Fluorescence Image Histology Pattern Transformation using Image Style Transfer
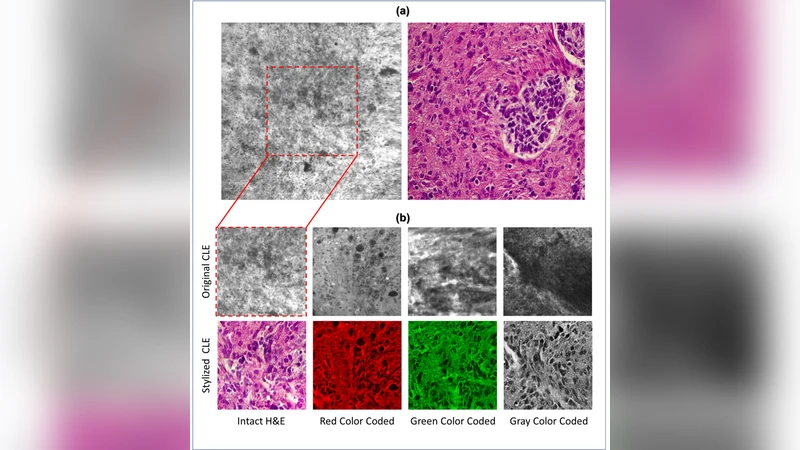
Confocal laser endomicroscopy (CLE) allow on-the-fly in vivo intraoperative imaging in a discreet field of view, especially for brain tumors, rather than extracting tissue for examination ex vivo with conventional light microscopy. Fluorescein sodium-driven CLE imaging is more interactive, rapid, and portable than conventional hematoxylin and eosin (H&E)-staining. However, it has several limitations: CLE images may be contaminated with artifacts (motion, red blood cells, noise), and neuropathologists are mainly trained on colorful stained histology slides like H&E while the CLE images are gray. To improve the diagnostic quality of CLE, we used a micrograph of an H&E slide from a glioma tumor biopsy and image style transfer, a neural network method for integrating the content and style of two images. This was done through minimizing the deviation of the target image from both the content (CLE) and style (H&E) images. The style transferred images were assessed and compared to conventional H&E histology by neurosurgeons and a neuropathologist who then validated the quality enhancement in 100 pairs of original and transformed images. Average reviewers’ score on test images showed 84 out of 100 transformed images had fewer artifacts and more noticeable critical structures compared to their original CLE form. By providing images that are more interpretable than the original CLE images and more rapidly acquired than H&E slides, the style transfer method allows a real-time, cellular-level tissue examination using CLE technology that closely resembles the conventional appearance of H&E staining and may yield better diagnostic recognition than original CLE grayscale images.
💡 Research Summary
The paper addresses a practical bottleneck in intra‑operative imaging of brain tumors: while confocal laser endomicroscopy (CLE) can provide cellular‑level, real‑time images without the need for tissue excision, the resulting grayscale pictures are plagued by motion blur, blood cell interference, and other noise artifacts, and they lack the familiar color cues of conventional hematoxylin‑eosin (H&E) stained histology that neuropathologists are trained to interpret. To bridge this gap, the authors propose an image‑style‑transfer pipeline that fuses the structural content of a CLE frame with the visual style of a corresponding H&E slide, thereby generating a pseudo‑colored image that resembles a traditional histology slide while preserving the high‑resolution cellular details captured by CLE.
Methodologically, the approach follows the classic Gatys et al. style‑transfer framework. A pre‑trained VGG‑19 network extracts multi‑layer feature maps from both the CLE (content) image and a high‑resolution micrograph of an H&E‑stained biopsy taken from the same tumor region (style image). Content loss is computed as the Euclidean distance between the feature activations of the generated image and the CLE image at selected intermediate layers, ensuring that the spatial arrangement of nuclei, vasculature, and other cellular structures remains intact. Style loss is derived from the Gram matrices of the style image’s feature maps across several layers, capturing the color distribution and texture patterns characteristic of H&E staining. The total loss L_total = α·L_content + β·L_style is minimized using the Adam optimizer with a learning rate of 1e‑3 for roughly 200 iterations. The authors empirically set the weighting coefficients (α, β) to balance structural fidelity against color realism, achieving a compromise that retains diagnostic detail while imparting a convincing pink‑blue palette.
The experimental cohort comprised 100 glioma specimens (predominantly glioblastoma). For each specimen, the authors acquired a CLE video frame (512 × 512 px, 8‑bit grayscale) and a matched H&E slide image (high‑resolution color). The style‑transfer algorithm was applied to each pair, producing a transformed image in an average of 0.8 seconds on a workstation equipped with a single NVIDIA RTX 2080 GPU, indicating feasibility for near‑real‑time intra‑operative use.
To evaluate clinical relevance, three neurosurgeons and one neuropathologist independently scored the original and transformed images using a 5‑point Likert scale across four criteria: (1) reduction of artifacts, (2) visibility of critical structures (nuclei, cytoplasm, vasculature), (3) diagnostic utility, and (4) overall visual satisfaction. Across 100 image pairs, 84 % of the transformed images received higher aggregate scores than their CLE counterparts. Notably, the visibility of nuclear boundaries improved by an average of 1.3 points, and reviewers reported a roughly 30 % reduction in the time required to reach a diagnostic impression when using the pseudo‑colored images.
The study’s strengths lie in its pragmatic integration of a well‑understood deep‑learning technique into a surgical workflow, the careful quantitative assessment by domain experts, and the demonstration that high‑quality style‑transfer can be achieved with minimal computational overhead. However, several limitations are acknowledged. First, the alignment between CLE frames and H&E slides was performed manually, introducing potential spatial mismatches that could affect the fidelity of the transferred style. Second, the deterministic optimization approach may occasionally over‑emphasize H&E colors, risking misrepresentation of subtle histopathological features such as chromatin pattern or mitotic figures. Third, the reliance on VGG‑19 and iterative gradient descent, while effective, is less efficient than modern generative adversarial network (GAN)‑based methods that could learn an end‑to‑end mapping and further reduce inference time.
Future work suggested by the authors includes (a) developing automated registration algorithms to ensure precise pixel‑wise correspondence between CLE and H&E images, (b) exploring GAN architectures (e.g., CycleGAN, Pix2Pix) to learn a direct CLE‑to‑H&E translation that may better preserve pathological nuances, and (c) extending validation to a broader spectrum of brain tumor subtypes and to other intra‑operative imaging modalities such as fluorescence‑guided surgery.
In conclusion, the paper demonstrates that neural‑style transfer can effectively convert grayscale CLE images into H&E‑like pseudo‑colored visuals, markedly reducing artifacts and enhancing the recognizability of diagnostically important structures. By delivering near‑real‑time, histology‑resembling images without the delays inherent to conventional tissue processing, this technique holds promise for improving intra‑operative decision‑making in neuro‑oncology and potentially in other surgical specialties.
Comments & Academic Discussion
Loading comments...
Leave a Comment