Deep Transfer Learning Methods for Colon Cancer Classification in Confocal Laser Microscopy Images
Purpose: The gold standard for colorectal cancer metastases detection in the peritoneum is histological evaluation of a removed tissue sample. For feedback during interventions, real-time in-vivo imaging with confocal laser microscopy has been propos…
Authors: Nils Gessert, Marcel Bengs, Lukas Wittig
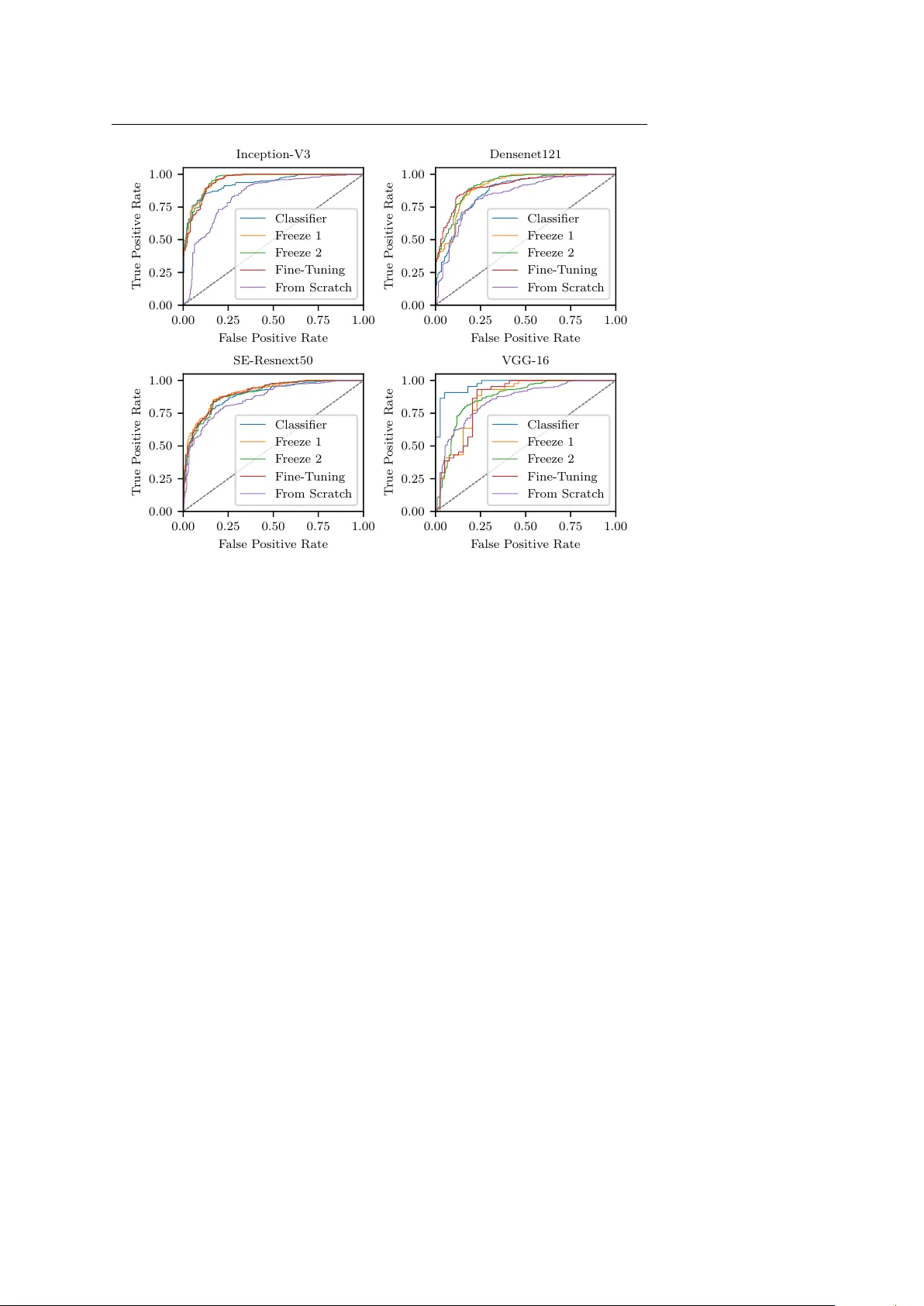
Deep T ransfer Learning Metho ds for Colon Cancer Classification in Confo cal Laser Microscop y Images Nils Gessert 1 ∗ , Marcel Bengs 1 ∗ , Luk as Wittig 2 , Daniel Dr¨ omann 2 , T obias Kec k 3 , Alexander Schlaefer 1 , David B. Ellebrec ht 3 Preprint. Accepted for publication in IJCARS. Abstract Purp ose The gold standard for colorectal cancer metastases detection in the peritoneum is histological ev aluation of a remo ved tissue sample. F or feedback during in terven- tions, real-time in-vivo imaging with confo cal laser microscopy has b een prop osed for differentiation of benign and malignant tissue by man ual exp ert ev aluation. Automatic image classification could impro ve the surgical w orkflow further by pro viding immediate feedback. Metho ds W e analyze the feasibility of classifying tissue from confo cal laser microscopy in the colon and p eritoneum. F or this purpose we adopt b oth classical and state- of-the-art conv olutional neural netw orks to directly learn from the images. As the a v ailable dataset is small, we in vestigate sev eral transfer learning strategies including partial freezing v ariants and full fine-tuning. W e address the distinction of different tissue t yp es, as w ell as b enign and malignant tissue. R esults W e presen t a thorough analysis of transfer learning strategies for colorectal cancer with confo cal laser microscopy . In the peritonuem, metastases are classified with an AUC of 97 . 1 and in the colon the primarius is classified with an AUC of 73 . 1. In general, transfer learning substan tially impro ves p erformance o ver training from scratc h. W e find that the optimal transfer learning strategy differs for models and classification tasks. Conclusions W e demonstrate that conv olutional neural netw orks and transfer learning can b e used to identify cancer tissue with confocal laser microscop y . W e sho w that there is no generally optimal transfer learning strategy and mo del as well as task- sp ecific engineering is required. Giv en the high p erformance for the peritoneum, Nils Gessert, E-mail: nils.gessert@tuhh.de, T el.: +49 (0)40 42878 3389, https://orcid.org/0000-0001-6325-5092 ∗ Authors con tributed equally 1 Institute of Medical T ec hnology , Ham burg Universit y of T ec hnology , Hamburg, German y 2 Department of Pulmology , Univ ersity Medical Centre Schleswig-Holstein, L ¨ ubeck, German y 3 Department of Surgery , Universit y Medical Centre Schleswig-Holstein, L ¨ ubeck, German y 2 Gessert et al. ev en with a small dataset, application for intraoperative decision supp ort could be feasible. Keyw ords Colon Cancer · Confo cal Laser Microscopy · T ransfer Learning · Con volution Neural Netw ork 1 Introduction Colorectal cancer is very common and it is often associated with metastatic spread [1]. In particular, p eritoneal carcinomatosis (PC) can arise in later stages of de- v elopment which often shortens patien t surviv al times substantially [2, 3]. Th us, early and reliable detection of metastases is crucial. Diagnosis with t ypical external imaging techniques such as computed tomography (CT) and magnetic resonance imaging (MRI) is difficult for PC as a very high resolution is required. F or example, preop erativ e CT has b een shown to b e ineffectiv e to detect individual p eritoneal tumor dep osits and the interobserv er v ariability among exp erts w as significant [4]. Also, integrated PET/CT did not provide sufficien t information for accurate as- sessmen t [5]. F or MRI, studies hav e sho wn impro vemen t ov er assessmen t with CT only [6, 7] but o verall, its resolution is still a limitation [8]. Therefore, exploratory laparoscop y is generally employ ed to inv estigate the presence of PC [9]. Recen tly , a new intraoperative device using confo cal laser microscopy (CLM) has b een in tro duced which provides submicrometer image resolution [10]. In the study , ten rats receiv ed colon carcionoma cell implants in the colon and p eri- toneum. After a growth p erio d, laparotom y with in-vivo CLM was performed. CLM images of healthy and malignant colon tissue, as well as healthy and malig- nan t p eritoneum were acquired. It was shown that exp erts are able to distinguish differen t tissue types as w ell as health y and malignan t tissue from CLM. This raises the question whether image pro cessing tec hniques can b e used to automatically classify different tissue types. This could enable faster and improv ed in traop erativ e decision supp ort with CLM. Recen tly , automatic tissue characterization has b een successfully addressed using deep learning methods such as con volutional neural net works (CNNs) for seman tic segmentation and classification [11, 12]. F or example, skin cancer classifi- cation at dermatologist-lev el performance w as ac hieved [13]. How ever, the datasets for this and related studies are large and commonly , datasets for medical learn- ing tasks are small [14]. This can b e problematic as insufficient data for optimal training might lead to ov erfitting and limited generalization. This is particularly imp ortan t for deep learning mo dels whic h can b e prone to ov erfitting due to their large num b er of trainable parameters. T o ov ercome this issue, transfer learning metho ds hav e been proposed where a deep learning mo del is first pretrained on a differen t, large dataset [15]. Then, information from the source domain can b e transferred to the (medical) target domain using strategies such as ”off-the-shelf” features, partial lay er freezing, or full fine-tuning [16]. While this has b een suc- cessfully applied for medical learning tasks [17], there is no single solution for all problems and the optimal transfer learning strategy is highly dependent on the imaging mo dality and dataset size [18]. Automatic analysis of CLM images has b een proposed for differen t tissue t yp es suc h as human skin [19], the cornea [20] or the oral cavit y [21]. Recently , deep Deep T ransfer Learning Metho ds for Colon Cancer Classification with CLM 3 learning metho ds ha ve b een applied to CLM and similar mo dalities. F or example, CNNs hav e b een used for oral squamous cell carcinoma classification [21] and motion correction with CLM [22]. Similarly , skin images from CLM hav e been used with CNN-based classification [23]. F or the gastroin testinal tract, CNNs hav e b een used to distinguish three classes of Barret’s esophagus [24]. Also, brain tumor classification with CNNs and CLM has shown promising results [25]. F or example, a CNN has b een used to differen tiate CLM images with and without diagnostic v alue for a physician during surgery [26]. Also, weakly-supervised localization has b een used to derive lo cal information in CLM images from image-level lab els only [27]. So far, deep learning-based classification of colorectal cancer from CLM images has not b een addressed. Also, while sev eral approaches ha ve used CLM and CNNs for other problems [28], there is no analysis of transfer learning prop erties for col- orectal cancer with CLM. Therefore, w e study deep learning-based colon cancer classification from CLM images with a v ariety of transfer learning metho ds from the ImageNet dataset. W e consider training from scratc h, partial la yer freezing, ”off-the-shelf” features and full fine-tuning to inv estigate how transferable Ima- geNet features are to CLM. W e p erform this study with the classic mo dels VGG- 16 [29] and Inception-V3 [30] as well as the state-of-the-art architectures Densenet [31] and squeeze-and-excitation net w orks [32] to analyze the consistency of transfer strategies across architectures. W e consider the classes health y colon (HC), ma- lignan t colon (MC), healthy p eritoneum (HP) and malignan t peritoneum (MP). Based on these classes, we address three binary classification tasks with CLM. First, we consider the differentiation of organs (HP vs. HC). Then, we study the detection of malignant tissue in tw o types of organs (HP vs. MP and HC vs. MC). This allows us to study v ariations across different classification tasks for CLM. A preliminary v ersion of this paper was presen ted at the BVM W orkshop 2019 [33]. W e substantially revised the pap er, extended the review of the literature and p erformed more experiments with additional transfer strategies and more archi- tectures. This pap er is structured as follows. First, we describ e our mo dels and transfer learning stratgies and the data set we use in Section 2. Then we rep ort our results in Section 3 and discuss them in Section 4. Last, w e conclude in Section 5. 2 Metho ds 2.1 Model Architectures and T raining Strategies First, w e consider the classic mo del VGG-16 [29] with the addition of batch normal- ization which enables faster training of the architecture by reducing the internal co v ariate shift [34]. The mo del itself is simple as it consists of sev eral stac ked con- v olutional lay ers without further augmentation. In b etw een blo c ks of tw o to three con volutional la yers with kernel sizes of 3 × 3 and 1 × 1, max p ooling reduces the spatial dimensions. Subsequent conv olutions double the num b er of feature maps. A building blo ck of the architecture is shown in Figure 1 (top left). Due to its simple structure, the architecture can serv e as a baseline. Second, w e employ Inception-V3 [30]. The mo del consists of multiple Inception blo c ks which follow t wo core design principles. First, the blo cks hav e a multi- path structure, i.e., the input feature maps are pro cessed in parallel by different 4 Gessert et al. Input F in Output F out ResNext H × W × C Avg. Pool FC - σ 1 × 1 × C 1 × 1 × C + Input F in ResBlock F in + k F in + N k ResBlock Compress Avg. Pool Output