Anatomically Consistent Segmentation of Organs at Risk in MRI with Convolutional Neural Networks
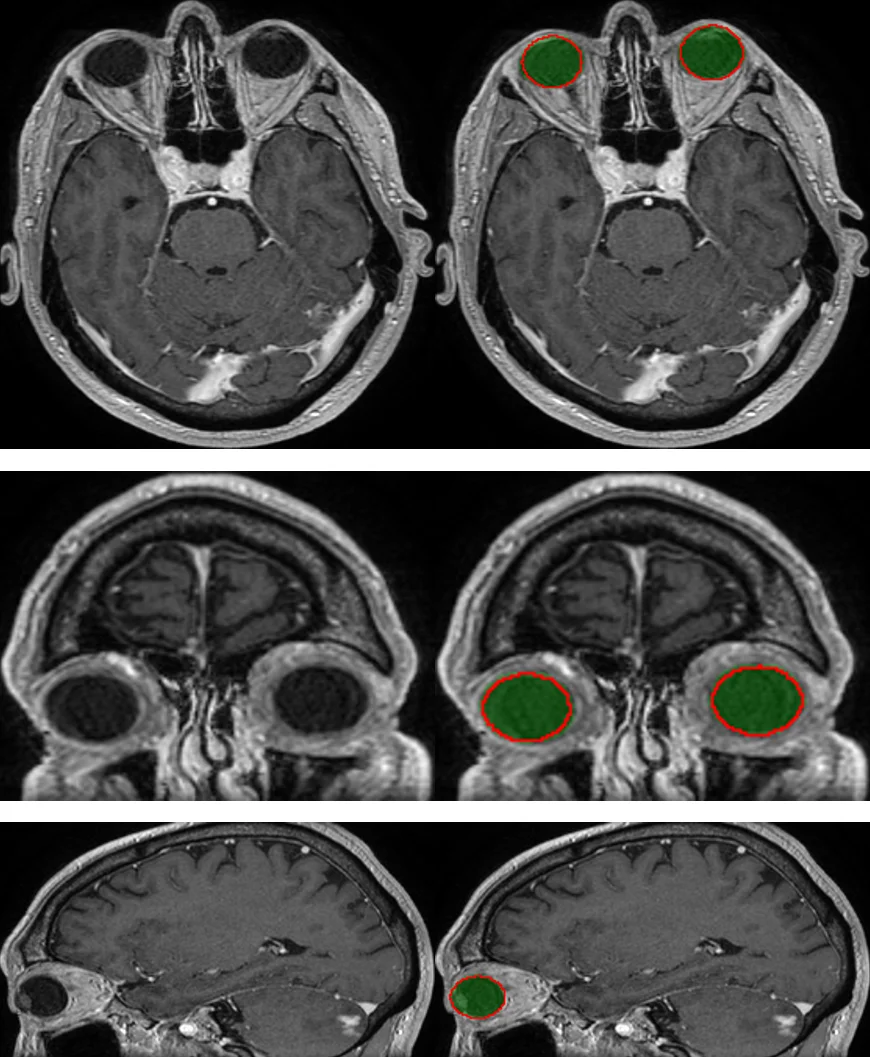
Planning of radiotherapy involves accurate segmentation of a large number of organs at risk, i.e. organs for which irradiation doses should be minimized to avoid important side effects of the therapy. We propose a deep learning method for segmentation of organs at risk inside the brain region, from Magnetic Resonance (MR) images. Our system performs segmentation of eight structures: eye, lens, optic nerve, optic chiasm, pituitary gland, hippocampus, brainstem and brain. We propose an efficient algorithm to train neural networks for an end-to-end segmentation of multiple and non-exclusive classes, addressing problems related to computational costs and missing ground truth segmentations for a subset of classes. We enforce anatomical consistency of the result in a postprocessing step, in particular we introduce a graph-based algorithm for segmentation of the optic nerves, enforcing the connectivity between the eyes and the optic chiasm. We report cross-validated quantitative results on a database of 44 contrast-enhanced T1-weighted MRIs with provided segmentations of the considered organs at risk, which were originally used for radiotherapy planning. In addition, the segmentations produced by our model on an independent test set of 50 MRIs are evaluated by an experienced radiotherapist in order to qualitatively assess their accuracy. The mean distances between produced segmentations and the ground truth ranged from 0.1 mm to 0.7 mm across different organs. A vast majority (96 %) of the produced segmentations were found acceptable for radiotherapy planning.
