Building Brain Invaders: EEG data of an experimental validation
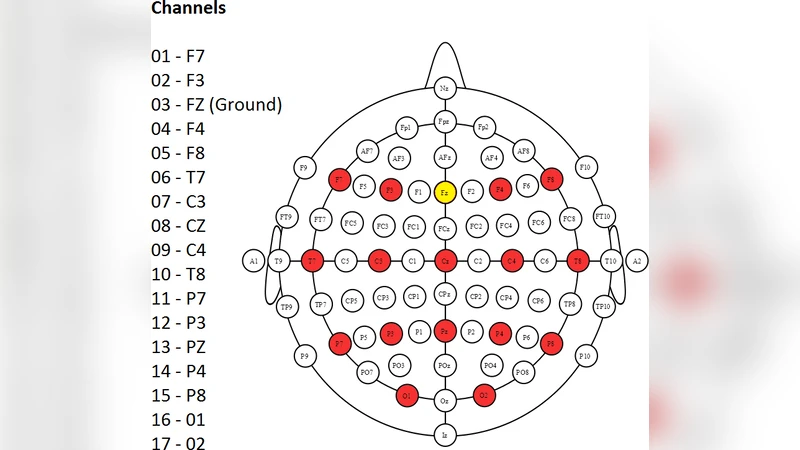
We describe the experimental procedures for a dataset that we have made publicly available at https://doi.org/10.5281/zenodo.2649006 in mat and csv formats. This dataset contains electroencephalographic (EEG) recordings of 25 subjects testing the Brain Invaders (Congedo, 2011), a visual P300 Brain-Computer Interface inspired by the famous vintage video game Space Invaders (Taito, Tokyo, Japan). The visual P300 is an event-related potential elicited by a visual stimulation, peaking 240-600 ms after stimulus onset. EEG data were recorded by 16 electrodes in an experiment that took place in the GIPSA-lab, Grenoble, France, in 2012 (Van Veen, 2013 and Congedo, 2013). Python code for manipulating the data is available at https://github.com/plcrodrigues/py.BI.EEG.2012-GIPSA. The ID of this dataset is BI.EEG.2012-GIPSA.
💡 Research Summary
The paper presents a comprehensive description of a publicly available electroencephalographic (EEG) dataset collected during a visual P300 brain‑computer interface (BCI) experiment called “Brain Invaders.” The dataset, identified as BI.EEG.2012‑GIPSA, comprises recordings from 25 healthy adult participants (both genders, average age ≈27 years) obtained in the GIPSA‑lab in Grenoble, France, in 2012. EEG was recorded with a 16‑channel cap following the standard 10‑20 layout (including Fz, Cz, Pz, etc.) plus two earlobe references, sampled at 250 Hz, and maintained with electrode impedances below 5 kΩ.
The experimental paradigm is a game‑inspired visual oddball task modeled after the classic arcade game Space Invaders. A 6 × 6 grid is displayed on a monitor; each cell normally shows a “alien” icon. Randomly, a “space‑invader” target appears for 100 ms in approximately 10 % of the trials. Participants are instructed to attend to the target and press a button as quickly as possible when it appears. Each stimulus is precisely timestamped, allowing extraction of epochs from –200 ms to +800 ms relative to stimulus onset. Reaction times and accuracy are logged alongside the EEG.
Data are provided in two interchangeable formats: MATLAB *.mat files and plain‑text *.csv files. The *.mat structure contains variables for raw EEG signals, event markers, and a metadata structure that records participant identifiers, session order, electrode locations, sampling parameters, and preprocessing settings. The *.csv files list a time column followed by 16 voltage columns (µV), an event type column, and a response column, making the data readily usable in any programming environment.
To facilitate immediate analysis, the authors release a Python repository (https://github.com/plcrodrigues/py.BI.EEG.2012‑GIPSA) that includes scripts for loading the data, performing baseline correction (−200 ms → 0 ms), epoch extraction, averaging, visualizing ERP waveforms, and training simple classifiers such as Linear Discriminant Analysis (LDA) and Support Vector Machines (SVM). The repository also demonstrates how to apply a 0.1–30 Hz band‑pass filter and Independent Component Analysis (ICA) for ocular and muscular artifact removal, providing a solid baseline pipeline that can be extended with more advanced signal‑processing or deep‑learning techniques.
Technical analysis of the dataset highlights several strengths. First, the game‑like stimulus design increases participant engagement and yields a more naturalistic attentional state compared with traditional flashing‑grid paradigms. Second, the 16‑channel spatial resolution combined with a relatively high sampling rate (250 Hz) supports investigations of source localization, high‑frequency noise characteristics, and time‑frequency analyses that are often limited in lower‑density datasets. Third, the precise event coding and inclusion of behavioral metrics (reaction time, hit/miss) enable multimodal studies linking neural dynamics to performance.
Potential applications are broad. Researchers can develop and benchmark P300‑based spellers, cursor control, or robotic arm interfaces, using the dataset to test classification algorithms, feature‑extraction methods (e.g., CSP, wavelet transforms), and adaptive training strategies. The data also serve as a testbed for artifact‑rejection techniques, transfer‑learning across subjects, and cross‑session stability assessments. From a cognitive‑neuroscience perspective, the dataset permits exploration of visual attention, decision‑making latency, and the neural correlates of target detection in a dynamic, game‑like environment.
By making the raw recordings, detailed metadata, and analysis code openly accessible, the authors address a critical need for reproducibility in BCI research. The dataset can be incorporated into multi‑site meta‑analyses, used for teaching EEG/BCI concepts, and serve as a standard benchmark for future algorithmic developments. In summary, the paper not only documents the experimental protocol and data structure with sufficient granularity for replication but also provides the community with a valuable resource that bridges the gap between laboratory‑grade P300 paradigms and ecologically valid BCI applications.
Comments & Academic Discussion
Loading comments...
Leave a Comment