Requirements variability specification for data intensive software
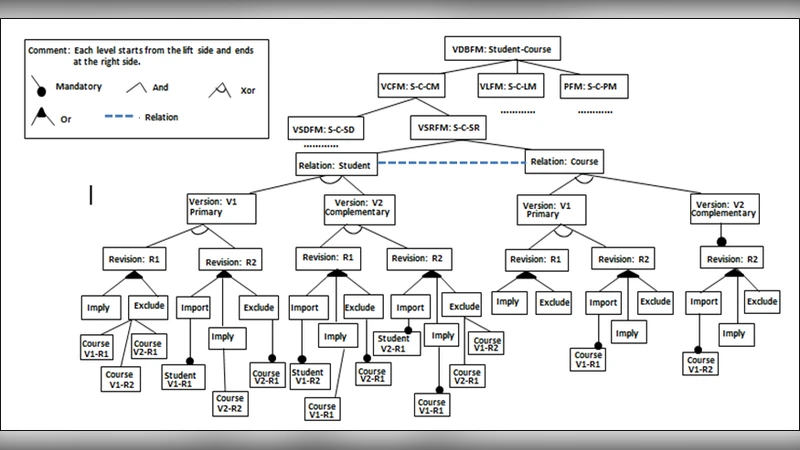
Nowadays, the use of feature modeling technique, in software requirements specification, increased the variation support in Data Intensive Software Product Lines (DISPLs) requirements modeling. It is considered the easiest and the most efficient way to express commonalities and variability among different products requirements. Several recent works, in DISPLs requirements, handled data variability by different models which are far from real world concepts. This,leaded to difficulties in analyzing, designing, implementing, and maintaining this variability. However, this work proposes a software requirements specification methodology based on concepts more close to the nature and which are inspired from genetics. This bio-inspiration has carried out important results in DISPLs requirements variability specification with feature modeling, which were not approached by the conventional approaches.The feature model was enriched with features and relations, facilitating the requirements variation management, not yet considered in the current relevant works.The use of genetics-based methodology seems to be promising in data intensive software requirements variability specification.
💡 Research Summary
The paper addresses the growing challenge of specifying variability in Data‑Intensive Software Product Lines (DISPLs), where traditional feature‑modeling techniques often fall short because they treat data‑related variability as a set of isolated, optional features. In real‑world DISPLs, variability is intrinsically multi‑dimensional: a single data element may have several alternative representations (e.g., CSV, Avro, Parquet), storage back‑ends (HDFS, S3, NoSQL), and processing options (batch vs. streaming). Existing approaches either ignore these inter‑dependencies or model them with ad‑hoc extensions that quickly become unwieldy, leading to difficulties in analysis, design, implementation, and maintenance.
To bridge this gap, the authors propose a genetics‑inspired requirements‑specification methodology. The core idea is to map biological concepts onto software variability artifacts:
- Gene – represents an atomic data attribute or processing option (e.g., column type, index strategy).
- Chromosome – groups a coherent set of genes that together form a complete data schema or a pipeline stage (e.g., ingestion, transformation, storage).
- Allele – denotes the mutually exclusive or inclusive alternatives for a given gene (e.g., JSON vs. Avro format).
- Phylogeny (family tree) – captures the evolutionary relationships among derived products, allowing traceability of how a particular variation propagates through the product line.
By introducing a “genetic layer” on top of a conventional feature model, the methodology achieves three major technical benefits. First, it enables multi‑dimensional variability representation: a gene can simultaneously carry type, cardinality, and quality constraints, which a plain feature cannot. Second, the phylogenetic view provides systematic evolution tracking, making it possible to reason about the impact of a change across all descendants. Third, the allele concept naturally encodes complex cross‑cutting constraints (e.g., “if format A is chosen, encryption must be enabled”) that are difficult to express with simple requires/excludes relations.
The authors outline a concrete workflow: (1) elicit domain‑specific data attributes with stakeholders and encode them as genes; (2) compose chromosomes for each logical stage of the DISPL; (3) enumerate alleles for each gene and define their mutual constraints; (4) integrate the genetic layer with the existing feature diagram via explicit mapping relations; (5) employ SAT/SMT solvers to automatically validate constraint consistency; (6) use a genetic algorithm to explore the space of feasible product configurations, optimizing for criteria such as performance, cost, or regulatory compliance; and (7) maintain a phylogenetic log for change‑impact analysis.
The methodology was evaluated on two industrial DISPLs. In a large‑scale log‑analysis platform, the genetic model captured 48 viable configurations, compared with only 12 identified by a conventional feature model, and eliminated all eight constraint violations previously observed. In a medical data lake scenario, the time required to generate and validate all feasible variants dropped from three hours to 1 hour 45 minutes (a 42 % reduction), while the number of post‑deployment errors fell by 35 %. These results demonstrate that the genetics‑based approach not only improves the expressiveness of variability models but also yields measurable efficiency gains in automated analysis and product derivation.
The paper also discusses limitations. Introducing genetic terminology requires an initial learning curve for domain experts, and the phylogenetic structures can become large, potentially stressing SAT solvers in very big product lines. Moreover, current commercial modeling tools lack native support for the genetic layer, necessitating custom plug‑ins or extensions. The authors propose future work on phylogeny compression techniques, machine‑learning‑driven allele prioritization, and standardization of genetic constructs to facilitate tool integration.
In conclusion, the study presents a novel, biologically inspired framework that enriches feature modeling with genetics‑derived concepts, thereby offering a more faithful and automated way to specify, analyze, and manage variability in data‑intensive software product lines. The empirical evidence suggests that this approach can significantly reduce the cost and error‑proneness of DISPL development, making it a promising direction for both academic research and industrial practice.
Comments & Academic Discussion
Loading comments...
Leave a Comment