F out b b b b b b b b b b b b (c) (d) Input F in C. 3 × 3 F 12 2 × C. 3 × 3 F 22 Output F out (b) Input F in C. 3 × 3 2 F in C. 3 × 3 2 F in Output F out (a) C. 1 × 1 F 21 C. 1 × 1 F 11 Pool F in Fig. 1: The building blocks of the models we use. The building blocks from CNN arc hitectures as indicated in Figure 2. W e employ VGG-16 (a), Inception-V3 (b), Densenet121 (c) and SE- Resnext50 (d). F denotes the num b er of feature maps in each blo c k. The Conv blo c ks also contain ReLU activ ations and batch normalization for VGG-16 (a). SE-Resnext50 is shown in simplified form without its b ottleneck in the SE mo dule. FC- σ is a fully-connected lay er with sigmoid activ ation. C. is an abbreviation for conv olutional layers. Note that Inception-V3 employs multiple blo c k v ariants and w e show one example. con volution and p o oling op erations. At the blo c k’s output, the feature maps from all paths are concatenated. Second, the conv olutional paths p erform a reduction op eration that downsizes the feature map dimension with 1 × 1 k ernels. Then, computationally more exp ensiv e 3 × 3 conv olutions pro cess the lo wer dimensional represen tations. The output feature map size is increased if the spatial dimensions are reduced inside the blo c k whic h a voids represen tational limitations. The idea of reduction and expansion has also found its w ay into the Resnet arc hitecture [35] which is a core comp onent of the next tw o mo dels. Resnets learn a residual instead of a full feature transformation by using skip connections. In detail, a Resnet blo ck (ResBlo c k) computes x ( l ) = a ( F ( x ( l − 1) , θ ( l ) ) + x ( l − 1) ) (1) where x ( l ) is the block output, x ( l − 1) is the block input, a is a ReLU activ ation [36] and F represen ts t wo conv olutional lay ers with parameters θ ( l ) . The skip connection enables b etter gradient propagation for improv ed training. Third, we consider Densenet121 [31], a state-of-the-art architecture whic h strives for more efficiency b y in tro ducing extensiv e feature reuse. In particular, within one DenseBlo c k, features computed in previous lay ers are also fed into the subsequent la yers. T o k eep the feature map sizes mo derate, compression blo cks reduce the feature maps b et w een DenseBlo cks. The DenseBlock is sho wn in Figure 1 (b ottom left). Deep T ransfer Learning Metho ds for Colon Cancer Classification with CLM 5 Input Crop Mo del Block b b b 1 Mo del Block N − 1 Mo del Block N FC La y er C = 2 Retrain Classifier Pa rtial F reeze 1 Pa rtial F reeze 2 F ull Fine-T uning Fig. 2: The different transfer learning scenarios w e inv estigate. Model Blo c k refers to one of the blo c ks shown in Figure 1. Green indicates that blo cks are retrained. Red indicates that blocks are frozen with their w eights ha ving b een trained on ImageNet. F ourth, we adopt the architecture SE-Resnext50 [32]. At its core, the mo del uses Resnext blo cks [37] which are an extension of Resnet. Here, the single con- v olutional path F is split into multiple paths with individual lay ers which in- creases representational p o wer. The key addition in SE-Resnext50 is the use of squeeze-and-excitation (SE) mo dules which recalibrate the f eature maps learned b y Resnext blo c ks. These mo dules ha ve shown improv ed p erformance with only a minimal increase in the n umber of parameters. The concept is sho wn in Figure 1 (b ottom right). Due to the small dataset size, we study sev eral transfer learning strategies where the ab o ve-men tioned mo dels are trained on ImageNet. W e cut off the last la yer of all mo dels and replace it with a fully-connected lay er with t wo outputs for binary classification. W e apply a softmax lay er on top and the final classification output is the class with the highest probabilit y . W e train a separate model for eac h of our binary classification tasks. As a baseline, we consider training from scratc h, i.e. all weigh ts are randomly initialized. Then, we use several different transfer learning strategies illustrated in Figure 2. The first transfer approach follows the ”off-the-shelf” features idea. Here, only the new classifier is retrained on features extracted by the pretrained CNN. W e also consider tw o partial freezing metho ds, where an initial part of the netw ork remains frozen and the part closer to the classifier is retrained. W e c hose the freezing p oin ts blo ck-wise, i.e. we do not cut in to building blo c ks. Last, w e consider full fine-tuning where all weigh ts in the netw ork are retrained with a small learning rate. The different strategies represent different abstractions of feature transfer b et ween ImageNet and CLM images. T o further impro ve generalization, we employ online data augmentation with random image flipping and random c hanges in brightness and contrast. F urther- more, w e use random cropping with crops of size 224 × 224 (299 × 299 for Inception- V3) taken from the full images of size 384 × 384. W e use the Adam algorithm for optimization. W e adapt learning rates and the num b er of training