💡 Research Summary
The paper addresses the clinically critical task of automatically segmenting organs at risk (OAR) in brain magnetic resonance imaging (MRI) for radiotherapy planning. Accurate delineation of eight structures—both eyes, lenses, optic nerves, optic chiasm, pituitary gland, hippocampi, brainstem and the whole brain—is required to minimize radiation exposure to healthy tissue. Manual segmentation is time‑consuming, variable between clinicians, and often incomplete because some organs are only annotated for a subset of patients. Existing automatic approaches fall into two categories: atlas‑based methods, which preserve anatomical consistency but struggle with anatomical variability and pathological deformations, and voxel‑wise machine‑learning methods (e.g., random forests, convolutional neural networks) that can learn complex image features but typically assign a single exclusive label per voxel, leading to anatomically implausible results and difficulty handling missing labels.
To overcome these limitations, the authors propose a modified 2‑dimensional U‑Net architecture that performs end‑to‑end multi‑class segmentation with non‑exclusive classes. Each of the eight target organs is assigned its own binary segmentation head, allowing a voxel to be simultaneously labeled as belonging to multiple structures (e.g., the eye and the lens). This design directly models the overlapping nature of many brain OARs.
A central contribution is a loss function that explicitly handles class imbalance and missing ground‑truth annotations. For each class c, the loss is a weighted pixel‑wise cross‑entropy where the weight of a positive pixel is t_c / N_c^1 and the weight of a negative pixel is (1‑t_c) / N_c^0; unknown pixels receive zero weight. The hyper‑parameter t_c (0 < t_c < 1) controls the relative importance of positives versus negatives, mitigating the severe imbalance typical of small structures. By setting unknown pixels to zero weight, the network can be trained on images where only a subset of organs is annotated, without penalizing the missing information.
Training efficiency is further enhanced by a carefully designed sampling strategy. Bounding boxes for each organ are pre‑computed. In each mini‑batch (size M = 10), the first C = 8 samples are forced to be centered on a positive region of each organ, guaranteeing that every class contributes both positive and negative voxels to the loss. The remaining samples are drawn randomly or centered on the largest organ (the brain) to provide context. This ensures that tiny structures such as lenses and optic nerves are consistently seen during training, despite their small spatial extent.
Because storing binary masks for many classes can be memory‑intensive, the authors encode the multi‑label ground truth as a single integer per voxel, with each bit representing the presence of a specific class. Bitwise operations retrieve individual binary masks on‑the‑fly, reducing I/O overhead.
After the network predicts probability maps for each organ, a post‑processing step enforces anatomical consistency, focusing on the optic nerves. The authors construct a graph where voxels are nodes and edge weights are derived from the predicted nerve probability. Using Dijkstra’s shortest‑path algorithm, they find the minimal‑cost path connecting each eye to the optic chiasm, thereby producing a continuous, anatomically plausible nerve segmentation. This graph‑based refinement corrects common errors such as fragmented or discontinuous nerve predictions that arise from purely voxel‑wise classification.
The method was evaluated on a dataset of 44 contrast‑enhanced T1‑weighted MRIs with ground‑truth segmentations (cross‑validation) and an independent test set of 50 MRIs. Quantitative metrics include Dice similarity coefficient, Hausdorff distance, and mean surface distance (MSD). Across all organs, the MSD ranged from 0.1 mm to 0.7 mm, indicating sub‑millimeter accuracy. Qualitative assessment by an experienced radiotherapist on the independent test set found that 96 % of the automatically generated segmentations were acceptable for treatment planning. These results surpass typical performances of both atlas‑based and prior CNN approaches applied to brain MRI OAR segmentation.
The authors discuss limitations: the 2‑D slice‑wise approach does not capture full 3‑D context, potentially affecting the continuity of structures not explicitly refined by the graph step. Nevertheless, the 2‑D design dramatically reduces GPU memory requirements, enabling training on high‑resolution images and small structures without down‑sampling. Future work could integrate 3‑D context, multi‑modal MRI sequences (e.g., T2, FLAIR), or more sophisticated anatomical priors directly into the loss function.
In conclusion, the paper presents a practical, high‑accuracy solution for automatic OAR segmentation in brain MRI. By combining a multi‑head 2‑D U‑Net with a loss that handles missing labels, a balanced sampling scheme, and a graph‑based post‑processing that restores anatomical connectivity, the method achieves clinically acceptable performance while remaining computationally efficient. This work has the potential to streamline radiotherapy planning workflows, reduce inter‑observer variability, and accelerate patient treatment timelines.
Comments & Academic Discussion
Loading comments...
Leave a Comment