Functional Segmentation through Dynamic Mode Decomposition: Automatic Quantification of Kidney Function in DCE-MRI Images
Quantification of kidney function in Dynamic Contrast-Enhanced Magnetic Resonance Imaging (DCE-MRI) requires careful segmentation of the renal region of interest (ROI). Traditionally, human experts are required to manually delineate the kidney ROI across multiple images in the dynamic sequence. This approach is costly, time-consuming and labour intensive, and therefore acts to limit patient throughout and acts as one of the factors limiting the wider adoption of DCR-MRI in clinical practice. Therefore, to address this issue, we present the first use of Dynamic Mode Decomposition (DMD) as a basis for automatic segmentation of a dynamic sequence, in this case, kidney ROIs in DCE-MRI. Using DMD coupled combined with thresholding and connected component analysis is first validated on synthetically generated data with known ground-truth, and then applied to ten healthy volunteers’ DCE-MRI datasets. We find that the segmentation result obtained from our proposed DMD framework is comparable to that of expert observers and very significantly better than that of an a-priori bounding box segmentation. Our result gives a mean Jaccard coefficient of 0.87, compared to mean scores of 0.85, 0.88 and 0.87 produced from three independent manual annotations. This represents the first use of DMD as a robust automatic data-driven segmentation approach without requiring any human intervention. This is a viable, efficient alternative approach to current manual methods of isolation of kidney function in DCE-MRI.
💡 Research Summary
Background
Quantifying renal function with dynamic contrast‑enhanced MRI (DCE‑MRI) requires precise delineation of the kidney region of interest (ROI) across all time‑frames. Conventional practice relies on expert manual tracing, which is labor‑intensive, time‑consuming, and prone to inter‑observer variability. Semi‑automatic approaches (k‑means, k‑NN, ICA, active contours) still need user‑defined seeds or templates and often involve complex preprocessing, motion correction, and parameter tuning, limiting their routine clinical adoption.
Objective
The authors propose a fully automatic, data‑driven segmentation pipeline based on Dynamic Mode Decomposition (DMD). By exploiting the intrinsic spatio‑temporal dynamics of the DCE‑MRI sequence, they aim to isolate the kidney ROI without any prior anatomical knowledge, manual intervention, or heuristic thresholds.
Methodology
- Dynamic Mode Decomposition – The DCE‑MRI series (N frames, each of size m × n) is reshaped into a matrix X ∈ ℝ^{mn × N}. A low‑rank approximation of the linear mapping A that advances the system from one frame to the next is obtained via singular value decomposition (SVD). The eigen‑values σ and eigen‑vectors (dynamic modes Ψ) of the reduced operator Ĥ are computed.
- Windowed‑Reconstruction DMD (WR‑DMD) – A preceding step (W‑DMD) separates low‑rank background (C1) from sparse contrast‑enhanced components (C2). The sparse component is fed to the standard DMD to generate a set of modes.
- Mode Selection – The authors examine the logarithm of |σ| and the phase angles of complex eigen‑values. Modes with frequencies near the origin (small growth/decay rates) are considered most significant. They discard one member of each complex‑conjugate pair and sort the remaining modes by ascending phase angle. Empirically, the second mode (mode‑2) consistently highlights the kidney region because it captures the dominant contrast‑agent dynamics within the renal parenchyma.
- Thresholding & Connected Component Analysis – Mode‑2 is normalized, a global intensity threshold is applied, and the resulting binary image is processed to retain the largest connected component, which serves as the kidney template.