ep ochs for the differen t transfer scenarios. W e use a cross-entrop y loss function with additional w eighting to accoun t for the slight class imbalance. In detail, w e multiply the loss of a training example b y N /n i where N is the total num b er of training examples in the current fold and n i is the num b er of examples b elonging to class i in the 6 Gessert et al. Fig. 3: Examples for the differen t classes. Malignant colon tissue, healthy colon tissue, malig- nant p eritoneum tissue and healthy p eritoneum tissue are shown from left to right. curren t fold. In this wa y , underrepresented classes receive a higher weigh ting in the loss function. During ev aluation, we use mutli-crop ev aluation with N c = 36 ev enly spread crops ov er the images. This ensures that all image regions are co v- ered with large o verlaps betw een crops. The final predictions are a veraged o ver the N c crops. W e implement our mo dels in PyT orch. 2.2 Dataset and Exp erimen ts The dataset w as collected in a previous study conducted at the Univ ersity Hospital Sc hleswig-Holstein in L¨ ub ec k where exp ert assessmen t of CLM images in the colon area was ev aluated [10]. A custom intraoperative device with integrated CLM (Karl Storz GmbH & Co KG, T uttlingen, Germany) was built. The image resolution w as 384 × 384 pixels whic h co vers a field of view of 300 µ m × 300 µ m. In the study , ten rats received colon adeno carcinoma cell implantation in the colon and p eritoneum with a growth time of seven days. Then, laparotomy was conducted and images of health y colon tissue, malignan t colon tissue, health y p eritoneum tissue and malignant p eritoneum tissue w ere obtained. Example CLM images for eac h tissue type are sho wn in Figure 3. After remov al of low qualit y images, 1577 images remained with 533 belonging to class HC, 309 b elonging to class MC, 343 b elonging to class HP and 392 b elonging to class MP . Note that some sub jects are missing classes such that, on av erage, six sub jects p er class remain. Ground-truth annotation of all images was obtained by tissue remov al of the scanned areas and subsequen t histological ev aluation. Due to the small dataset size, we chose a cross-v alidation scheme where im- ages from one sub ject are left for ev aluation and training is p erformed on the remaining ones. Thus, all rep orted results are the mean v alue of, on av erage, six training scenarios with six different folds. Based on the four classes, we address three binary classification problems. First, we consider HC vs. HP , i.e., w e in- v estigate the feasibilit y of distinguishing the different organs in CLM. Then, we consider the differentiation of healthy and malignant tissue with the tw o binary classification problems HP vs. MP and HC vs. MC. W e report the accuracy , sen- sitivit y , sp ecificit y , F1-score and AUC. W e use the AUC as the main metric as it is threshold-indep endent. Deep T ransfer Learning Metho ds for Colon Cancer Classification with CLM 7 1 2 3 4 5 T raining T yp e 0 . 20 0 . 25 0 . 30 0 . 35 0 . 40 0 . 45 0 . 50 0 . 55 0 . 60 0 . 65 0 . 70 0 . 75 0 . 80 0 . 85 0 . 90 0 . 95 1 . 00 A UC HC vs. HP DenseNet SE-RX Inception V GG16BN 1 2 3 4 5 T raining T yp e 0 . 20 0 . 25 0 . 30 0 . 35 0 . 40 0 . 45 0 . 50 0 . 55 0 . 60 0 . 65 0 . 70 0 . 75 0 . 80 0 . 85 0 . 90 0 . 95 1 . 00 HP vs. MP DenseNet SE-RX Inception V GG16BN 1 2 3 4 5 T raining T yp e 0 . 20 0 . 25 0 . 30 0 . 35 0 . 40 0 . 45 0 . 50 0 . 55 0 . 60 0 . 65 0 . 70 0 . 75 0 . 80 0 . 85 0 . 90 0 . 95 1 . 00 HC vs. MC DenseNet SE-RX Inception V GG16BN Fig. 4: AUC v alues of all applied architectures for the different classification problems. W e ev aluate the follo wing training t yp es: (1) retrain classifier, (2) partial freeze 1, (3) partial freeze 2, (4) full fine-tuning, (5) training from scratch. F or each v alue the standard deviation ov er multiple folds is represented b y an error bar. 3 Results First, we compare the different transfer learning scenarios describ ed in Section 2 across all arc hitectures for eac h classification scenario, see Figure 4. In general, the A UC is very high for the differentiation of different health y tissue types and health y and malignant p eritoneum tissue. The AUC for classifying malignant colon tissue is substan tially low er. Also, the standard deviation is higher for this task. T raining from scratch p erforms worst for all architectures and classification scenarios. Regarding the transfer learning scenarios, training from scratc h performs worst for all classification scenarios. F or tw o of the three scenarios, only retraining the classifier shows substantially low er p erformance than other transfer scenarios. There are no clear trends betw een the partial freezing and fine-tuning scenarios. Second, we go in to more details for the classification task HP vs. MP . Figure 5 sho ws the ROC curves for all mo dels with all transfer learning scenarios for the classification task. Op erating points with a go od trade-off in the upp er left corner v ary for each mo del. F or VGG-16, retraining the classifier only stands out. F or Densenet121, partial freezing p erforms well. F or Inception-V3 and SE-Resnext50, partial freezing and fine-tuning p erform similar. Third, an ov erview of the b est p erforming transfer strategies is shown in T a- ble 1. Comparing individual results for each architecture, no mo del clearly out- p erforms the others consistently . In general, Densenet121 p erforms slightly b etter across the tasks. The optimal transfer strategy differs across mo dels and classifi- cation tasks. F or HC vs. HP and for Densenet121 in general, the partial freezing metho d p erforms b est. 8 Gessert et al. 0 . 