∈ I R k × m + , with k < min( m, n ) , such that A ≈ W H (7) If we see matrix A as having m m ultiv ariate ob jects column-stack ed, the straigh tforward interpretation of (7) is as follo ws a j ≈ W h j , (8) 6 D. Bacciu and L. Di Sotto with a j and h j corresp onding to j -th columns of A and H . The appro xima- tion (8) entails that each multi-v ariate ob ject is a linear com bination of columns of W w eighted b y co efficien ts in h j . Thus W is referred to as the basis ma- trix or equiv alently the cluster centroids matrix if w e in tend to in terpret NMF as a clustering metho d. Matrix H can be seen, instead, as a low-dimensional represen tation of the input data making th us NMF also useful for dimensional- it y reduction. Laten t representation, in the clustering p erspective, may indicate whether a sample ob ject b elongs to a cluster. F or example, w e could constrain eac h data-p oin t to b elong to a single cluster at a time: namely , eac h data-p oin t is assigned to the closest cluster x j ≈ u j . W e generally lo ok for non-trivial en- co dings to explain comm unity evolution in graphs. Th us, the problem could b e relaxed to a soft-clustering problem in that each data-p oin t can b elong to k o verlapping clusters [28]. F ormulation (7) requires to define a metric to measure the quality of the approximation, and Kullback-Leibler (KL-) divergence or the more common F rob enius norm (F-norm) are common choices. Man y techniques from n umerical linear algebra can b e used to minimize problem (7) whatev er the cost function we use, although its inheren tly non-conv ex nature does not giv e an y guarantee on global minimum [13]. In [19] w ere first prop osed multiplicativ e and additiv e update rules that ensure monotone descrease under KL- or F-norm. Th us, our prop osed solution can b e summarized into tw o main steps. First, w e enco de the input adjacency matrix to learn soft-assignments of no des, and that could accomplished via exact NMF of the adjacency matrix. Second, we apply soft-assignments as linear op erators to coarse adjacency matrix and no de em b eddings. T o this end, we refer to algebraic op erations seen in [32] for de- comp osing adjacency matrices and we extend it using equations widely used for graph coarsening [31], for they tak e into account em b edding matrix reduction and no des connectivit y strength. F or a complete picture, consider ` NMFP o ol la yers interlea v ed with at least ` + 1 stac ked Graph Conv olutions (GCs) as illus- trated in Figure 1, where the graph conv olutions are computed according to (6). Then, let Z ( i ) ∈ I R n i × d b e the output of i -th GC, namely the conv olv ed no de em b eddings at la yer i -th, defined as Z ( i ) = ReLU A ( i ) Z ( i − 1) Θ ( i ) (9) with adjacency matrix A ( i ) ∈ I R n i × n i , with n i n umber of no des at previous lay er, and Θ ( i ) ∈ I R d × d matrix of weigh ts. Observ e that we are assuming, without loss of generality , each GC lay er (9) as having the same n umber of neurons. Observe also that Z (0) = X ∈ I R n × d , namely the initial no de lab els, and the initial adjacency matrix is set to A (0) = ˆ D − 1 / 2 ˆ A ˆ D − 1 / 2 , i.e. the normalized adjacency matrix with ˆ A = A + I , A ∈ I R n × n , and ˆ D is a diagonal matrix of no de degrees [17]. The i -th NMFPool la yer solv es the problem in (7), i.e. the decomposition of the symmetric and p ositiv e A ( i ) , by minimizing the following loss || A ( i ) − W ( i ) H ( i ) || F (10) A NMF approach to no de p