An attention-based multi-resolution model for prostate whole slide imageclassification and localization
Histology review is often used as the `gold standard’ for disease diagnosis. Computer aided diagnosis tools can potentially help improve current pathology workflows by reducing examination time and interobserver variability. Previous work in cancer grading has focused mainly on classifying pre-defined regions of interest (ROIs), or relied on large amounts of fine-grained labels. In this paper, we propose a two-stage attention-based multiple instance learning model for slide-level cancer grading and weakly-supervised ROI detection and demonstrate its use in prostate cancer. Compared with existing Gleason classification models, our model goes a step further by utilizing visualized saliency maps to select informative tiles for fine-grained grade classification. The model was primarily developed on a large-scale whole slide dataset consisting of 3,521 prostate biopsy slides with only slide-level labels from 718 patients. The model achieved state-of-the-art performance for prostate cancer grading with an accuracy of 85.11% for classifying benign, low-grade (Gleason grade 3+3 or 3+4), and high-grade (Gleason grade 4+3 or higher) slides on an independent test set.
💡 Research Summary
This paper addresses the challenging problem of automated Gleason grading for prostate cancer whole‑slide images (WSIs) by proposing a two‑stage, attention‑based multiple‑instance learning (MIL) framework that requires only slide‑level labels. The authors first note that existing approaches either rely on manually annotated regions of interest (ROIs) or need massive amounts of fine‑grained tile annotations, both of which are impractical for routine clinical workflows. To overcome these limitations, they treat each WSI as a “bag” and each extracted tile as an “instance” within a MIL setting, and they embed a learnable attention mechanism to weight the contribution of each tile to the slide‑level prediction.
Stage 1 – Low‑resolution screening (5×):
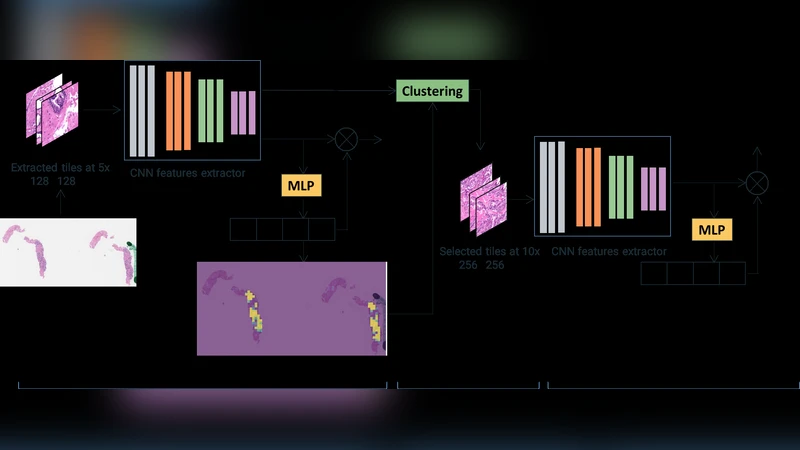
Tiles of size 128 × 128 px are extracted at 5× magnification. A VGG‑11 with batch normalization (pre‑trained on ImageNet) serves as the backbone to produce 1024‑dimensional feature vectors for each tile. An MLP‑based attention module computes a softmax‑normalized weight αi for each instance, indicating its relevance to the binary cancer vs. non‑cancer decision. To prevent the model from focusing exclusively on a few highly discriminative tiles, the authors adopt “instance dropout” during training: randomly selected tiles have their pixel values replaced by the dataset mean, forcing the network to learn from a broader set of regions.
Stage 2 – High‑resolution grading (10×):
From the attention map and the instance features of Stage 1, the authors perform dimensionality reduction (PCA to 32 dimensions) followed by K‑means clustering (k = 4). For each cluster, the average attention score is calculated; clusters with higher average attention are presumed to contain diagnostically relevant tissue. A proportionate number of tiles from these high‑attention clusters are then re‑extracted at 10× magnification (same spatial locations) and fed into a second MIL model that classifies the slide into three categories: benign, low‑grade (Gleason 3+3 or 3+4), and high‑grade (Gleason 4+3 or higher).
Datasets and preprocessing:
Two data sources are used. A small, pixel‑level annotated dataset from Cedars‑Sinai (≈12 k tiles) is employed to pre‑train the tile‑level classifier and to initialize the feature extractor. The main training set consists of 3,521 prostate biopsy slides from UCLA (718 patients) with only slide‑level labels extracted from pathology reports. Tissue masks are generated by converting the lowest‑magnification image to HSV, thresholding the hue channel, and applying morphological operations; tiles with <80 % tissue are discarded. Color normalization follows the method of Macenko et al.
Implementation details:
The first‑stage model is trained with the feature extractor frozen for 10 epochs (learning rate 1e‑4 for attention and classifier), then fine‑tuned for 90 epochs (learning rate 1e‑5 for the backbone, 1e‑4 for the classifier). Instance dropout is set to 0.5. The attention hidden dimension h is 512. For the second stage, the same backbone is initialized with the pre‑trained weights, and the model is trained for five epochs with the extractor frozen. Hyper‑parameters (number of clusters, dropout rate, learning rates) are tuned on a validation split.
Results:
On an independent test set of 860 slides (227 patients), the two‑stage system achieves 85.11 % overall accuracy for the three‑class problem (benign, low‑grade, high‑grade). This outperforms previously reported methods: Zhou et al. (75 %), Xu et al. (79 %), Nagpal et al. (70 %). The confusion matrix shows particularly strong separation between low‑ and high‑grade cases. Visualizations of the attention maps demonstrate that the model reliably highlights cancerous regions, providing interpretable localization without explicit ROI supervision.
Key contributions and limitations:
- Demonstrates that slide‑level labels alone are sufficient for high‑performing Gleason grading when combined with attention‑based MIL and instance dropout.
- Introduces a clinically inspired two‑stage workflow that mimics pathologists’ low‑magnification scanning followed by high‑magnification examination.
- Provides interpretable saliency maps that can serve as a second reader for pathologists.
Limitations include sensitivity to clustering hyper‑parameters, reliance on fixed magnification steps (5× → 10×), and the need for external validation on other organs or cancer types. Future work could explore multi‑scale pyramids, self‑supervised pre‑training, and broader multi‑institutional datasets to improve generalizability.
Comments & Academic Discussion
Loading comments...
Leave a Comment