00 0 . 25 0 . 50 0 . 75 1 . 00 F alse P ositiv e Rate 0 . 00 0 . 25 0 . 50 0 . 75 1 . 00 T rue P ositiv e Rate Inception-V3 Classifier F reeze 1 F reeze 2 Fine-T uning F rom Scratc h 0 . 00 0 . 25 0 . 50 0 . 75 1 . 00 F alse P ositiv e Rate 0 . 00 0 . 25 0 . 50 0 . 75 1 . 00 T rue P ositiv e Rate Densenet121 Classifier F reeze 1 F reeze 2 Fine-T uning F rom Scratc h 0 . 00 0 . 25 0 . 50 0 . 75 1 . 00 F alse P ositiv e Rate 0 . 00 0 . 25 0 . 50 0 . 75 1 . 00 T rue P ositiv e Rate SE-Resnext50 Classifier F reeze 1 F reeze 2 Fine-T uning F rom Scratc h 0 . 00 0 . 25 0 . 50 0 . 75 1 . 00 F alse P ositiv e Rate 0 . 00 0 . 25 0 . 50 0 . 75 1 . 00 T rue P ositiv e Rate V GG-16 Classifier F reeze 1 F reeze 2 Fine-T uning F rom Scratc h Fig. 5: ROC curv e for the different architectures and the different training types, sho wn for the classification of HP vs. MP . Last, w e pro vide training times for all architectures and training scenarios, see Figure 6. In general, freezing more w eights during training reduces the o verall training time. F urthermore, training time lo osely scales with the num b er of train- able parameters as VGG-16 contains the most parameters and shows the longest training times, follow ed by SE-Resnext50. 4 Discussion W e study deep transfer learning metho ds for CLM images for three binary clas- sification problems. Automatic decision supp ort with CLM during interv entions could impro ve workflo w with immediate feedback on the tissue properties. F or this purp ose we inv estigate the use of CNNs with four differen t architectures and fiv e training scenarios. The three classification tasks. As a baseline, differentiating healthy colon and peritoneum tissue w orks w ell with an A UC o ver 0 . 90 for partial freezing across all mo dels, see Figure 4. This indicates that discriminative features for different organs can b e learned from CLM images. Similarly , for classification of metastases in the p eritoneum the AUC is around 0 . 90 for all transfer learning scenarios. How- ev er, classifying healthy and malignant colon tissue p erforms substantially worse with an AUC of ≈ 0 . 70 for partial freezing and fine-tuning. The task app ears to Deep T ransfer Learning Metho ds for Colon Cancer Classification with CLM 9 T able 1: The b est p erforming transfer learning method for each model and classification task. Densenet refers to the Densenet121 mo del, SE-RX50 refers to the SE-Resnext50 mo del. F or each training scenario, the b est performing configuration is marked bold. All v alues are given in percent. The sensitivity is given with resp ect to the cancer class and for the case of organ differentiation it is given with respect to the p eritoneum class. Type Accuracy Sensitivity Specificity F1-Score A UC HC vs. HP Inception F reeze 1 87.7 79.9 94.4 90.4 95.7 Densenet F reeze 1 91.2 82.8 95.3 91.9 92.6 SE-RX50 F reeze 1 85.8 78.5 96.3 91.3 91.9 VGG-16 F reeze 1 82.5 74.9 91.8 87.2 91.6 HP vs. MP Inception F reeze 2 85.9 86.6 87.0 86.8 95.6 Densenet F reeze 2 83.3 84.6 83.2 84.0 91.9 SE-RX50 F reeze 1 81.7 84.6 83.2 84.0 90.9 VGG-16 Classifier 88.0 91.0 84.6 87.9 97.1 HC vs. MC Inception Fine-T uning 63.1 71.0 57.0 63.7 68.0 Densenet F reeze 1 70.0 72.9 64.1 69.1 73.1 SE-RX50 Fine-T uning 63.7 66.7 65.9 69.1 71.8 VGG-16 F reeze 2 63.5 67.6 64.2 68.1 72.0 b e more difficult which is also reflected in a slightly higher standard deviation. This indicates higher uncertaint y of mo del predictions. This could b e caused by the heterogeneous app earance of colon tissue in different parts of the colon which complicates the learning task in conjunction with the small dataset size. F urther- more, during developmen t, colon carcinoma cells transform from a healthy stage to adenoma and then carcinoma. At earlier stages, healthy and malignant cells can still hav e similar app earance whic h complicates the learning task. T ransfer learning scenarios. Figure 4 also provides an o verview of the trans- fer strategies across all models. Clearly , transfer learning substantially outp erforms training from scratch across all classification tasks which supp orts the effective- ness of transfer learning for medical image classification problems [38]. The results indicate that meaningful feature transfer from the natural image domain to CLM images is p ossible, although the images hav e a v astly different appearance. How- ev er, comparing transfer strategies, only retraining