Imaging cytometry without image reconstruction (ghost cytometry)
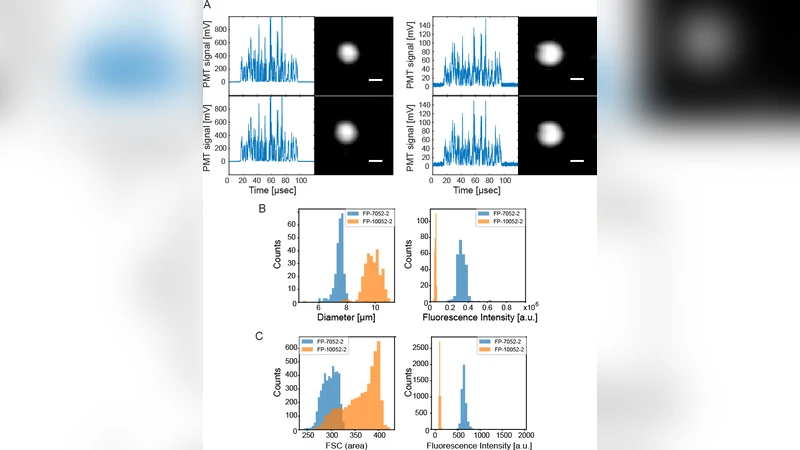
Imaging and analysis of many single cells hold great potential in our understanding of heterogeneous and complex life systems and in enabling biomedical applications. We here introduce a recently realized image-free “imaging” cytometry technology, which we call ghost cytometry. While a compressive ghost imaging technique utilizing object’s motion relative to a projected static light pattern allows recovery of their images, a key of this ghost cytometry is to achieve ultrafast cell classification by directly applying machine learning methods to the compressive imaging signals in a temporal domain. We show the applicability of our method in the analysis of flowing objects based on the reconstructed images as well as in that based on the imaging waveform without image production.
💡 Research Summary
The paper introduces “ghost cytometry,” a novel image‑free flow cytometry platform that leverages compressive sensing and machine‑learning to achieve ultrafast cell classification without reconstructing images. In the system, a static, random light pattern is projected onto a microfluidic channel. As cells flow through the channel, they modulate the transmitted or fluorescent light according to the spatial pattern. A high‑speed photodetector records the resulting intensity as a one‑dimensional temporal waveform. This waveform is a compressed representation of the cell’s interaction with the pattern, containing sufficient information to infer morphology and fluorescence characteristics.
Two analytical pathways are explored. The first follows conventional compressive imaging: the recorded waveform is inverted using L1‑regularized algorithms to reconstruct a two‑dimensional image, which is then fed to a convolutional neural network (CNN) for classification. The second, and the core contribution of ghost cytometry, bypasses reconstruction entirely. The raw waveforms are directly used as input to machine‑learning models (1‑D CNNs, LSTMs, support vector machines, random forests). By training on thousands of labeled waveforms, the models learn to map temporal features to cell types.
Experimental validation involved several biological specimens: human leukocytes, cancer cell lines, and fluorescently labeled bacteria. For each class, >10 000 waveforms were collected at flow rates corresponding to thousands of cells per second. Both reconstruction‑based and direct‑waveform classifiers achieved >95 % accuracy, but the direct approach delivered classification latencies on the order of 1 ms, compared with hundreds of milliseconds for the image‑reconstruction pipeline. Moreover, the waveform‑only method proved more robust to optical noise, pattern drift, and variations in illumination, because it does not amplify reconstruction artifacts.
Key technical considerations include pattern design and detector bandwidth. The authors demonstrate that a binary random matrix generated from a bivariate distribution provides an optimal trade‑off between spatial diversity (information content) and temporal sparsity (signal‑to‑noise ratio). The detector must support >1 GHz sampling and high‑resolution analog‑to‑digital conversion; the implementation used a 12‑bit ADC and FPGA‑based streaming to ensure real‑time data handling.
The significance of ghost cytometry lies in eliminating the computational bottleneck of image reconstruction, thereby enabling true real‑time, high‑throughput cytometry. This opens avenues for large‑scale cell screening, rapid clinical diagnostics, and dynamic monitoring of cellular processes where latency is critical. Future directions proposed include multi‑spectral pattern projection for simultaneous morphological, fluorescence, Raman, and scattering measurements, three‑dimensional flow control to enrich spatial encoding, and integration of on‑chip AI accelerators to perform classification entirely within the instrument. Such extensions would further increase throughput, reduce system cost and footprint, and position ghost cytometry as a disruptive alternative to conventional imaging flow cytometers.
Comments & Academic Discussion
Loading comments...
Leave a Comment