RinQ Fingerprinting: Recurrence-informed Quantile Networks for Magnetic Resonance Fingerprinting
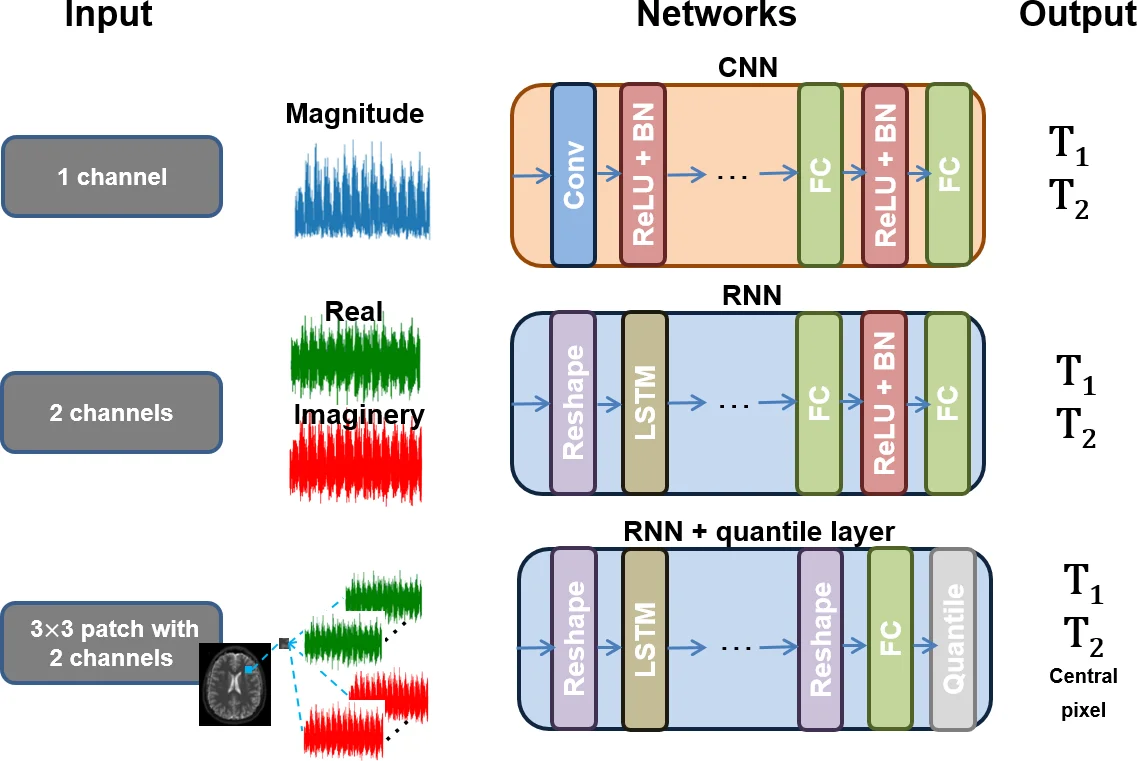
Recently, Magnetic Resonance Fingerprinting (MRF) was proposed as a quantitative imaging technique for the simultaneous acquisition of tissue parameters such as relaxation times $T_1$ and $T_2$. Although the acquisition is highly accelerated, the state-of-the-art reconstruction suffers from long computation times: Template matching methods are used to find the most similar signal to the measured one by comparing it to pre-simulated signals of possible parameter combinations in a discretized dictionary. Deep learning approaches can overcome this limitation, by providing the direct mapping from the measured signal to the underlying parameters by one forward pass through a network. In this work, we propose a Recurrent Neural Network (RNN) architecture in combination with a novel quantile layer. RNNs are well suited for the processing of time-dependent signals and the quantile layer helps to overcome the noisy outliers by considering the spatial neighbors of the signal. We evaluate our approach using in-vivo data from multiple brain slices and several volunteers, running various experiments. We show that the RNN approach with small patches of complex-valued input signals in combination with a quantile layer outperforms other architectures, e.g. previously proposed CNNs for the MRF reconstruction reducing the error in $T_1$ and $T_2$ by more than 80%.