the classifier p erforms worse than other scenarios in tw o out of three classification tasks. This agrees with re- sults of a previous study on transfer learning with CLM images in neurosurgery [28]. Here, the authors found that full fine-tuning outp erforms retraining of the classifier only . How ever, in our case, retraining the classifier only also shows a high p erformance for the task HP vs. MP . This could be caused b y fragile co-adaptation of w eights [39] which leads to large p erformance differences b et ween the different classification tasks. F or some tasks (e.g. HP vs. MP) recov ery and reuse of p oten- tially co-adapted weigh ts might b e feasible while reuse is impaired for other tasks (e.g. HC vs. MC). The partial freezing and fine-tuning strategies appear to b e more consistent across tasks, how ever, the optimal strategy still differs. Overall, our results indicate that the transferability of features not only dep ends on the imaging mo dalit y but also the classification task. This adds to previous insights on transfer learning in the medical domain where the optimal transfer strategy w as found to b e mo dalit y and dataset size dep endent [18]. Comparing the partial 10 Gessert et al. Classifier F reeze 1 F reeze 2 F ull T raining T raining T yp e 0 . 0 0 . 5 1 . 0 1 . 5 2 . 0 2 . 5 3 . 0 3 . 5 4 . 0 4 . 5 5 . 0 5 . 5 6 . 0 6 . 5 7 . 0 7 . 5 8 . 0 8 . 5 9 . 0 9 . 5 10 . 0 10 . 5 11 . 0 11 . 5 12 . 0 12 . 5 13 . 0 13 . 5 T raining time in min utes DenseNet SE-RX Inception V GG16BN Fig. 6: T raining times for 90 ep ochs of all applied architectures for the different training scenarios for the classification task HP vs HC. Note that for training from scratch the same num b er of parameters is trained as for full-fine tuning. Thus, training times are equivalen t for the tw o cases. freezing and fine-tuning strategies, p erformance is very close and there is no op- timal strategy for each of the tasks. How ever, training times are also an asp ect to consider for the differen t transfer learning strategies. As shown in Figure 6, freezing more parameters inside the architecture leads to reduced training times. Th us, partial freezing can b e generally seen as adv antageous as it often achiev es similar p erformance as full tine-tuning while requiring less training time. F or ap- plication, this insight could b e useful when adopting and retraining mo dels f or cancer classification in other organs or when new er arc hitectures are introduced. Differen t architectures for CLM. T o analyze the different transfer strate- gies further, w e consider the R OC curves of each architecture for the HP vs. MP task, see Figure 5. F or this task, using ”off-the-shelf” features and only re- training the classifier p erformed considerably b etter than for the other tasks. As discussed b efore, this indicates that transfer learning scenarios are classification task-dep enden t. In detail, the R OC curv es rev eal that VGG-16 stands out in partic- ular where retraining the classifier only performs b est out of all transfer strategies. In transfer learning researc h, VGG-16 is still a p opular general purp ose feature extractor for n umerous tasks [40, 11]. F or the other arc hitectures, the optimal strat- egy differs. F or example, for Densenet121, the partial freezing metho ds show go o d op erating points in the upp er, left corner of the ROC-curv e. F or Inception-V3 and SE-Resnext50, partial freezing and fine-tuning p erform similar with no clearly su- Deep T ransfer Learning Metho ds for Colon Cancer Classification with CLM 11 p erior metho d. This indicates that the choice of transfer learning metho d dep ends on the architecture. This should b e exp ected, as the mo dels hav e very different blo c k types and each freezing type fixes a different num b er of parameters. The de- tailed results in T able 1 with additional metrics underline this insight. There is no optimal transfer learning strategy and the best p erforming strategy v aries for dif- feren t architectures and classification tasks. Overall, we demonstrate that transfer learning has an impact on p erformance, how ev er, there is no simple rule-of-thum b for optimal transfer learning with CLM. Our results show that examining different freezing strategies can considerably improv e p erformance for individual mo dels. 5 Conclusion W e inv estigate the feasibilit y of colon cancer classification in CLM images using CNNs and multiple transfer learning scenarios. Using in-vivo images of healthy and malignan t colon and p eritoneum tissue obtained from ten sub jects, we adopt four arc hitectures and five transfer learning scenarios for three classification problems with CLM. Our results show that differen t organs as well as health y and malignan t p eritoneum tissue can be classified with deep transfer learning. W e sho w that transfer learning from ImageNet is successful with CLM but the transferability of features is limited. W e find that there is no single optimal mo del or transfer strategy for all CLM classification problems and that