Muscle Activity Analysis using Higher-Order Tensor Models: Application to Muscle Synergy Identification
Higher-order tensor decompositions have hardly been used in muscle activity analysis despite multichannel electromyography (EMG) datasets naturally occurring as multi-way structures. Here, we seek to demonstrate and discuss the potential of tensor de…
Authors: Ahmed Ebied, Eli Kinney-lang, Loukianos Spyrou
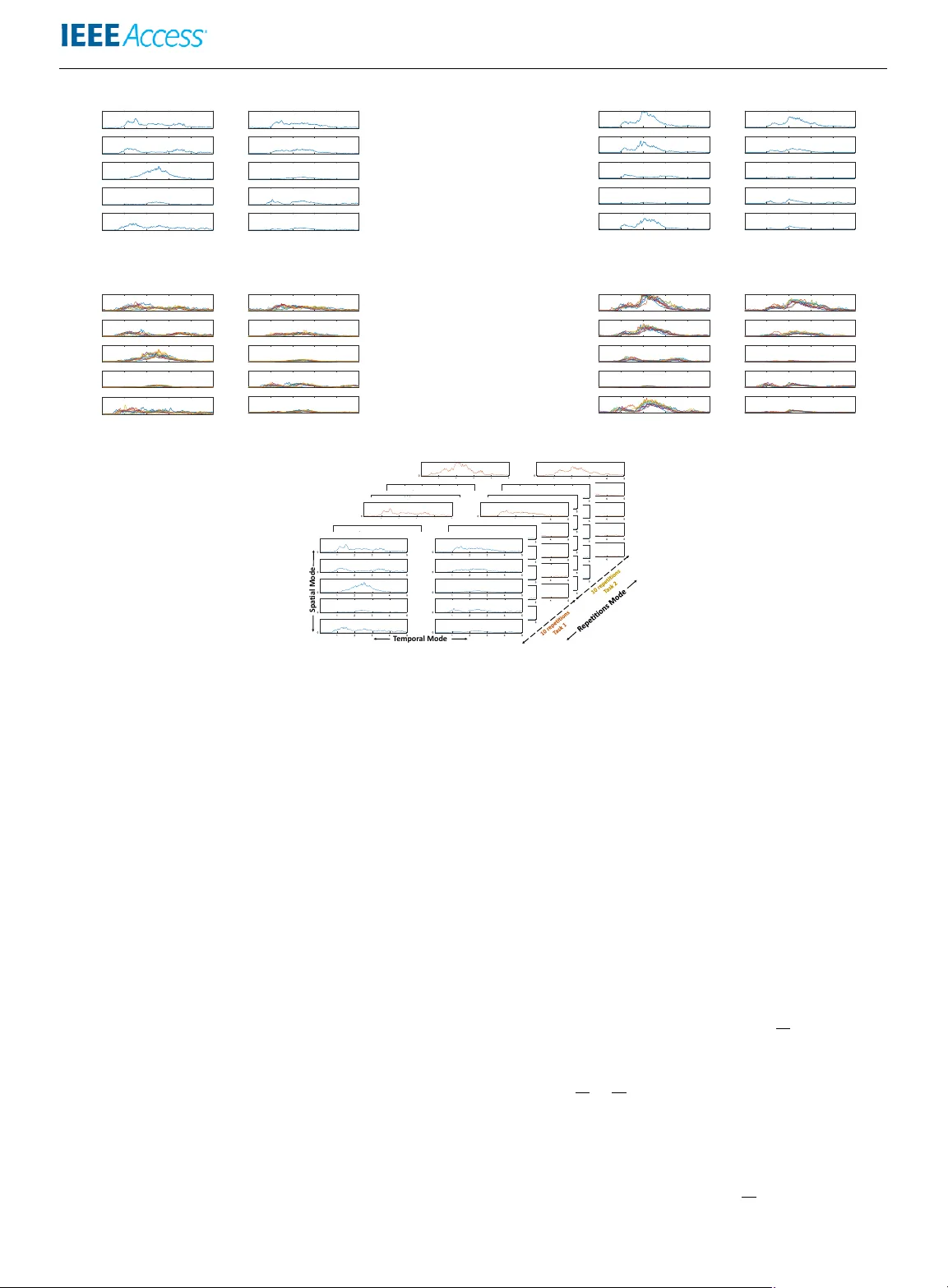
Received December 3, 2018, accepted F ebruar y 5, 2019, date of publication F ebruar y 28, 2019, date of current version March 13, 2019. Digital Object Identifier 10.1109/ACCESS.2019.2902122 Musc le Activity Anal ysis using Higher -Or der T ensor Decomposition: Application to Musc le Synergy Extraction AHMED EBIED 1 , (student member , IEEE), ELI KINNEY-LANG 1 , (student member , IEEE), LOUKIANOS SPYROU 1 , and JA VIER ESCUDERO 1 ,(Member , IEEE) 1 School of Engineering, Institute for Digital Communications, Univ ersity of Edinburgh, Edinb urgh EH9 3FB, United Kingdom Corresponding author: Ahmed Ebied (e-mail: ahmed.ebied@ed.ac.uk). ABSTRA CT Higher-order tensor decompositions hav e hardly been used in muscle activity analysis despite multichannel electromyography (EMG) datasets naturally occurring as multi-way structures. Here, we seek to demonstrate and discuss the potential of tensor decompositions as a framework to estimate muscle synergies from 3 rd -order EMG tensors built by stacking repetitions of multi-channel EMG for se veral tasks. W e compare the two most widespread tensor decomposition models – Parallel Factor Analysis (P ARAF A C) and T ucker – in muscle synergy analysis of the wrist’ s three main Degree of Freedoms (DoFs) using the public first Ninapro database. Furthermore, we proposed a constrained Tuck er decomposition (consTD) method for ef ficient synergy extraction building on the po wer of tensor decompositions. This method is proposed as a direct nov el approach for shared and task-specific synergy estimation from two biomechanically related tasks. Our approach is compared with the current standard approach of repetiti vely applying non-negati ve matrix factorisation (NMF) to a series of the movements. The results show that the consTD method is suitable for syner gy e xtraction compared to P ARAF A C and T ucker . Moreo ver , e xploiting the multi-way structure of muscle activity , the proposed methods successfully identified shared and task- specific synergies for all three DoFs tensors. These were found to be rob ust to disarrangement with regard to task-repetition information, unlike the commonly used NMF. In summary , we demonstrate ho w to use tensors to characterise muscle activity and dev elop a ne w consTD method for muscle synergy e xtraction that could be used for shared and task-specific synergies identification. W e expect that this study will pav e the way for the de velopment of no vel muscle acti vity analysis methods based on higher-order techniques. INDEX TERMS Muscle synergy , NMF , P ARAF AC, Shared synergies, T ask-specific synergies, T ensor Decomposition, T ucker Decomposition. I. INTRODUCTION H IGHER-ORDER tensors are the generalisation of vec- tors ( 1 st -order tensors) and matrices ( 2 nd -order ten- sors). When analysing and extracting patterns from higher- order data (data indexed by more than two variables), tensor decompositions may provide se veral adv antages, such as compactness, uniqueness of decomposition, and generality of the identified components, ov er classical matrix (i.e., 2 nd -order) factorisations [1]. T ucker [2] and Parallel Factor Analysis (P ARAF A C) [3] are the most widespread methods to factorise the tensor into its main components. Recently , higher-order tensor decompositions hav e re- ceiv ed substantial attention in biomedical signal processing applications. For instance, they ha ve been utilised frequently in brain activity analysis [4]. Some applications include analysing electroencephalogram data to classify epileptic patients [5] and analysis of magnetoencephalogram activity in Alzheimer’ s disease [6]. Surprisingly , tensor factorisation had hardly been used in electromyography (EMG) analysis [7]. Recently , we classified wrist movements using the com- ponents of a 4 th -order muscle acti vity tensor to pro vide a proof-of-concept for the use higher -order tensor decompo- sition in muscle synergy analysis [7]. Another study used T ucker decomposition for feature extraction from a 2-channel VOLUME 4, 2016 1 Ebied et al. : Preparation of Papers f or IEEE TRANSACTIONS and JOURNALS EMG for classification [8]. Moreo ver , Delis el al. [9]–[11] proposed a space-by-time decomposition model to extract concurrent spatial and temporal components from single-trail EMG recordings using a Sample-Based Non-neg ativ e Matrix T rifactorization algorithm that resembles a T ucker2 tensor decomposition model [2]. Ho wever , a detailed e valuation of the potential of different tensor factorisation models for EMG analysis is lacking. Multichannel EMG data are most often represented in matrix form with time and channels as indices along each mode (dimension) so that two-way signal processing meth- ods (i.e., matrix factorisations) are used for muscle acti vity analysis. Howe ver , in most EMG studies, data are naturally structured with more modes than the tempor al ( samples ) and spatial ( channel s ) indices. For instance, the EMG datasets usually includes repetitions of subjects and/or move- ments. This means that the muscle activity naturally fits into a higher -order tensor model including additional modes to the tempor al and spatial ones. This is what the studies [7], [8] illustrated and shows that current 2 nd -order approaches do not take adv antage of the natural data structure. This means that some information about the interaction between modes may be lost in those approaches. Thus, we hypothesise that higher-order tensor decomposition will be beneficial for muscle activity analysis. Therefore, our main aim is to explore the use of higher- order tensor decomposition in muscle activity analysis. W e propose a constrained T ucker decomposition (consTD) model for muscle synergy analysis and compare it with the most prominent tensor decomposition models (P ARAF A C and T ucker). Hence, we formulate an appropriate and ef fi- cient approach for consistent and meaningful muscle syner gy extraction. Our secondary objecti ve is to demonstrate the possible application and benefits of tensor decomposition in muscle synergy analysis. Thus, the consTD model is utilised to identify shared and task-specific muscle synergies as an illustration for adv antages of higher -order tensor decomposi- tion in muscle synergy e xtraction. A. MUSCLE SYNERGY EXTRA CTION The muscle synergy concept [12]–[14] pro vides an explana- tion for ho w the central nervous system (CNS) deals with the complexity and high dimensionality of motor control for the musculoskeletal system across multiple Degree of Freedoms (DoFs) [15]. The concept posits that the CNS reduces