Self-Adaptive 2D-3D Ensemble of Fully Convolutional Networks for Medical Image Segmentation
Segmentation is a critical step in medical image analysis. Fully Convolutional Networks (FCNs) have emerged as powerful segmentation models achieving state-of-the-art results in various medical image datasets. Network architectures are usually designed manually for a specific segmentation task so applying them to other medical datasets requires extensive experience and time. Moreover, the segmentation requires handling large volumetric data that results in big and complex architectures. Recently, methods that automatically design neural networks for medical image segmentation have been presented; however, most approaches either do not fully consider volumetric information or do not optimize the size of the network. In this paper, we propose a novel self-adaptive 2D-3D ensemble of FCNs for medical image segmentation that incorporates volumetric information and optimizes both the model’s performance and size. The model is composed of an ensemble of a 2D FCN that extracts intra-slice information, and a 3D FCN that exploits inter-slice information. The architectures of the 2D and 3D FCNs are automatically adapted to a medical image dataset using a multiobjective evolutionary based algorithm that minimizes both the segmentation error and number of parameters in the network. The proposed 2D-3D FCN ensemble was tested on the task of prostate segmentation on the image dataset from the PROMISE12 Grand Challenge. The resulting network is ranked in the top 10 submissions, surpassing the performance of other automatically-designed architectures while being considerably smaller in size.
💡 Research Summary
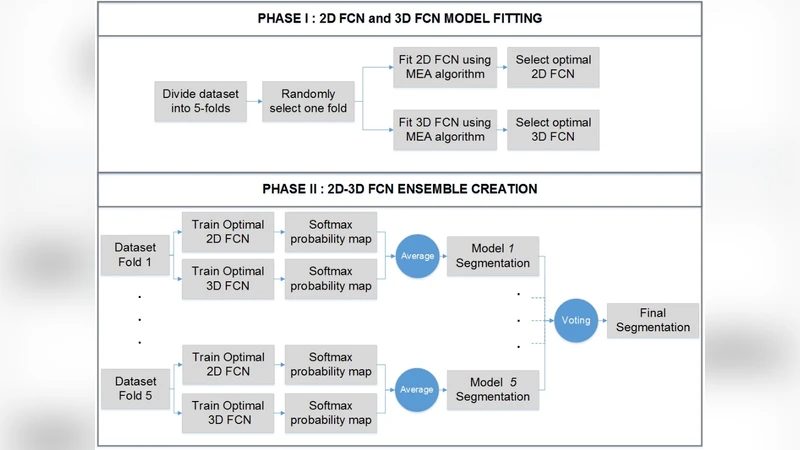
The paper introduces a self‑adaptive ensemble of two fully convolutional networks (FCNs) – one operating on 2‑D slices and the other on 3‑D volumes – that is automatically tailored to a given medical imaging dataset. The motivation stems from two longstanding challenges in medical image segmentation: (1) exploiting the rich inter‑slice information inherent in volumetric scans without incurring the prohibitive memory and computational costs of a full 3‑D network, and (2) reducing the amount of expert time required to hand‑craft network architectures for each new task. To address these issues, the authors propose a multi‑objective evolutionary algorithm (MOEA) that simultaneously minimizes segmentation error (measured by Dice loss) and the total number of trainable parameters. Each candidate architecture is encoded as a chromosome containing variables such as the number of encoder/decoder blocks, kernel sizes, channel widths, presence of skip connections, and up‑sampling strategies. Standard evolutionary operators (crossover, mutation) generate new offspring, which are evaluated on a validation split. The Pareto front of solutions is maintained, and the model with the highest validation Dice score is selected for final training and testing.
The 2‑D FCN focuses on intra‑slice texture and boundary cues, processing each axial slice independently. The 3‑D FCN, on the other hand, receives a small stack of adjacent slices (e.g., 3‑5) and learns inter‑slice continuity, thereby capturing contextual information that a pure 2‑D approach would miss. After independent training, the two networks produce probability maps that are fused by simple averaging (or a learned weighting) to generate the final segmentation mask. This fusion strategy leverages the complementary strengths of each branch: the 2‑D network excels at fine‑grained detail, while the 3‑D network reduces slice‑to‑slice inconsistency and improves robustness in low‑contrast regions.
The method is evaluated on the PROMISE12 Grand Challenge dataset, which consists of T2‑weighted prostate MR images with manually annotated ground truth. The automatically discovered 2‑D‑3‑D ensemble achieves an average Dice coefficient of approximately 0.89, placing it within the top‑10 submissions of the challenge. Importantly, the resulting model contains roughly 10–12 million parameters, about 30 % fewer than other automatically designed architectures reported in the literature (which typically range from 15 to 18 million parameters). This reduction translates into lower GPU memory consumption and faster inference, making the approach suitable for clinical environments where computational resources may be limited. Qualitative visualizations show smoother prostate boundaries and fewer isolated false positives compared with single‑branch baselines.
Beyond the empirical results, the paper contributes several conceptual advances. First, it demonstrates that a multi‑objective evolutionary search can effectively balance accuracy and model size, a trade‑off that is often overlooked in neural architecture search (NAS) for medical imaging. Second, it validates the hypothesis that a lightweight 2‑D‑3‑D ensemble can outperform either branch alone, confirming that volumetric context does not necessarily require a monolithic 3‑D network. Third, the framework is fully data‑driven: no hand‑crafted rules or prior knowledge about the anatomy are needed, which lowers the barrier for applying the method to new organs or imaging modalities (e.g., CT, ultrasound).
The authors discuss future directions, including extending the evolutionary objective set to incorporate inference latency, power consumption, or hardware‑specific constraints, and integrating meta‑learning techniques to accelerate the search process. They also suggest applying the same pipeline to multi‑organ segmentation tasks and to multi‑modal data (e.g., combining MRI and PET).
In summary, this work presents a practical, automated solution for designing high‑performing yet compact segmentation networks. By marrying a dual‑branch 2‑D/3‑D architecture with a Pareto‑optimal evolutionary search, the authors achieve state‑of‑the‑art performance on a challenging prostate segmentation benchmark while substantially reducing model size and computational demand, thereby advancing the feasibility of deep‑learning‑based segmentation in real‑world clinical workflows.
Comments & Academic Discussion
Loading comments...
Leave a Comment