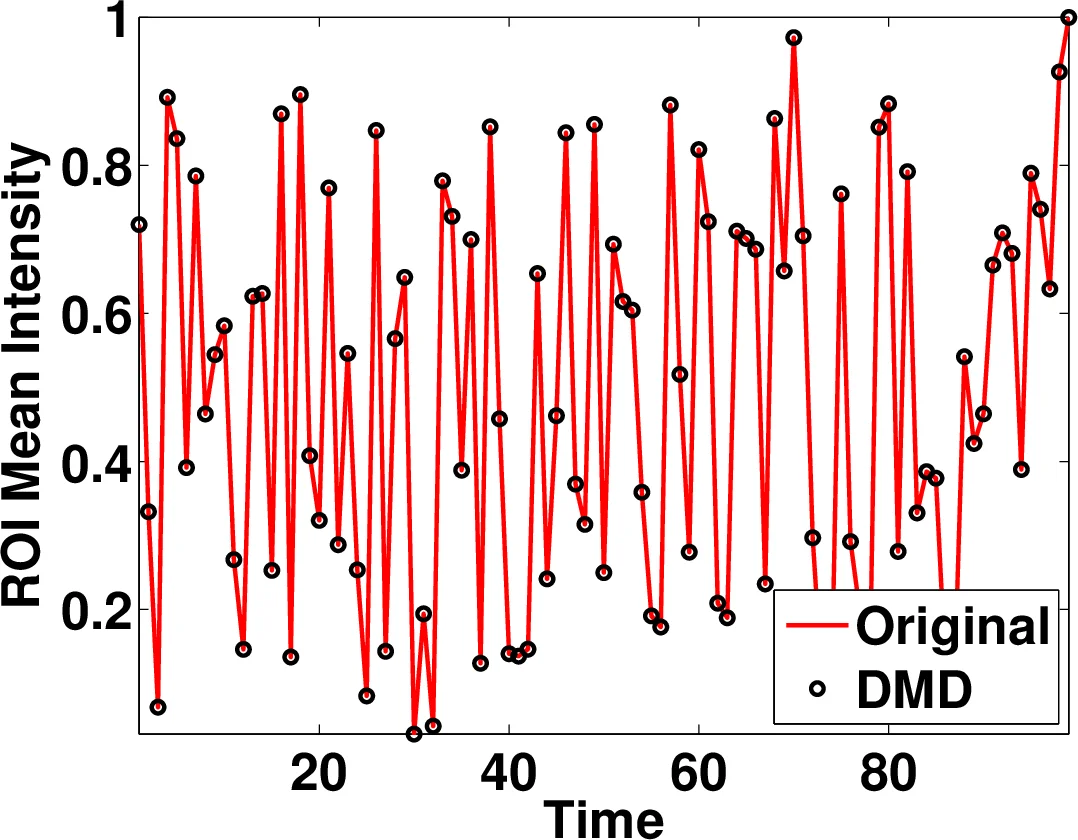
- Quantification – The binary mask is projected back onto each original frame; mean pixel intensity within the mask is computed to derive renal perfusion curves and subsequent kinetic parameters.
Datasets
- Synthetic data with known ground truth was first used to validate the pipeline.
- Clinical data comprised 10 healthy volunteers scanned on a 1.5 T Siemens Avanto system (32‑channel coil), yielding 120 temporal frames per subject covering both kidneys and surrounding anatomy.
Evaluation
Three experienced radiologists manually segmented the kidneys, producing masks e₁, e₂, e₃. For each expert, a consensus ground truth Gᵢ was defined as the intersection of the other two experts’ masks. The Jaccard similarity index J(A,B)=|A∩B|/|A∪B| was computed between each expert’s mask and its Gᵢ (Eᵢ) and between the DMD‑derived mask d and each Gᵢ (Dᵢ). The overall DMD performance D is the average of D₁, D₂, D₃.
Results
- The DMD framework achieved a mean Jaccard index of 0.87 across the three consensus ground truths.
- Expert Jaccard scores were 0.85, 0.88, and 0.87 respectively, indicating that the automatic method matches or slightly exceeds human performance.
- Compared with a naïve a‑priori bounding‑box segmentation (reported as significantly lower, exact numbers not provided), the DMD approach shows a clear advantage.
Discussion
The study demonstrates that DMD can isolate functional kidney tissue purely from temporal contrast dynamics, eliminating the need for anatomical priors or extensive preprocessing. Computationally, the method requires only an SVD of the snapshot matrix (size mn × (N‑1)) and eigen‑decomposition of a small reduced operator, making it feasible on standard CPUs without GPU acceleration.
Limitations include the small sample size (10 healthy subjects) and the exclusive focus on mode‑2; pathological cases with altered perfusion patterns may shift the dominant mode, necessitating adaptive mode selection. The global threshold is fixed across subjects, which could be refined with an adaptive or learning‑based strategy.
Conclusion & Future Work
The authors present the first application of DMD for fully automatic kidney ROI segmentation in DCE‑MRI, achieving expert‑level accuracy and offering a fast, reproducible alternative to manual delineation. Future research should (i) test the pipeline on patients with renal disease, (ii) explore multi‑mode fusion or data‑driven mode selection to handle atypical dynamics, and (iii) integrate the method with downstream kinetic modeling or deep‑learning post‑processing to further improve robustness.
Overall, the work positions DMD as a promising tool for clinical functional imaging pipelines, potentially reducing workload, standardizing measurements, and accelerating the translation of DCE‑MRI into routine nephrology practice.
Comments & Academic Discussion
Loading comments...
Leave a Comment