the motor tasks into a lo wer-dimensional subspace in a modular form. Simply put, the nervous system acti vates muscles in groups (syner gies) for motor control rather than activ ating each muscle individually . Hence, the multichannel EMG signal is considered as a linear mixture of muscle synergies with weighting function or activ ation coef ficients across time as illustrated in Figure 1. Despite the debate about the neural origin of muscle syner- gies [17]–[19], they hav e proved useful for many applications such as clinical research [20], prosthesis control [21], [22], and biomechanical studies [23], [24]. The time-in variant mathematical modelling of muscle syn- ergies [12], [13] expresses the multi-channel EMG as a combination of synchronous synergies scaled by a set of respectiv e weighting functions. This leads to the formula- tion of a blind source separation problem represented as a matrix factorisation where several techniques have been explored, including non-ne gati ve matrix factorisation (NMF) [25], PCA [26] and ICA [27]. Among them, NMF is the most prominent and suitable method [7], [28] since the non- negati vity constraint makes it more appropriate and easily interpretable due to the additiv e nature of synergies [29]. Howe ver , all these approaches are 2 nd -order analysis meth- ods. This may limit them when dealing with situations where repetitiv e analysis are in vestigated such as the identification of shared muscle synergies. B. SHARED SYNERGIES IDENTIFICA TION The notion of shared syner gies deri ves directly from the muscle synergy concept. This implies that shared synergies can be found in diverse motor tasks sharing some mechanical or physical characteristics. Support for this idea comes from animal studies (frogs [30], [31] and cats [32]) as well as hu- man studies where shared and task-specific synergies distinc- tiv e for one motor task or mo vement have been in vestigated across acti vities such as walking and cycling [33], postural balance positions [34], [35], and normal walking and slipping [23], [24]. The current approach to estimate the shared and task- specific syner gies is to apply NMF on the multi-channel EMG signals recorded during the tasks in question. This is done for several repetitions of each task and usually for a number of different subjects. Then, the synergies are rearranged across tasks, repetitions and subjects (in some cases) in order to maximise the similarity between a set of synergies, which is assumed to be shared across tasks and/or subjects. Most of the shared or common synergies identification studies rely only on correlation coef ficients as a similarity metric to dif ferentiate between shared and task- FIGURE 1: A schematic illustration of muscle syner gies. T wo muscle synergies (red and green) and their weighting function generate 5-channel EMG signal as linear combina- tion. The two colours (red and green) in the EMG recording shows ho w each synergy contributes to the wav eform (black line) of each channel. Figure from [16]. 