ooling in GCNs 7 GC (null) (null) (null) (null) fc (null) (null) (null) (null) P o ol (null) (null) (null) (null) GC (null) (null) (null) (null) GC (null) (null) (null) (null) P o ol (null) (null) (null) (null) Fig. 1: High level arc hitecture of a 3-la yers GCN in terleav ed with 2 NMF P o oling lay ers. with W ( i ) ∈ I R n i × k i + and H ( i ) ∈ I R k i × n i + , and k i n umber of o verlapping com- m unities to p o ol the n i no des in to, and k . k F the F rob enius norm. Observ e that k i ’s are hyper-parameters to con trol graph coarsening scale. The algorithm to minimize (10) depends on the underlying NMF implemen tation. Then NMFP o ol applies the enco ding H ( i ) to coarsen graph top ology and its conten t as follows Z ( i +1) = H ( i ) T Z ( i ) ∈ I R k i × d (11) A ( i +1) = H ( i ) T A ( i ) H ( i ) ∈ I R k i × k i . (12) A graphical interpretation of the inner workings of the NMFPool lay er is pro vided in Figure 2, highlighting the interpretation of po oling as a matrix de- comp osition op erator. It is crucial to p oin t out that NMFPool la yers are inde- p enden t of the n umber of nodes in the graph, whic h is essen tial to deal with graphs with v arying top ologies. 4 Exp erimen ts W e assess the effectiv eness of using the exact NMF of the adjacency matrix A as a po oling mechanism in graph con volutional neural netw orks. T o this end, we consider five popular graph classification b enchmarks and w e further compare the p erformance of our approac h, referred to as NMFPool in the following, with that of DiffPool, with the goal of showing ho w a simple and general metho d may easily compare to differen tiable and parameterized p ooling op erators such as DiffP o ol. Results were gathered on graph classification tasks for solving biological problems on the ENZYMES ([6], [25]), NCI1 [27], PR OTEINS ([6], [12]), and D&D ([12], [26]) datasets and the scientific collab oration dataset COLLAB [30]. In T able 1 are summarized statistics on b enc hmark datasets. In our exp erimen ts, the baseline graph conv olution is the v anilla lay er in (6). F or b oth mo dels, we employ ed the interlea ving of po oling and con volutional 8 D. Bacciu and L. Di Sotto 2 4 3 5 (null) (null) (null) (null) (null) (null) (null) (null) W (null) (null) (null) (null) H (null) (null) (null) (null) n (null) (null) (null) (null) k (null) (null) (null) (null) k (null) (null) (null) (null) n (null) (null) (null) (null) NMFP o ol Lay er (null) (null) (null) (null) p ooled graph: (null) (null) (null) (null) input graph: (null) (null) (null) (null) Fig. 2: The NMFPool la yer. Orange circles represen t no des of input graph, and solid lines the edges. Dashed lines are the predicted edges in b etw een no des p ooled together. Colored dashed circles represen t disco vered comm unities. T able 1: Statistics on b enchmark datasets. Dataset Graphs Classes Nodes (avg) Edges (a vg) COLLAB 5000 3 74.49 2457.78 D&D 1178 2 284.32 715.66 ENZYMES 600 6 32.63 62.14 NCI1 4110 2 29.87 32.30 PR OTEINS 1113 2 39.06 72.82 A NMF approach to no de p ooling in GCNs 9 la yers depicted in the architecture in Figure 1, v arying the n umber of p ooling- con volution lay er pairs to assess the effect of netw ork depth on task performance. Note that the n um b er of lay ers in the con volutional arc hitecture influences the con text spreading across the no des in the graph. Implemen tation of NMFP o ol and Diffp ool is based on the Pytorc h Geometric library [14], complemented b y the NMF implemen tation a