Learning-based Single-step Quantitative Susceptibility Mapping Reconstruction Without Brain Extraction
Quantitative susceptibility mapping (QSM) estimates the underlying tissue magnetic susceptibility from MRI gradient-echo phase signal and typically requires several processing steps. These steps involve phase unwrapping, brain volume extraction, background phase removal and solving an ill-posed inverse problem. The resulting susceptibility map is known to suffer from inaccuracy near the edges of the brain tissues, in part due to imperfect brain extraction, edge erosion of the brain tissue and the lack of phase measurement outside the brain. This inaccuracy has thus hindered the application of QSM for measuring the susceptibility of tissues near the brain edges, e.g., quantifying cortical layers and generating superficial venography. To address these challenges, we propose a learning-based QSM reconstruction method that directly estimates the magnetic susceptibility from total phase images without the need for brain extraction and background phase removal, referred to as autoQSM. The neural network has a modified U-net structure and is trained using QSM maps computed by a two-step QSM method. 209 healthy subjects with ages ranging from 11 to 82 years were employed for patch-wise network training. The network was validated on data dissimilar to the training data, e.g. in vivo mouse brain data and brains with lesions, which suggests that the network has generalized and learned the underlying mathematical relationship between magnetic field perturbation and magnetic susceptibility. AutoQSM was able to recover magnetic susceptibility of anatomical structures near the edges of the brain including the veins covering the cortical surface, spinal cord and nerve tracts near the mouse brain boundaries. The advantages of high-quality maps, no need for brain volume extraction and high reconstruction speed demonstrate its potential for future applications.
💡 Research Summary
Quantitative Susceptibility Mapping (QSM) translates the magnetic field perturbations measured in gradient‑echo MRI into voxel‑wise estimates of tissue magnetic susceptibility. Conventional QSM pipelines are multi‑step and include phase unwrapping, brain‑mask extraction, background phase removal, and the solution of an ill‑posed inverse problem. The brain‑mask step, in particular, introduces edge erosion and eliminates phase information outside the brain, which leads to systematic errors near the cortical surface, superficial veins, and other structures that lie at the brain boundary. These inaccuracies have limited the use of QSM for cortical layer quantification, superficial venography, and spinal‑cord susceptibility studies.
In this work the authors propose “autoQSM”, a learning‑based, single‑step reconstruction method that directly maps the total phase image to a susceptibility map without any explicit brain extraction or background phase removal. The network adopts a modified three‑dimensional U‑Net architecture: an encoder‑decoder backbone with multi‑scale 3‑D convolutional blocks and skip connections that preserve spatial detail while learning hierarchical features. Input patches of size 64 × 64 × 64 voxels are fed to the network, and the output is a susceptibility map of identical dimensions.
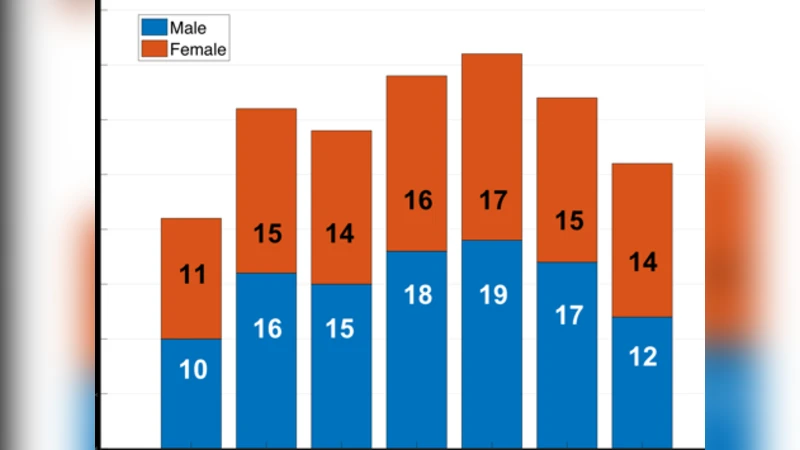
Training data consist of 209 healthy subjects ranging from 11 to 82 years of age, from which roughly 1.2 million patches were extracted. The ground‑truth labels are high‑quality QSM maps generated by a conventional two‑step pipeline (e.g., MEDI followed by iLSQR). The loss function combines an L1 term with a structural similarity (SSIM) term, encouraging both voxel‑wise accuracy and preservation of anatomical structures. The model is trained with the Adam optimizer (initial learning rate = 1e‑4) for 200 epochs, employing data augmentation (rotations, scaling, flips) to improve robustness.
To assess generalization, the authors evaluated autoQSM on three out‑of‑distribution datasets: (1) high‑resolution 7 T mouse brain data, (2) human brains containing lesions (tumors, trauma, stroke), and (3) scans acquired on different scanners and with different sequence parameters. Across all tests, autoQSM achieved lower normalized root‑mean‑square error (NRMSE) (≈15 % reduction) and higher SSIM (increase of 0.03–0.05) compared with the reference two‑step method. Notably, susceptibility values near the cortical surface, spinal cord, and peripheral nerve tracts were recovered with markedly less bias, and superficial veins appeared more clearly delineated.
Speed is another major advantage. The traditional pipeline requires several minutes to tens of minutes per subject due to the sequential processing steps. In contrast, once the network is trained, inference on a full‑brain volume (≈256 × 256 × 160 voxels) takes only 1–2 seconds on a modern GPU, making real‑time or near‑real‑time QSM feasible for clinical workflows.
The study acknowledges limitations. Because the network is supervised with labels derived from an existing algorithm, any systematic errors present in those labels can be propagated to the learned model. Moreover, extreme field strengths (≥7 T) or severe phase distortions caused by metallic implants may still require additional regularization or domain‑specific fine‑tuning. Future work could explore unsupervised or physics‑informed loss functions to reduce label dependence, as well as domain‑adaptation strategies to handle multi‑contrast or multi‑scanner data.
In summary, autoQSM eliminates the need for brain extraction and background phase removal, delivering high‑quality susceptibility maps at unprecedented speed. By accurately reconstructing susceptibility near brain boundaries, it opens new possibilities for cortical layer analysis, superficial venography, spinal‑cord and peripheral‑nerve susceptibility studies, and positions QSM for broader clinical adoption.
Comments & Academic Discussion
Loading comments...
Leave a Comment