task-sp ecific engineering is lik ely required for application. F or future work, our results could b e extended to more classification problems with CLM. Compliance with Ethical Standards F unding: The authors hav e no funding to declare. Conflict of Interest: The authors declare that they hav e no conflict of interest. Ethical Approv al: All pro cedures p erformed in studies inv olving animals were in ac- cordance with the ethical standards of the institution or practice at which the studies were conducted. Informed Consent: Informed consent w as obtained from all individual participants in- cluded in the study . References 1. T orre, L.A., Bray , F., Siegel, R.L., F erlay , J., Lortet-Tieulent, J., Jemal, A. (2015) Global cancer statistics, 2012. CA: a cancer journal for clinicians 65 (2), 87–108 2. V erw aal, V.J., v an Ruth, S., Witk amp, A., Bo ot, H., v an Slo oten, G., Zoetmulder, F.A. (2005) Long-term surviv al of p eritoneal carcinomatosis of colorectal origin. Annals of surgical oncology 12 (1), 65–71 3. F rank o, J., Shi, Q., Goldman, C.D., Pock a j, B.A., Nelson, G.D., Goldb erg, R.M., Pitot, H.C., Grothey , A., Alberts, S.R., Sargent, D.J. (2012) T reatment of colorectal p eritoneal carcinomatosis with systemic chemotherap y: a p ooled analysis of north central cancer treatment group phase iii trials n9741 and n9841. Journal of Clinical Oncology 30 (3), 263 4. de Bree, E., Koops, W., Kr¨ oger, R., v an Ruth, S., Witk amp, A.J., Zoetmulder, F.A. (2004) Peritoneal carcinomatosis from colorectal or appendiceal origin: correlation of preoperative ct with intraoperative findings and ev aluation of interobserv er agreement. Journal of surgical oncology 86 (2), 64–73 12 Gessert et al. 5. Dromain, C., Leb oulleux, S., Aup erin, A., Go ere, D., Malk a, D., Lumbroso, J., Sch um- berger, M., Sigal, R., Elias, D. (2008) Staging of p eritoneal carcinomatosis: enhanced ct vs. p et/ct. Abdominal imaging 33 (1), 87–93 6. Low, R.N., Semelk a, R.C., W oraw attanakul, S., Alzate, G.D. (2000) Extrahepatic ab dom- inal imaging in patients with malignancy: comparison of mr imaging and helical ct in 164 patients. Journal of Magnetic Resonance Imaging: An Official Journal of the International Society for Magnetic Resonance in Medicine 12 (2), 269–277 7. Iafrate, F., Ciolina, M., Sammartino, P ., Baldassari, P ., Rengo, M., Lucchesi, P ., Sibio, S., Accarpio, F., Di Giorgio, A., Laghi, A. (2012) Peritoneal carcinomatosis: imaging with 64-mdct and 3t mri with diffusion-weigh ted imaging. Ab dominal imaging 37 (4), 616–627 8. Gonz´ alez-Moreno, S., Gonz´ alez-Bay´ on, L., Ortega-P ´ erez, G., Gonz´ alez-Hernando, C. (2009) Imaging of p eritoneal carcinomatosis. The Cancer Journal 15 (3), 184–189 9. Ishigami, S., Uenosono, Y., Arigami, T., Y anagita, S., Okumura, H., Uchik ado, Y., Kita, Y., Kurahara, H., Kijima, Y., Nak a jo, A., Maemura, K., Natsugo e, S. (2014) Clinical utility of periop erativ e staging laparoscopy for adv anced gastric cancer. W orld journal of surgical oncology 12 (1), 350 10. Ellebrech t, D.B., Kuemp ers, C., Horn, M., Keck, T., Kleemann, M. (2018) Confo cal laser microscopy as nov el approach for real-time and in-vivo tissue examination during minimal- inv asive surgery in colon cancer. Surgical endoscopy pp. 1–7 11. Litjens, G., Kooi, T., Bejnordi, B.E., Setio, A.A.A., Ciompi, F., Ghafo orian, M., v an der Laak, J.A., V an Ginneken, B., S´ anchez, C.I. (2017) A surv ey on deep learning in medical image analysis. Medical image analysis 42 , 60–88 12. Goceri, E., Goceri, N. (2017) Deep learning in medical image analysis: recen t adv ances and future trends. In: International Conferences Computer Graphics, Visualization, Computer Vision and Image Pro cessing, pp. 305–311 13. Estev a, A., Kuprel, B., Nov oa, R.A., Ko, J., Swetter, S.M., Blau, H.M., Thrun, S. (2017) Dermatologist-level classification of skin cancer with deep neural netw orks. Na- ture 542 (7639), 115 14. Shen, D., W u, G., Suk, H.I. (2017) Deep learning in medical image analysis. Ann ual review of biomedical engineering 19 , 221–248 15. Bengio, Y. (2012) Deep learning of representations for unsupervised and transfer learning. In: Pro ceedings of ICML W orkshop on Unsup ervised and T ransfer Learning, pp. 17–36 16. Hoo-Chang, S., Roth, H.R., Gao, M., Lu, L., Xu, Z., Nogues, I., Y ao, J., Mollura, D., Summers, R.M. (2016) Deep conv olutional neural netw orks for computer-aided detection: Cnn architectures, dataset characteristics and transfer learning. IEEE transactions on medical imaging 35 (5), 1285 17. Gessert, N., Lutz, M., Heyder, M., Latus, S., Leistner, D.M., Abdelwahed, Y.S., Schlae- fer, A. (2019) Automatic plaque detection in ivoct pullbacks using conv olutional neural netw orks. IEEE transactions on medical imaging 38 (2), 426–434 18. T a jbakhsh, N., Shin, J.Y., Gurudu, S.R., Hurst, R.T., Kendall, C.B., Got wa y , M.B., Liang, J. (2016) Conv olutional neural net works for medical image analysis: F ull training or fine tuning? IEEE transactions on medical imaging 35 (5), 1299–1312 19. Ra jadhy aksha, M., Grossman, M., Esterowitz, D., W