v ailable in the Scikit library . Mo dels configurations w ere run on a multi-core arc hitecture equipped with 4 NUMA nodes eac h with 18 cores (Intel(R) Xeon(R) Gold 6140M @ 2.30GHz) capable of running 2 threads eac h for a total of 144 pro cessing units av ailable. W e had access also to 4 T esla V100 GPUs accelerators. Mo del selection w as p erformed for exploring a v ariet y of configurations using stratified 3-fold cross v alidation. F ollowing standard practice in graph conv olu- tion neural net works, learning rate w as set with an initial v alue of 0 . 1 and then decreased b y a factor of 0 . 1 whenev er v alidation error did not sho w any impro ve- men t after 10 ep o c hs w ait. The num b er of neurons is the same for each graph con volutional lay er and it has b een selected in { 16 , 32 , 64 , 128 } as part of the cross-v alidation pro cedure. When applying the p ooling op erator b oth NMFPool and Diffpo ol require to define the num ber of communities k , similarly to ho w the p ooling operator on images requires the definition of the p o oling windows size (and stride). Here, follo wing the idea indicated in the original DiffP o ol pap er [31], w e choose different k for each dataset as a fraction of the av erage n umber of no des in the samples. Thus during cross-v alidation we intended to study how NMFP o ol and DiffPool b eha ve as a function of the cluster sizes k i at each lay er. T o this end, p ooling size has been selected from the set { k 1 , k 2 } . In particular, for mo dels with a single p ooling lay er, we tested b oth sizes k 1 and k 2 . Instead, for deep er architectures, we restricted to use the largest k i for the first lay er, follo wing up in decreasing order of k i . T able 2 summarizes the num b er of clusters used for the first and second po oling lay er in the arc hitectures considered in this empirical assessment. T able 2: k 1 is computed using formula k 1 = b n av g · p c with p v arying in [21% − 25%], and n av g a verage num b er of no des (see T able 1). Then k 2 = k 1 / 2. F ractions are chosen dep ending on the size of task at hand and to previous empirical observ ation. Except for the D&D dataset where p = 5% , 1%, b eing the bigger dataset we needed a go od compromise b etw een abstraction capabilit y and computational time. Dataset k 1 k 2 p COLLAB 16 8 22% D&D 14 2 5% - 1% ENZYMES 8 4 25% NCI1 6 3 24% PR OTEINS 8 4 21% 10 D. Bacciu and L. Di Sotto The outcome of the empirical ass essmen t is summarized in T able 3, where it is rep orted the mean classification accuracy of the differen t mo dels av eraged on the dataset folds. T able 3 rep orts results for a v anilla GCN (no p o oling) and a v arying num ber of graph conv olution la yers: results show how at most t wo lay ers are sufficient to guarantee go o d p erformances, while three la yers are only required for the COLLAB dataset and a single la y er net work obtains the b est p erformance on the NCI1 dataset. In the exp erimen ts w e th us decided to emplo y at most three GCN lay ers, namely at most tw o NMF and DiffPool p ool- ing lay ers. It is still eviden t how adding more conv olutional and po oling lay ers do es not alwa ys result into better p erformances. The analysis of the results for NMFP o ol sho ws ho w the addition of the simple NMF po oling allo ws a consisten t increase of the classification accuracy with resp ect to the non-p ooled mo del for all the benchmark datasets. Note how a single p ooling lay er is sufficien t, on most datasets, to