Bridging the gap between individual-based and continuum models of growing cell populations
Continuum models for the spatial dynamics of growing cell populations have been widely used to investigate the mechanisms underpinning tissue development and tumour invasion. These models consist of nonlinear partial differential equations that describe the evolution of cellular densities in response to pressure gradients generated by population growth. Little prior work has explored the relation between such continuum models and related single-cell-based models. We present here a simple stochastic individual-based model for the spatial dynamics of multicellular systems whereby cells undergo pressure-driven movement and pressure-dependent proliferation.We show that nonlinear partial differential equations commonly used to model the spatial dynamics of growing cell populations can be formally derived from the branching random walk that underlies our discrete model. Moreover, we carry out a systematic comparison between the individual-based model and its continuum counterparts, both in the case of one single cell population and in the case of multiple cell populations with different biophysical properties. The outcomes of our comparative study demonstrate that the results of computational simulations of the individual-based model faithfully mirror the qualitative and quantitative properties of the solutions to the corresponding nonlinear partial differential equations. Ultimately, these results illustrate how the simple rules governing the dynamics of single cells in our individual-based model can lead to the emergence of complex spatial patterns of population growth observed in continuum models.
💡 Research Summary
This paper addresses a longstanding gap between individual‑based (agent‑based) descriptions of growing cell populations and the continuum partial differential equation (PDE) models that are commonly used to study tissue development and tumor invasion. The authors first introduce a stochastic lattice‑based model in which each cell occupies a site on a one‑dimensional grid. Time is discretised with step τ and space with step χ. At each time step a cell can (i) proliferate, die, or remain quiescent with probabilities that depend on the local pressure p, and (ii) move left, right, or stay put with probabilities that depend on the pressure differences between the current site and its neighbours. The pressure itself is defined through a barotropic relation p = Π(ρ), where ρ is the total cell density and Π is a smooth, strictly increasing function (often taken as Π(ρ)=K γ ρ^γ with γ>1). Proliferation is pressure‑limited: if p exceeds a homeostatic pressure P, the net growth rate G(p) becomes non‑positive, so cells only die or stay quiescent. Movement is biased down pressure gradients: the probability of moving toward a neighbour is proportional to the positive part of the pressure difference, scaled by a mobility parameter ν_h for each cell type h.
From this microscopic rule set the authors write a mass‑balance equation for the expected number of cells of type h at site i. By expanding the discrete update using Taylor series and taking the formal limit τ→0, χ→0 while keeping the ratio χ²/τ finite, they obtain the continuum limit. The diffusion‑like term emerges as μ_h ∇·(ρ_h ∇p) with μ_h = ν_h χ²/(2τ), while the reaction term becomes G_h(p) ρ_h. Thus the familiar PDE ∂_t ρ_h − μ_h ∇·(ρ_h ∇p) = G_h(p) ρ_h, with total pressure p = Π(∑_h ρ_h), is rigorously derived from the underlying stochastic process. For a single population the equation reduces to the classic form ∂_t ρ − μ ∇·(ρ ∇p) = G(p) ρ.
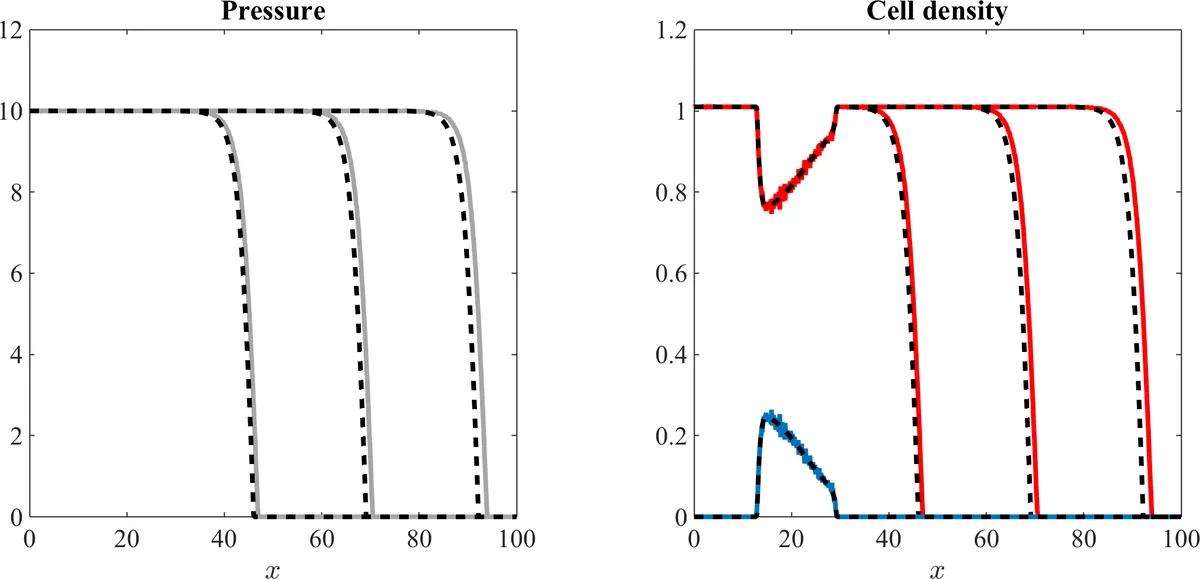
The paper then presents extensive numerical comparisons. In the single‑population case, the authors initialise a compact high‑density region and observe a travelling‑wave front. Stochastic simulations of thousands of cells are performed, and the resulting density profiles are compared with finite‑difference solutions of the PDE. The wave speed, shape, and front steepness match closely, confirming that the macroscopic PDE accurately captures the aggregate behaviour of the microscopic model. In the two‑population scenario (one proliferating, one non‑proliferating) the authors assign different mobilities μ₁≠μ₂ and set the growth term to zero for the second population. Simulations show spatial segregation: the more motile proliferating cells advance ahead, while the less motile non‑proliferating cells lag behind, forming a sharp interface. The PDE system reproduces this segregation and predicts the same interface dynamics. Parameter sweeps illustrate how varying ν_h or the pressure‑growth function G(p) influences wave speed and interface thickness.
Key insights include: (1) a minimal set of biologically motivated rules—pressure‑dependent proliferation and pressure‑gradient‑driven biased random walk—suffices to generate the nonlinear diffusion‑reaction structure observed in continuum models; (2) the derivation clarifies the physical meaning of the mobility coefficient μ as a combination of microscopic jump frequency and step size; (3) the agreement between stochastic and deterministic simulations validates the continuum approximation even at moderate cell numbers, while also highlighting regimes (very low density) where stochastic fluctuations could be important; (4) the framework is readily extensible to multiple cell types, higher spatial dimensions, and additional mechanisms such as chemotaxis or adhesion.
In conclusion, the authors successfully bridge the micro‑macro divide for pressure‑driven growing cell populations, providing both a rigorous derivation and a thorough computational validation. This work offers a solid foundation for future modelling efforts that require both the analytical tractability of PDEs and the detailed realism of individual‑based simulations, and it suggests clear pathways for incorporating more complex biological processes into a unified modelling hierarchy.
Comments & Academic Discussion
Loading comments...
Leave a Comment