Unsupervised Deformable Image Registration Using Cycle-Consistent CNN
Medical image registration is one of the key processing steps for biomedical image analysis such as cancer diagnosis. Recently, deep learning based supervised and unsupervised image registration methods have been extensively studied due to its excellent performance in spite of ultra-fast computational time compared to the classical approaches. In this paper, we present a novel unsupervised medical image registration method that trains deep neural network for deformable registration of 3D volumes using a cycle-consistency. Thanks to the cycle consistency, the proposed deep neural networks can take diverse pair of image data with severe deformation for accurate registration. Experimental results using multiphase liver CT images demonstrate that our method provides very precise 3D image registration within a few seconds, resulting in more accurate cancer size estimation.
💡 Research Summary
This paper introduces a novel unsupervised deep learning framework for deformable registration of three‑dimensional (3‑D) medical volumes, specifically targeting multiphase liver CT scans. The core idea is to enforce cycle‑consistency between two registration networks, G_AB and G_BA, which predict forward (A→B) and backward (B→A) deformation fields (φ_AB and φ_BA) respectively. Both networks share a U‑Net‑style architecture derived from VoxelMorph‑1, consisting of an encoder‑decoder with skip connections, and output a dense 3‑D displacement field. A 3‑D Spatial Transformer layer (tri‑linear interpolation) warps the moving image according to the predicted field, enabling end‑to‑end differentiable training.
The loss function combines four terms: (1) registration loss (L_register) for each direction, composed of a similarity term (cross‑correlation) and a smoothness regularizer (L2 norm of the field); (2) cycle loss (L_cycle) that penalizes the L1 distance between the original image and the image obtained after a forward‑backward deformation cycle, thereby encouraging inverse consistency; (3) identity loss (L_identity) that forces the network to produce near‑zero deformation when the moving and fixed images are identical, stabilising stationary regions; and (4) weighting coefficients α and β to balance the contributions. This formulation eliminates the need for ground‑truth deformation fields, making the training fully unsupervised.
Experiments were conducted on a dataset of 605 four‑phase liver CT scans (555 for training, 50 for testing) collected at Asan Medical Center, South Korea. Each scan consists of 5 mm thick volumes covering the liver. Pre‑processing involved liver segmentation, zero‑padding around the liver centroid, intensity normalization, and random data augmentations (flipping, 90° rotations). During training, volumes were down‑sampled to 128³ to fit GPU memory (NVIDIA GTX 1080 Ti), while testing used the full 512³ resolution. The optimizer was Adam with a learning rate of 1e‑4, batch size 1, and training lasted 50 epochs.
Performance was evaluated using (i) target registration error (TRE) measured on 20 anatomical landmarks, (ii) tumor size estimation error, (iii) the proportion of voxels with non‑positive Jacobian determinant (a measure of folding), and (iv) normalized mean square error (NMSE) between original and re‑warped images. Compared with the classical Elastix (state‑of‑the‑art intensity‑based registration) and the baseline VoxelMorph‑1, the proposed method achieved: – an average TRE reduction of ~30 % relative to VoxelMorph‑1, while being only slightly higher than Elastix; – a dramatic speedup, processing a full 3‑D volume in ~10 seconds versus ~19.6 minutes for Elastix; – a lower folding rate (0.0175 % vs. 0.0327 % for VoxelMorph‑1), demonstrating superior topology preservation; – the most accurate tumor size measurements in the delayed‑to‑portal phase registration, and comparable accuracy in the arterial‑to‑portal phase.
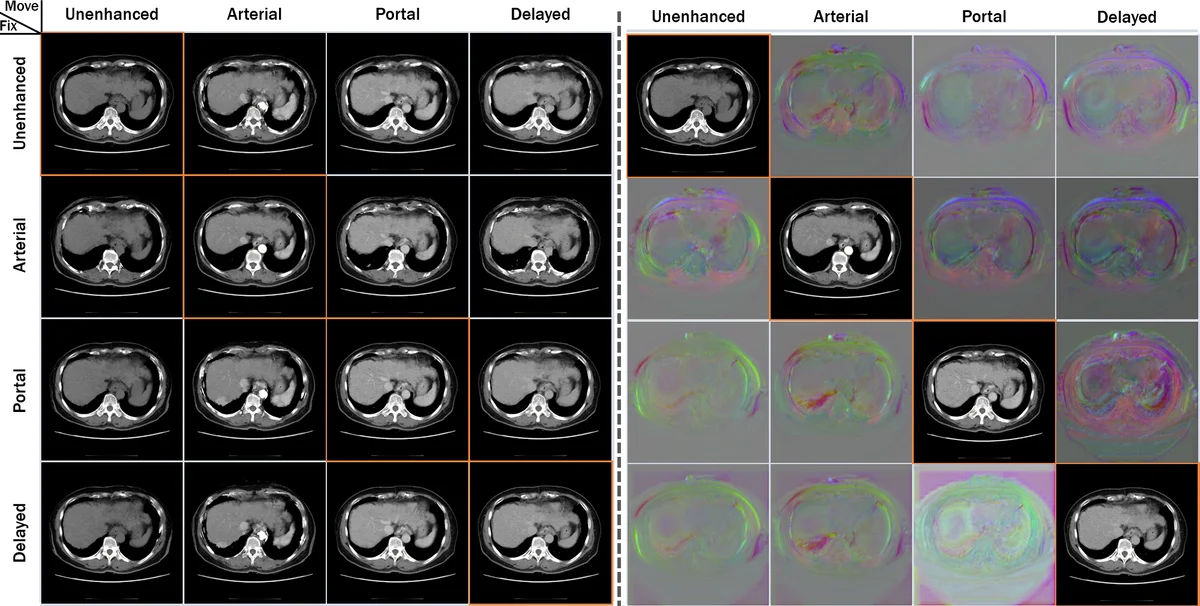
Ablation studies confirmed the importance of each component: removing the cycle loss increased folding and degraded TRE, while omitting the identity loss reduced stability in stationary regions. Visualizations of deformation fields showed smooth, anatomically plausible transformations without noticeable discontinuities.
The paper’s contributions are threefold: (1) introducing cycle‑consistency into unsupervised deformable registration, which enforces inverse consistency and reduces folding; (2) adding an identity loss to preserve stationary anatomy; (3) demonstrating that a single trained model can register any pair of phases in a multiphase CT study, achieving near‑real‑time performance suitable for clinical workflows.
Limitations include reliance on down‑sampling during training (potential loss of fine‑scale detail), evaluation limited to liver CT (generalization to other organs or modalities remains to be shown), and duplicated network parameters that could be streamlined for memory efficiency. Future work may explore multi‑scale cycle architectures, probabilistic deformation modeling, extension to other anatomical regions (brain, heart), and integration into end‑to‑end diagnostic pipelines.
Comments & Academic Discussion
Loading comments...
Leave a Comment