Bayesian Volumetric Autoregressive generative models for better semisupervised learning
Deep generative models are rapidly gaining traction in medical imaging. Nonetheless, most generative architectures struggle to capture the underlying probability distributions of volumetric data, exhibit convergence problems, and offer no robust indices of model uncertainty. By comparison, the autoregressive generative model PixelCNN can be extended to volumetric data with relative ease, it readily attempts to learn the true underlying probability distribution and it still admits a Bayesian reformulation that provides a principled framework for reasoning about model uncertainty. Our contributions in this paper are two fold: first, we extend PixelCNN to work with volumetric brain magnetic resonance imaging data. Second, we show that reformulating this model to approximate a deep Gaussian process yields a measure of uncertainty that improves the performance of semi-supervised learning, in particular classification performance in settings where the proportion of labelled data is low. We quantify this improvement across classification, regression, and semantic segmentation tasks, training and testing on clinical magnetic resonance brain imaging data comprising T1-weighted and diffusion-weighted sequences.
💡 Research Summary
The paper introduces 3DPixelCNN, the first volumetric autoregressive generative model that extends the 2‑D PixelCNN architecture to three‑dimensional brain MRI data. To avoid the “blind‑spot” problem inherent in masked convolutions, the authors design three complementary stacks—horizontal, depth, and vertical—each conditioned on appropriate subsets of previously generated voxels. These stacks are combined using gated activation units (tanh × sigmoid) and residual connections, enabling deep networks to be trained efficiently while preserving the strict autoregressive ordering required for exact likelihood estimation.
A key contribution is the Bayesian reformulation of the model. By inserting SpatialDropout after every convolutional layer and performing multiple stochastic forward passes (T = 20) at test time, the network yields voxel‑wise mean (µ) and standard deviation (σ) estimates. This procedure approximates a deep Gaussian process, providing a principled measure of epistemic uncertainty without altering the underlying generative objective. The model is trained with continuous negative log‑likelihood using the Adam optimizer (learning rate = 0.001, batch size = 1) for 20 epochs.
Experiments are conducted on two large neuro‑imaging cohorts. The first consists of 2,315 diffusion‑weighted imaging (DWI) scans from acute stroke patients, with manually curated lesion masks. The second comprises 13,287 T1‑weighted grey‑matter segmentations from the UK Biobank and UCLH, together with age and sex labels. All volumes are down‑sampled to 3 mm isotropic resolution (52 × 64 × 52 voxels) to keep training feasible.
Reconstruction quality, measured by log‑likelihood, shows that 3DPixelCNN outperforms a Bayesian auto‑encoder baseline (0.360 vs 0.378 on DWI and 0.105 vs 0.222 on grey‑matter), especially on the higher‑detail T1 data where it achieves a 111 % relative improvement. Uncertainty maps (σ) derived from the model trained on non‑lesioned brains highlight regions that deviate from the learned “healthy” distribution; a simple threshold on σ improves Dice overlap for stroke lesion detection from 14.7 % (using raw intensities) to 23.7 %, surpassing the auto‑encoder’s 17.3 % result.
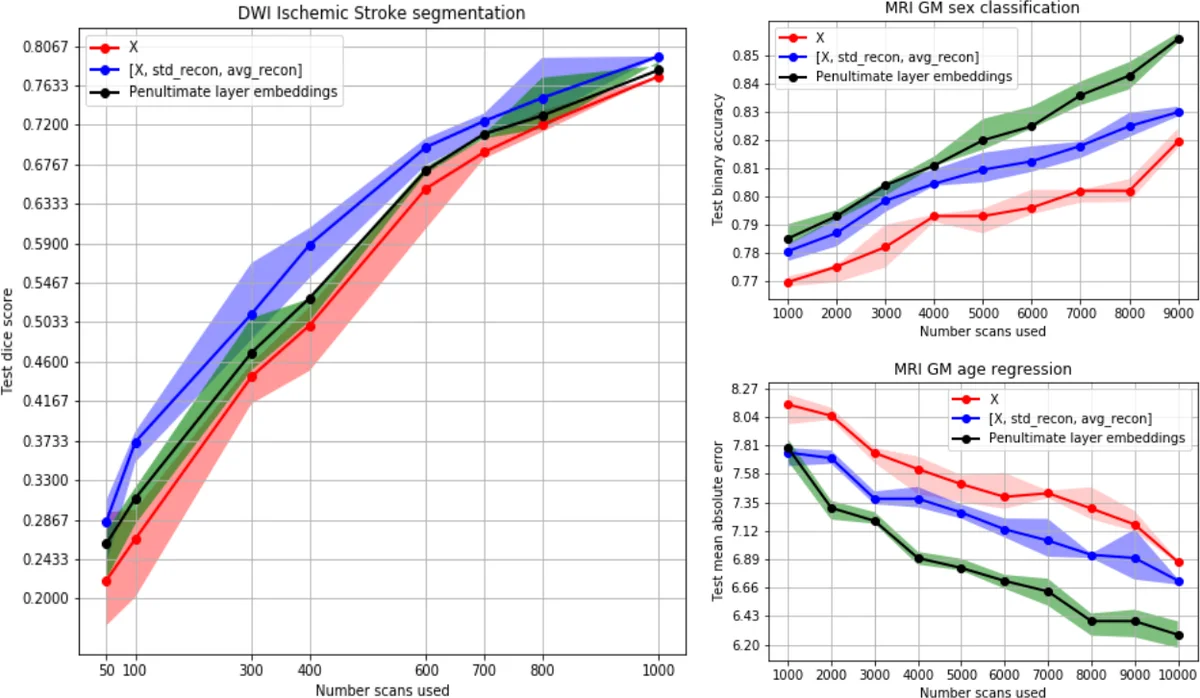
For semi‑supervised learning, the authors augment two downstream classifiers: a 3D U‑Net for lesion segmentation and an ASC network (a shallow CNN) for age regression and sex classification. Three input variants are evaluated: (χ) raw volumes only, (ξ) raw volumes concatenated with µ and σ, and (ψ) the activations of the final 3DPixelCNN layer (a high‑dimensional embedding). Across all tasks, adding uncertainty information (ξ) consistently boosts performance when labeled data are scarce. In lesion segmentation, ξ yields a 0.082 Dice increase (25.6 % relative gain) for N < 500 training samples and an average gain of 0.056 (15.2 %). The high‑dimensional embedding ψ provides smaller but still positive gains (average Dice +0.025). For age regression, ψ reduces mean absolute error by 0.68 years (9.09 %); ξ reduces error by 0.30 years (3.98 %). For sex classification, ψ improves accuracy by 3.36 % versus 1.87 % for ξ. By contrast, the Bayesian auto‑encoder’s uncertainty (ξ) degrades performance on all tasks, likely due to higher noise in its uncertainty estimates.
These findings suggest that voxel‑wise uncertainty is most beneficial for tasks with localized signals (e.g., lesion segmentation), whereas the richer latent representation of the autoregressive model better supports global predictions (age, sex). The work demonstrates that a volumetric autoregressive model can simultaneously learn a full probability distribution over high‑dimensional medical images and provide useful Bayesian uncertainty estimates, thereby enhancing semi‑supervised learning when annotated data are limited. The authors note that future work could explore full‑resolution training, alternative Bayesian approximations, and integration with other downstream architectures.
Comments & Academic Discussion
Loading comments...
Leave a Comment