ebb, R.H., Anderson, R.R. (1995) In vivo confo cal scanning laser microscop y of human skin: melanin provides strong con trast. Journal of Inv estigative Dermatology 104 (6), 946–952 20. Niederer, R. L., Perumal, D., Sherwin, T., McGhee, C.N. (2007) Age-related differences in the normal h uman cornea: a laser scanning in vivo confo cal microscopy study . British Journal of Ophthalmology 21. Aubreville, M., Knipfer, C., Oetter, N., Jaremenk o, C., Rodner, E., Denzler, J., Bohr, C., Neumann, H., Stelzle, F., Maier, A. (2017) Automatic classification of cancerous tissue in laserendomicroscopy images of the oral cavit y using deep learning. Scientific rep orts 7 (1), 11,979 22. Aubreville, M., Stoeve, M., Oetter, N., Goncalves, M., Knipfer, C., Neumann, H., Bohr, C., Stelzle, F., Maier, A. (2018) Deep learning-based detection of motion artifacts in prob e- based confocal laser endomicroscopy images. International journal of computer assisted radiology and surgery pp. 1–12 23. Wiltgen, M., Bloice, M. (2016) Automatic in terpretation of melanocytic images in confocal laser scanning microscopy . In: Microscopy and Analysis. InT ech 24. Hong, J., Park, B.y ., Park, H. (2017) Conv olutional neural netw ork classifier for distin- guishing barrett’s esophagus and neoplasia endomicroscopy images. In: Engineering in Medicine and Biology So ciety (EMBC), 2017 39th Annual In ternational Conference of the IEEE, pp. 2892–2895. IEEE Deep T ransfer Learning Metho ds for Colon Cancer Classification with CLM 13 25. Izadyyazdanabadi, M., Belykh, E., Mo oney , M.A., Esch bacher, J.M., Nak a ji, P ., Y ang, Y., Preul, M.C. (2018) Prosp ects for theranostics in neurosurgical imaging: Emp o wering confocal laser endomicroscop y diagnostics via deep learning. F rontiers in Oncology 8 , 240 26. Izadyyazdanabadi, M., Belykh, E., Martirosy an, N., Esch bacher, J., Nak a ji, P ., Y ang, Y., Preul, M.C. (2017) Improving utilit y of brain tumor confocal laser endomicroscopy: ob jec- tive v alue assessment and diagnostic frame detection with conv olutional neural netw orks. In: Medical Imaging 2017: Computer-Aided Diagnosis, vol. 10134, p. 101342J. Interna- tional So ciet y for Optics and Photonics 27. Izadyyazdanabadi, M., Belykh, E., Cav allo, C., Zhao, X., Gandhi, S., Moreira, L.B., Es- ch bacher, J., Nak a ji, P ., Preul, M.C., Y ang, Y. (2018) W eakly-supervised learning-based feature lo calization for confo cal laser endomicroscopy glioma images. In: International Conference on Medical Image Computing and Computer-Assisted Interv ention, pp. 300– 308. Springer 28. Izadyyazdanabadi, M., Belykh, E., Mo oney , M., Martirosyan, N., Esch bacher, J., Nak a ji, P ., Preul, M.C., Y ang, Y. (2018) Conv olutional neural netw orks: ensem ble modeling, fine- tuning and unsupervised semantic localization for neurosurgical cle images. Journal of Visual Communication and Image Represen tation 54 , 10–20 29. Simony an, K., Zisserman, A. (2014) V ery deep conv olutional net works for large-scale image recognition. arXiv preprint 30. Szegedy , C., V anhouck e, V., Ioffe, S., Shlens, J., W o jna, Z. (2016) Rethinking the inception architecture for computer vision. In: CVPR, pp. 2818–2826 31. Huang, G., Liu, Z., W einberger, K.Q., v an der Maaten, L. (2016) Densely connected con- volutional networks. arXiv preprint 32. Hu, J., Shen, L., Sun, G. (2018) Squeeze-and-excitation netw orks. In: Pro ceedings of the IEEE Conference on Computer Vision and Pattern Recognition, pp. 7132–7141 33. Gessert, N., Wittig, L., Dr¨ omann, D., Keck, T., Schlaefer, A., Ellebrech t, D.B. (2019) F easibility of colon cancer detection in confo cal laser microscopy images using conv olution neural netw orks. In: Bildverarbeitung f ¨ ur die Medizin 2019 34. Ioffe, S., Szegedy , C. (2015) Batch normalization: Accelerating deep net work training by reducing internal co v ariate shift. In: ICML 35. He, K., Zhang, X., Ren, S., Sun, J. (2016) Deep residual learning for image recognition. In: CVPR, pp. 770–778 36. Nair, V., Hinton, G.E. (2010) Rectified linear units improv e restricted boltzmann machines. In: ICML, pp. 807–814 37. Xie, S., Girshick, R., Doll´ ar, P ., T u, Z., He, K. (2017) Aggregated residual transformations for deep neural netw orks. In: 2017 IEEE Conference on Computer Vision and Pattern Recognition (CVPR), pp. 5987–5995. IEEE 38. Shin, H.C., Roth, H.R., Gao, M., Le Lu, Xu, Z., Nogues, I., Y ao, J., Mollura, D., Summers, R.M. (2016) Deep conv olutional neural networks for computer-aided detection: CNN ar- chitectures, dataset characteristics and transfer learning. IEEE T ransactions on Medical Imaging 35 (5), 1285–1298 39. Y osinski, J., Clune, J., Bengio, Y., Lipson, H. (2014) Ho w transferable are features in deep neural netw orks? In: Adv ances in neural information processing systems, pp. 3320–3328 40. Herath, S., Harandi, M., Porikli, F. (2017) Going deep er into action recognition: A surv ey . Image and vision computing 60 , 4–21
Original Paper
Loading high-quality paper...
Comments & Academic Discussion
Loading comments...
Leave a Comment