obtain the b est results, confirming the fact that p o oling allo ws to effectiv ely fasten the process of context spreading betw een the nodes. When com- pared to DiffPool, our approach achiev es accuracies whic h are only marginally lo wer than DiffPool on few datasets. This despite the fact that DiffP o ol em- plo ys a solution p erforming an task-sp ecific parameterized decomp osition of the graph, while our solution simply looks for quasi-symmetrical product matrices b y kno wing nothing of the underlying task. T able 3: Mean and standard deviation (in brack ets) of graph classification accuracies on the different b enc hmarks, for the v anilla GCN with ` con volutional lay ers ( ` -GC), for NMFPool and DiffPool with ` p p ooling la yers and ` p + 1 con volutional lay ers (i.e. ` p 1 -NMFP o ol and ` p 2 -DiffP o ol, resp ectiv ely). Mo del ENZYMES NCI1 PR OTEINS D&D COLLAB 1-GC 0.222 (0 . 023) 0.625 (0 . 014) 0.713 (0 . 019) 0.681 (0 . 045) 0.671 (0 . 007) 2-GCs 0.228 (0 . 023) 0.620 (0 . 057) 0.720 (0 . 034) 0.704 (0 . 048) 0.678 (0 . 007) 3-GCs 0.182 (0 . 022) 0.628 (0 . 031) 0.688 (0 . 024) 0.692 (0 . 032) 0.681 (0 . 002) 1-NMFP o ol 0.241 (0 . 039) 0.662 (0 . 026) 0.721 (0 . 031) 0.760 (0 . 015) 0.650 (0 . 004) 2-NMFP o ol 0.175 (0 . 023) 0.655 (0 . 013) 0.724 (0 . 020) 0.753 (0 . 010) 0.658 (0 . 002) 1-DiffP o ol 0.259 (0 . 069) 0.661 (0 . 017) 0.743 (0 . 011) 0.770 (0 . 007) 0.659 (0 . 005) 2-DiffP o ol 0.239 (0 . 064) 0.632 (0 . 017) 0.744 (0 . 026) 0.761 (0 . 003) 0.667 (0 . 022) 5 Conclusions W e in tro duced a p o oling mechanism based on the NMF of the adjacency matrix of the graph, discussing how this approach can b e used to yield a hierarchical soft-clustering of the nodes and to induce a coarsening of the graph structure. A NMF approach to no de p ooling in GCNs 11 W e ha ve empirically assessed our NMPool approac h with the task-sp ecific adap- tiv e p ooling mechanism put forward by the DiffPool mo del on a num b er of state-of-the-art graph classification b enchmarks. W e argue that our approach can yield to p oten tially more general and scalable p o oling mec hanisms than DiffP o ol, allowing to choose weather the p ooling mechanism has to consider the no de embeddings computed b y the mo del and the task-related information when p erforming the decomp osition (as in DiffPool), but also allowing to directly de- comp ose the graph structure a-priori with no knowledge of the node embeddings adaptiv ely computed by the con volutional lay er. This latter asp ect, in particular, allo ws to pre-compute the graph decomp osition and results in a multiresolution represen tation of the graph structure whic h do es not change with the particular task at hand. F uture works will consider the use of symmetric and optimized NMF v ari- an ts to increase prediction p erformances. It also would b e of particular in terest to improv e the quality and quan tity of information NMFPool retains into the enco ding matrix. NMFPool could evolv e out of its general purp ose form, for example, making it a generative end-to-end differentiable la yer using probabilis- tic approaches. See [8] for an attempt to solve NMF using probabilistic mo dels. W e could refer to the p opular probabilistic generative mo del of the V ariational Auto-Enco ders (V AEs) [16], [24] p ossibly extended to graphs [18]. The under- lying hierarc hical structure of graph data may also be taken in to account b y imp osing laten t enco ding to match priors referring to h yp erb olic spaces [20]. In