Adversarial optimization for joint registration and segmentation in prostate CT radiotherapy
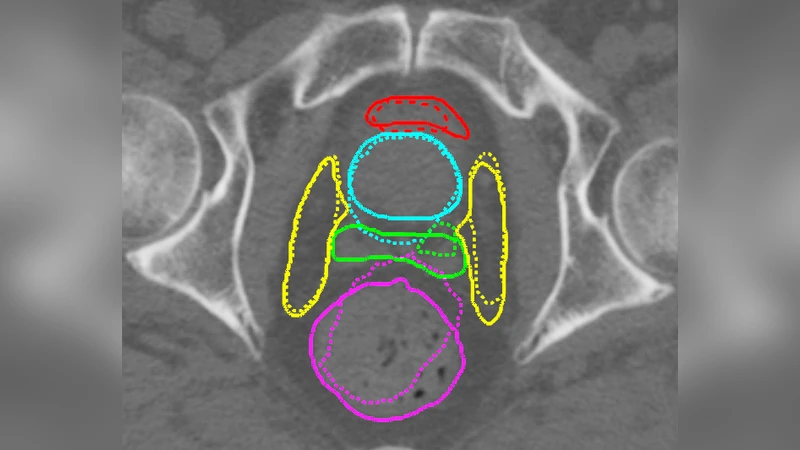
Joint image registration and segmentation has long been an active area of research in medical imaging. Here, we reformulate this problem in a deep learning setting using adversarial learning. We consider the case in which fixed and moving images as well as their segmentations are available for training, while segmentations are not available during testing; a common scenario in radiotherapy. The proposed framework consists of a 3D end-to-end generator network that estimates the deformation vector field (DVF) between fixed and moving images in an unsupervised fashion and applies this DVF to the moving image and its segmentation. A discriminator network is trained to evaluate how well the moving image and segmentation align with the fixed image and segmentation. The proposed network was trained and evaluated on follow-up prostate CT scans for image-guided radiotherapy, where the planning CT contours are propagated to the daily CT images using the estimated DVF. A quantitative comparison with conventional registration using \texttt{elastix} showed that the proposed method improved performance and substantially reduced computation time, thus enabling real-time contour propagation necessary for online-adaptive radiotherapy.
💡 Research Summary
This paper addresses the long‑standing challenge of performing image registration and segmentation jointly, with a particular focus on prostate CT images used in image‑guided radiotherapy. The authors propose a 3‑dimensional generative adversarial network (GAN) that learns to estimate a deformation vector field (DVF) between a fixed and a moving CT scan in an unsupervised manner, while simultaneously leveraging segmentation masks during training. Fixed images (I_f), moving images (I_m), and their corresponding segmentations (S_f, S_m) are available for training; however, at test time only the image pair is required, reflecting the practical scenario where daily CT scans lack manual contours.
The architecture consists of two convolutional neural networks. The generator follows a 3‑D U‑Net design, taking the concatenated fixed and moving images as input and outputting a DVF Φ. A resampling module (adopted from NiftyNet) warps both the moving image and its segmentation using Φ, enabling end‑to‑end back‑propagation. The discriminator is built on the PatchGAN concept and receives three channels: the fixed image, the warped moving image, and the warped moving segmentation. Two variants are explored: (a) feeding the segmentation as an independent channel (JRS‑GAN a) and (b) multiplying the segmentation with the corresponding image to emphasize anatomical structures (JRS‑GAN b).
Training employs a Wasserstein GAN (WGAN) loss for stability, but the authors observed slow convergence when using WGAN alone. Consequently, they augment the generator loss with a similarity term L_sim that combines (1 – Dice similarity coefficient) for the warped and fixed segmentations and (1 – normalized cross‑correlation) for the warped and fixed images. A smoothness regularizer on the DVF (bending energy) is also added, yielding the final generator loss: L_G = L_sim + λ₁ L_smooth + λ₂ L_GAN. During the first 25 training epochs the discriminator is updated 100 times per generator update; thereafter a 1:5 update ratio is used, with weight clipping as prescribed for WGAN.
The method was evaluated on a dataset of 18 prostate cancer patients, each with a planning CT and 7–10 daily repeat CTs, resulting in 111 image pairs for training/validation and 50 pairs for testing. Ground‑truth contours for prostate, seminal vesicles, lymph nodes, bladder, and rectum were available for evaluation. Performance was measured using mean surface distance (MSD) and 95 % Hausdorff distance (HD). Comparisons included traditional iterative registration (elastix with NCC and MI), two non‑adversarial deep learning baselines (Reg‑CNN using only NCC loss, and JRS‑CNN using NCC + Dice loss), and an adversarial baseline without segmentation input (Reg‑GAN).
Results show that all GAN‑based approaches significantly outperform elastix in both MSD and HD across all structures. The joint registration‑segmentation models (JRS‑GAN a and JRS‑GAN b) achieve the lowest errors, with average MSD values around 1.1 mm for the prostate and 1.0 mm for seminal vesicles, compared to 1.8 mm (elastix‑NCC) and 1.7 mm (elastix‑MI). HD improvements are especially pronounced for organs‑at‑risk (bladder, rectum), where JRS‑GAN reduces 95 % HD from ~11 mm (elastix) to ~3 mm. The inclusion of segmentation masks in the discriminator yields a modest but consistent gain over Reg‑GAN, confirming that anatomical priors help the discriminator learn a more meaningful alignment metric. Runtime analysis demonstrates a dramatic speedup: the proposed pipeline processes a 256³ voxel volume in 0.6 seconds on an NVIDIA V100 GPU, versus 13 seconds for elastix on a 4‑core CPU.
The authors discuss that while the Jacobian determinant’s standard deviation is higher for the learned DVFs (indicating greater flexibility), the resulting deformations remain diffeomorphic enough for clinical use. They note that the benefit of the two discriminator designs is minor, possibly due to the small size and irregular shape of the seminal vesicles. Future work is suggested to explore improved GAN objectives (e.g., gradient‑penalty regularization), multi‑scale discriminators, and extension to other anatomical sites or multimodal imaging.
In conclusion, the paper presents a fast, accurate, and fully unsupervised method for joint image registration and segmentation of prostate CT scans, leveraging adversarial training to incorporate segmentation information during learning while requiring no manual contours at inference time. The approach achieves real‑time performance suitable for online‑adaptive radiotherapy, marking a significant step toward clinically viable deep‑learning‑based contour propagation.
Comments & Academic Discussion
Loading comments...
Leave a Comment