💡 Research Summary
Magnetic Resonance Fingerprinting (MRF) promises rapid, quantitative mapping of tissue relaxation parameters (T1, T2) by acquiring highly undersampled, time‑varying signal evolutions. Conventional reconstruction relies on template matching against a discretized dictionary of simulated signals, which is computationally intensive and limited by the finite resolution of the dictionary. Recent deep‑learning (DL) approaches replace the exhaustive search with a regression network that maps measured signals directly to parameter values, but most existing DL solutions either use fully‑connected networks (prone to over‑fitting) or convolutional neural networks (CNNs) that are not optimal for processing long temporal sequences.
In this work, the authors propose a novel architecture called RinQ Fingerprinting, which combines a recurrent neural network (RNN) based on Long Short‑Term Memory (LSTM) cells with a newly introduced quantile layer. The key design choices are:
-
Complex‑valued input – Instead of discarding phase information and feeding only magnitude data, the full complex signal (real and imaginary parts) is used. Each MRF time series contains 3000 points; the authors reshape it into 30 blocks of 100 complex samples (200 real‑valued features) to keep the sequence length manageable for the LSTM and to mitigate vanishing/exploding gradients.
-
Temporal modeling with LSTM – The LSTM processes the reshaped sequence, capturing the temporal dependencies that are intrinsic to MRF signals more effectively than a CNN can.
-
Spatial context via patches – For each voxel, a 3 × 3 neighborhood of signals is extracted, yielding nine temporally‑aligned sequences. This assumes that neighboring voxels belong to the same tissue class and therefore share similar T1/T2 values.
-
Quantile layer – After the RNN produces nine independent predictions (one per patch element), a 0.5 quantile (median) operation aggregates them. The quantile is implemented as a sparse matrix Q that selects the median element; during back‑propagation the gradient is simply Qᵀ, making the layer differentiable. This median pooling is more robust to outliers caused by undersampling artefacts than average or max pooling, and it preserves edges.
The network is trained with a mean‑squared‑error loss using the ADAM optimizer. Ground‑truth parameter maps are generated by a very fine dictionary (≈ 691 k entries) after compressing both the dictionary and measured signals to 50 principal components via SVD. Data were acquired from eight volunteers (12 axial brain slices) on a 3 T scanner using a fast imaging with steady‑state precession (FISP) sequence with a spiral readout and an undersampling factor of 48. An additional set of 16 slices from four more volunteers was used to test the effect of larger training data.
Four experimental configurations were evaluated:
- Magnitude vs. complex input – Switching from magnitude (Sm) to complex (Sc) reduced errors for both CNN and RNN (CNN error ↓ 62 %, RNN error ↓ 50 %).
- CNN vs. RNN – With comparable numbers of trainable parameters, the RNN outperformed the CNN, achieving up to a 53 % reduction in relative mean error (RME). The CNN failed to produce meaningful tissue contrast, especially when using magnitude inputs.
- Patch size and quantile layer – Adding a 3 × 3 spatial patch and the median quantile layer (RNN 3) further lowered T1 error by 57 % and T2 error by 43 % relative to an RNN without the quantile operation. Visual inspection showed that the quantile layer dramatically reduced artefacts at tissue boundaries, acting as an edge‑preserving denoiser.
- Training data size – Training the best model (RNN 3) on the larger dataset (RNN * 3) yielded the lowest validation loss (≈ 195 ms √MSE), confirming that the architecture benefits from more data.
Overall, the proposed RinQ architecture achieved a reduction of more than 80 % in T1/T2 estimation error compared with the baseline CNN, while handling severely undersampled, noisy in‑vivo data that are far more challenging than synthetic or fully sampled dictionary signals used in prior work.
Limitations and future directions: The study is constrained by a relatively small number of subjects and slices; cross‑subject generalization remains to be rigorously tested. The reliance on a finely sampled dictionary for ground truth inflates the required training data volume. The authors suggest incorporating physics‑based layers (e.g., Bloch simulation modules) to reduce model complexity and improve interpretability, as well as expanding the dataset to include completely unseen volunteers for a more robust validation.
In summary, RinQ Fingerprinting demonstrates that a recurrent network equipped with complex‑valued inputs, spatial patch context, and a differentiable median‑quantile pooling layer can dramatically accelerate MRF reconstruction while delivering highly accurate T1 and T2 maps, even under aggressive undersampling conditions. This work paves the way for real‑time quantitative MRI in clinical practice.
Comments & Academic Discussion
Loading comments...
Leave a Comment