terestingly , laten t matrix enco ding ma y not b e forced to matc h o verimposed priors, for they could make the mo del to o biased ov er particular graph geome- tries. Instead, such priors could be directly learned from relational data using adv ersarial approaches [21] extended also to graph auto-enco ders [23]. Another in teresting feature w ould be to make NMFPool indep enden t of hyper-parameter k . Ac knowledgmen ts This work has been supported b y the Italian Ministry of Education, Universit y , and Research (MIUR) under pro ject SIR 2014 LIST-IT (grant n. RBSI14STDE). References 1. Bacciu, D., Micheli, A., Sp erduti, A.: Comp ositional generative mapping for tree- structured data - part II: T op ographic pro jection model. IEEE T rans. Neural Netw. Learning Syst. 24 (2), 231–247 (2013) 2. Bacciu, D., Micheli, A., Sp erduti, A.: Generative k ernels for tree-structured data. IEEE T ransactions on Neural Netw orks and Learning Systems 29 (10), 4932–4946 (Oct 2018) 3. Bacciu, D., Errica, F., Micheli, A.: Contextual graph Mark ov mo del: A deep and generativ e approach to graph pro cessing. In: Dy , J., Krause, A. (eds.) Pro ceedings of the 35th In ternational Conference on Mac hine Learning. Pro ceedings of Machine Learning Research, vol. 80, pp. 294–303. PMLR, Sto ckholmsmssan, Stockholm Sw eden (2018) 12 D. Bacciu and L. Di Sotto 4. Belkin, M., Niy ogi, P .: Laplacian eigenmaps and spectral tec hniques for em b edding and clustering. In: Proceedings of the 14th In ternational Conference on Neural Information Processing Systems: Natural and Syn thetic. pp. 585–591. NIPS’01, MIT Press, Cambridge, MA, USA (2001) 5. Bianc hi, F.M., Grattarola, D., Livi, L., Alippi, C.: Graph neural netw orks with con volutional ARMA filters. CoRR abs/1901.01343 (2019) 6. Borgw ardt, K.M., Ong, C.S., Schonauer, S., Vishw anathan, S.V.N., Smola, A.J., Kriegel, H.P .: Protein function prediction via graph kernels. Bioinformatics 21 (Suppl 1), i47–i56 (2005) 7. Bronstein, M.M., Bruna, J., LeCun, Y., Szlam, A., V andergheynst, P .: Geometric deep learning: going b ey ond euclidean data. CoRR abs/1611.08097 (2016) 8. Brou wer, T., F rellsen, J., Li` o, P .: F ast ba yesian non-negativ e matrix factorisation and tri-factorisation (12 2016), nIPS 2016 : Adv ances in Appro ximate Bay esian Inference W orkshop ; Conference date: 09-12-2016 Through 09-12-2016 9. Bruna, J., Zaremba, W., Szlam, A., Lecun, Y.: Sp ectral netw orks and lo cally con- nected net works on graphs. In: International Conference on Learning Represen ta- tions (ICLR2014), CBLS, April 2014 (2014) 10. Cangea, C., V eliˇ ck ovi ´ c, P ., Jov anovi ´ c, N., Kipf, T., Li` o, P .: T ow ards Sparse Hier- arc hical Graph Classifiers. arXiv e-prints arXiv:1811.01287 (Nov 2018) 11. Defferrard, M., Bresson, X., V andergheynst, P .: Conv olutional neural netw orks on graphs with fast lo calized sp ectral filtering. CoRR abs/1606.09375 (2016) 12. Dobson, P .D., Doig, A.J.: Distinguishing enzyme structures from non-enzymes without alignments. Journal of Molecular Biology 330 (4), 771 – 783 (2003) 13. F a v ati, P ., Lotti, G., Menchi, O., Romani, F.: Adaptiv e computation of the Sym- metric Nonnegative Matrix F actorization (NMF). arXiv e-prints (Mar 2019) 14. F ey , M., Lenssen, J.E.: F ast graph representation learning with pytorc h geometric. CoRR abs/1903.02428 (2019) 15. Hamilton, W.L., Ying, R., Lesko vec, J.: Inductive representation learning on large graphs. CoRR abs/1706.02216 (2017) 16. Kingma, D.P ., W elling, M.: Auto-Enco ding V ariational Bay es. arXiv e-prints arXiv:1312.6114 (Dec 2013) 17. Kipf, T.N., W elling, M.: Semi-sup ervised classification with graph con volutional net works. CoRR abs/1609.02907 (2016) 18. Kipf, T.N., W elling, M.: V ariational Graph Auto-Enco ders. arXiv e-prints arXiv:1611.07308 (Nov 2016) 19. Lee, D.D., Seung, H.S.: Algorithms for non-negative matrix factorization. In: Leen, T.K., Dietteric h, T.G., T resp, V. (eds.) Adv ances in Neural Information Pro cessing Systems 13, pp. 556–562. MIT Press (2001) 20. Mathieu, E., Le Lan, C., Maddison, C.J., T omiok a, R., Wh ye T eh, Y.: Hierar- c hical Representations with Poincar \ ’e V ariational Auto-Enco ders. arXiv e-prin ts arXiv:1901.06033 (Jan 2019) 21. Mesc heder, L.M., Now ozin, S., Geiger, A.: Adversarial v ariational bay es: Uni- fying v ariational auto enco ders and generative adversarial netw orks. CoRR abs/1701.04722 (2017) 22. Mic heli, A.: Neural netw ork for graphs: A contextual constructiv e approach. IEEE T ransactions on Neural Netw orks 20 (3), 498–511 (March 2009) 23. P an, S., Hu, R., Long, G., Jiang, J., Y ao, L., Zhang, C.: Adv ersarially regularized graph auto enco der. CoRR abs/1802.04407 (2018) A NMF approach to no de p ooling in GCNs 13 24. Rezende, D.J., Mohamed, S., Wierstra, D.: Stochastic bac kpropagation and ap- pro ximate inference in deep generative mo dels. In: Xing, E.P ., Jebara, T. (eds.) Pro ceedings of the 31st In ternational Conference on Mac hine Learning. Pro ceed- ings of Machine Learning Research, vol. 32, pp. 1278–1286. PMLR, Bejing, China (22–24 Jun 2014) 25. Sc homburg, I., Chang, A., Eb eling, C., Gremse, M., Heldt, C., Huhn, G., Schom- burg, D.: Brenda, the enzyme database: Up dates and ma jor new dev elopments. Nucleic acids research 32 , D431–3 (01 2004). https://doi.org/10.1093/nar/gkh081 26. Sherv ashidze, N., Sc h weitzer, P ., v an Leeu wen, E.J., Mehlhorn, K., Borgwardt, K.M.: W eisfeiler-lehman graph k ernels. J. Mac h. Learn. Res. 12 , 2539–2561 (No v 2011) 27. W ale, N., W atson, I.A., Karypis, G.: Comparison of descriptor spaces for chemical comp ound retriev al and classification. Kno wl. Inf. Syst. 14 (3), 347–375 (Mar 2008). h ttps://doi.org/10.1007/s10115-007-0103-5 28. W att, J., Borhani, R., Katsaggelos, A.K.: Machine Learning Refined: F oundations, Algorithms, and Applications. Cambridge Universit y Press, New Y ork, NY, USA, 1st edn. (2016) 29. Xu, K., Hu, W., Lesko vec, J., Jegelk a, S.: Ho w pow erful are graph neural netw orks? CoRR abs/1810.00826 (2018) 30. Y anardag, P ., Vishw anathan, S.: Deep graph kernels. In: Pro ceedings of the 21th ACM SIGKDD In ternational Conference on Knowledge Discov ery and Data Mining. pp. 1365–1374. KDD ’15, ACM, New Y ork, NY, USA (2015). h ttps://doi.org/10.1145/2783258.2783417 31. Ying, Z., Y ou, J., Morris, C., Ren, X., Hamilton, W., Lesko vec, J.: Hierarchical graph represen tation learning with differentiable p o oling. In: Bengio, S., W allach, H., Laro chelle, H., Grauman, K., Cesa-Bianc hi, N., Garnett, R. (eds.) Adv ances in Neural Information Processing Systems 31, pp. 4804–4814. Curran Asso ciates, Inc. (2018) 32. Y u, K., Y u, S., T resp, V.: Soft clustering on graphs. In: W eiss, Y., Sc h¨ olk opf, B., Platt, J.C. (eds.) Adv ances in Neural Information Pro cessing Systems 18, pp. 1553–1560. MIT Press (2006), http://papers.nips.cc/paper/ 2948- soft- clustering- on- graphs.pdf
Original Paper
Loading high-quality paper...
Comments & Academic Discussion
Loading comments...
Leave a Comment