Exploiting bilateral symmetry in brain lesion segmentation
Brain lesions, including stroke and tumours, have a high degree of variability in terms of location, size, intensity and form, making automatic segmentation difficult. We propose an improvement to existing segmentation methods by exploiting the bilateral quasi-symmetry of healthy brains, which breaks down when lesions are present. Specifically, we use nonlinear registration of a neuroimage to a reflected version of itself (“reflective registration”) to determine for each voxel its homologous (corresponding) voxel in the other hemisphere. A patch around the homologous voxel is added as a set of new features to the segmentation algorithm. To evaluate this method, we implemented two different CNN-based multimodal MRI stroke lesion segmentation algorithms, and then augmented them by adding extra symmetry features using the reflective registration method described above. For each architecture, we compared the performance with and without symmetry augmentation, on the SISS Training dataset of the Ischemic Stroke Lesion Segmentation Challenge (ISLES) 2015 challenge. Using affine reflective registration improves performance over baseline, but nonlinear reflective registration gives significantly better results: an improvement in Dice coefficient of 13 percentage points over baseline for one architecture and 9 points for the other. We argue for the broad applicability of adding symmetric features to existing segmentation algorithms, specifically using nonlinear, template-free methods.
💡 Research Summary
The paper introduces a novel way to improve automatic brain lesion segmentation by explicitly exploiting the bilateral quasi‑symmetry of the healthy brain. The authors observe that, in a normal brain, each voxel has a homologous counterpart in the opposite hemisphere with very similar intensity values, whereas lesions (stroke or tumor) typically break this symmetry, producing large intensity differences between a lesion voxel and its mirror voxel. To capture this information, they propose a “reflective registration” pipeline: each subject’s multimodal MRI (FLAIR, DWI, T1, T1‑contrast) is first reflected across the left‑right axis, then registered back to the original image. Two registration strategies are examined: (i) a linear rigid + affine registration (LSymm) and (ii) a full nonlinear SyN registration (NLSymm) using the ANTs SynQuick tool. The resulting transformation T(x,y,z) maps every voxel to its mirrored location. For each modality a Symmetry Difference Image (SDI) is computed as the voxel intensity minus the intensity of its mirrored voxel: S_r = I_r – I_r∘T. In healthy tissue these SDIs are near zero; in lesion tissue they show large absolute values.
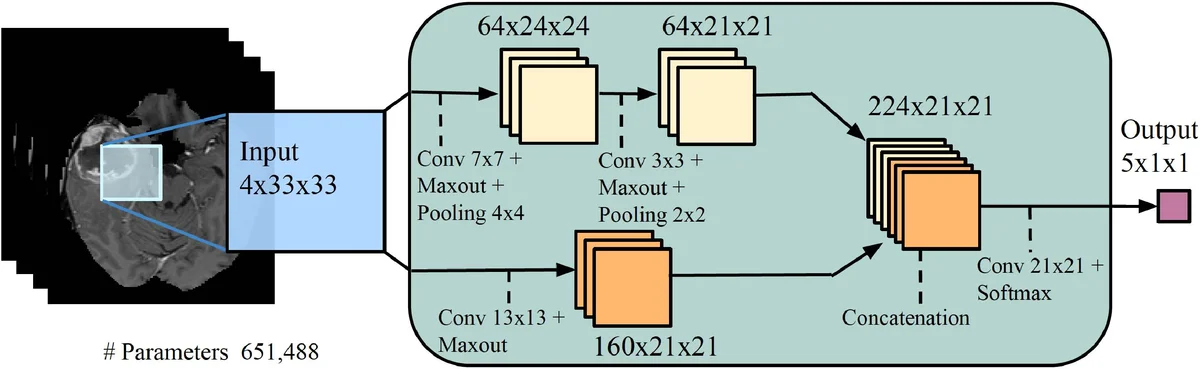
These SDIs are concatenated with the original four modalities, yielding an eight‑channel input that is fed unchanged into two existing 2‑D convolutional neural network (CNN) architectures: TwoPathCNN (a shallow network with a local and a global path) and Wider2dSeg (a deeper, multi‑scale network derived from the 3‑D DeepMedic architecture). The authors keep all other training details identical to the baseline models, including loss functions, regularization, data augmentation, and the 7‑fold cross‑validation protocol on the ISLES‑2015 SISS training set (28 patients, isotropic 1 mm³ resolution).
Results show that adding symmetry features consistently improves segmentation performance across all metrics (Dice coefficient, recall, precision). For TwoPathCNN, Dice improves from 0.45 ± 0.25 (baseline) to 0.52 ± 0.23 with linear symmetry and to 0.54 ± 0.21 with nonlinear symmetry. For the deeper Wider2dSeg, the gains are even larger: Dice rises from 0.49 ± 0.25 (baseline) to 0.61 ± 0.22 with linear symmetry and to 0.62 ± 0.22 with nonlinear symmetry—a 13‑percentage‑point increase over baseline. Recall and precision also show modest gains, indicating that the symmetry features help both detection (recall) and reduction of false positives (precision). Visual examples illustrate that the nonlinear registration more accurately aligns homologous structures, leading to sharper SDI maps and better discrimination of lesion voxels.
The authors compare their work with prior symmetry‑based methods (Shen et al., Wang et al.) that relied on template‑based affine transformations and often required the mirrored voxel to lie in the same axial slice. Their reflective registration is template‑free and operates directly on each subject, allowing for more precise alignment of complex anatomical variations. Moreover, the method is modality‑agnostic; while they used FLAIR for the reflection step, any modality could be employed, and the approach can be extended to 3‑D CNNs without architectural changes.
Limitations include the additional computational cost of the registration step (approximately 1–2 minutes per subject) and the fact that experiments were limited to 2‑D CNNs. The authors suggest future work on integrating the symmetry mapping into end‑to‑end trainable networks, reducing registration overhead (e.g., via learned deformation fields), and testing on other lesion types such as tumors or traumatic injuries. They also note that while the current study used a modest dataset (28 subjects), the consistent improvements across two distinct network architectures indicate that the symmetry augmentation is robust and likely to generalize.
In summary, the paper demonstrates that exploiting the natural bilateral symmetry of the brain through nonlinear reflective registration yields informative “symmetry difference” features that, when added to existing multimodal CNNs, significantly boost stroke lesion segmentation performance. This template‑free, modality‑agnostic technique offers a practical and broadly applicable enhancement for medical image segmentation pipelines.
Comments & Academic Discussion
Loading comments...
Leave a Comment