Cross-Modal Data Programming Enables Rapid Medical Machine Learning
Labeling training datasets has become a key barrier to building medical machine learning models. One strategy is to generate training labels programmatically, for example by applying natural language processing pipelines to text reports associated with imaging studies. We propose cross-modal data programming, which generalizes this intuitive strategy in a theoretically-grounded way that enables simpler, clinician-driven input, reduces required labeling time, and improves with additional unlabeled data. In this approach, clinicians generate training labels for models defined over a target modality (e.g. images or time series) by writing rules over an auxiliary modality (e.g. text reports). The resulting technical challenge consists of estimating the accuracies and correlations of these rules; we extend a recent unsupervised generative modeling technique to handle this cross-modal setting in a provably consistent way. Across four applications in radiography, computed tomography, and electroencephalography, and using only several hours of clinician time, our approach matches or exceeds the efficacy of physician-months of hand-labeling with statistical significance, demonstrating a fundamentally faster and more flexible way of building machine learning models in medicine.
💡 Research Summary
The paper tackles a central bottleneck in medical AI: the high cost and inflexibility of manually labeled training data. While modern deep learning architectures are readily available, building clinically useful models still requires massive expert annotation, often amounting to months or years of physician effort. The authors propose “cross‑modal data programming,” a weak‑supervision framework that leverages an auxiliary modality—typically the unstructured text report that accompanies an imaging or monitoring study—to generate training labels for a model that operates solely on the target modality (e.g., X‑ray images, CT volumes, EEG time series).
In practice, clinicians write a small set of labeling functions (LFs) that take a report as input and either emit a binary label (e.g., “abnormal”) or abstain. These LFs can be simple keyword matches, regular‑expression patterns, ontology look‑ups, or even calls to existing classifiers. When applied to a large pool of unlabeled report‑image pairs, the LFs produce a noisy label matrix Λ, where entries may overlap, conflict, or be correlated. To combine these noisy signals, the authors extend the Snorkel generative‑model approach: they define an exponential‑family model pθ(Y,Λ) and estimate the parameters θ̂ by minimizing the negative log marginal likelihood over Λ (Equation 1). This step automatically learns each LF’s accuracy and pairwise correlations, even when the LFs are imperfect or dependent.
The learned generative model yields a probabilistic label ŷ(i)=P(Y=1|Λi;θ̂) for each sample. These confidence‑weighted labels are then used to train a discriminative deep network on the target modality, employing a noise‑aware loss that accounts for label uncertainty (Equation 2). Theoretically, under mild assumptions—most LFs are better than random and a sufficient number of LF pairs are approximately independent—the estimation error of θ̂ and the test error of the discriminative model both decay as O(n⁻¹ᐟ²), where n is the number of unlabeled examples. Consequently, as more unlabeled data become available, performance improves at the same asymptotic rate as adding hand‑labeled examples.
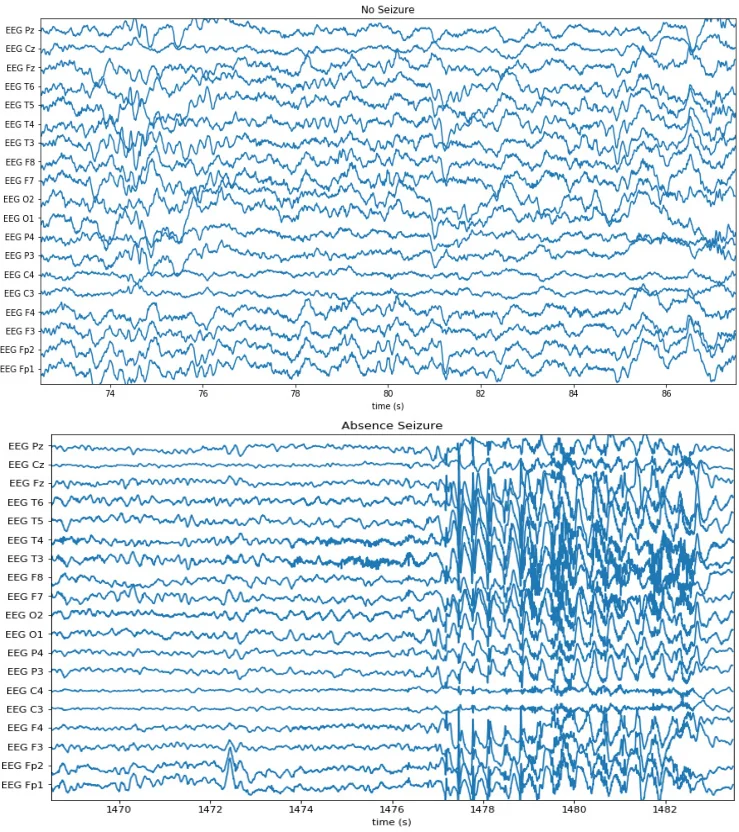
The authors evaluate the framework on four real‑world clinical tasks: (1) chest‑radiograph triage, (2) knee‑radiograph series triage, (3) intracranial hemorrhage detection on head CT, and (4) seizure‑onset detection on EEG. For each task they assembled a large fully‑labeled dataset representing physician‑months or physician‑years of effort, and a separate development set of a few hundred examples used only for LF tuning. Clinicians wrote on average 14 LFs per task (≈6 lines of Python each) in less than eight cumulative hours. Using these LFs and the generative model, the authors produced probabilistic labels for thousands of unlabeled studies and trained standard deep architectures (e.g., ResNet‑18 with attention for CT, CNNs for X‑ray, and temporal CNNs for EEG).
Results show that cross‑modal data programming consistently matches or exceeds the performance of models trained on the massive hand‑labeled baselines. On average, the weak‑supervision approach improves the area under the ROC curve (AUC) by 8.5 points relative to physician‑months of labeling and comes within 3.75 AUC points of the physician‑years baseline, with statistical significance in every application. Moreover, when the amount of unlabeled data is increased (2×, 4×), AUC continues to rise, confirming the predicted O(n⁻¹ᐟ²) scaling.
Key advantages of the method are: (i) drastic reduction in expert labeling time, (ii) intuitive LF authoring that leverages clinicians’ natural expertise in interpreting reports, and (iii) seamless scalability with abundant unlabeled data. Limitations include the need for sufficiently expressive and approximately independent LFs; complex multi‑label or continuous outcomes may require more sophisticated LF design, and violations of the independence assumption could degrade label quality. Future work may explore automated LF synthesis, extensions to multi‑label settings, and applying LFs directly to the target modality.
In summary, cross‑modal data programming offers a theoretically grounded, practically efficient pipeline that transforms textual reports into high‑quality supervision for medical imaging and signal models. By requiring only a few hours of clinician effort yet achieving performance comparable to extensive manual annotation, it paves the way for faster, more flexible deployment of AI systems across diverse medical domains.
Comments & Academic Discussion
Loading comments...
Leave a Comment