2 VOLUME 4, 2016 Ebied et al. : Preparation of Papers f or IEEE TRANSACTIONS and JOURNALS specific syner gies. Nonetheless, this approach is limited by the fact that the rearrangement of synergies would ha ve a significant effect on identifying shared synergies [23], [24], [33]. Thus, the second objecti ve of this manuscript is to devise the consTD method to take advantage of the multi-way structure in EMG acti vity to e xtract shared muscle synergies. Strictly speaking, the concept of muscle synergy is applicable to 2 nd -order data but we are here inspired by it to extract anal- ogous syner gies via tensor factorisation. The consTD method will be compared against the current traditional method that uses 2 nd -order analysis model (NMF). W e illustrate this with wrist mov ements. II. MA TERIALS In this study , we analyse surf ace EMG datasets from the publicly-av ailable Ninapro first database [36], [37]. The data were collected from 27 healthy subjects instructed to perform 53 wrist, hand and finger movements with 10 repetitions for each mov ement. The “stimulus" time series in the Ninapro dataset is used to set the segment’ s start and end points for each movement repetition. Each segment consists of 10-channel surface EMG signals recorded by a MyoBock 13E200-50 system (Otto Bock HealthCare GmbH), rectified by root mean square and sampled at 100Hz. A selected number of wrist movements are included in this study since we will use upper-limb myoelectric control as an impactful vehicle to demonstrate the techniques. This application of muscle synergy has recei ved substantial atten- tion recently [29], [38]–[40] and we expect that its selection to illustrate our study will promote uptake of the methods. Biomechanically related tasks/mo vements were chosen to identify shared and task-specific synergies between them. Related tasks for this study focused specifically on wrist motion and its three main DoFs. Concerning to the wrist mov ements, 3 DoFs are always considered in myoelectric control radial and ulnar de viation (DoF1) , wrist extension and flexion (DoF2) and finally wrist supination and pronation (DoF3). The three DoFs represent the horizontal, vertical and rotation DoFs or mov ements respectiv ely . III. METHODS A. HIGHER-ORDER TENSOR DECOMPOSITION 1) T ensor construction T ensors are a higher -order generalisation of vectors (1 st - order) and matrices (2 nd -order). The first step to create a higher-order syner gy model is to prepare the data in higher order tensor form. This process of transformation or mapping lower -order data to higher-order data is kno wn as “tensorisa- tion". Se veral stochastic and deterministic techniques hav e been used for tensorisation [41]. “Se gmentation" is one of the deterministic techniques where lower -order tensors are reshaped into higher-order form by segmenting the data into smaller segments and stacking them after each other . T o create 3 rd -order tensor for EMG dataset, the multi- channel EMG recordings of several mov ements and/or tasks can be represented as a matrix with time and channels are its dimensions or modes. This matrix are se gmented into equal epochs where each epoch contains one repetition of one mov ement or task. By stacking these epochs across the ne w r epetition mode, a 3 rd -order tensor is created with modes tempor al × spatial × r epetition . A gi ven wrist’ s DoF tensor is constructed by stacking repetitions of wrist movements. For example, a 1-DoF tensor is created using ulnar (Figure 2c) and radial (Figure 2d) de viation mov ements repetitions to form a 3 rd -order tensor as shown in Figure 2. The 2- DoFs tensor consists of 4 wrist mov ements repetitions; the ulnar and radial de viation in addition to wrist extension and flexion movements. Both tensors are used in the comparison between tensor decomposition models. Howe ver , only the 1- DoF tensors are used for shared and task-specific synergies comparison against NMF for simplicity as we introduce this application as a proof of concept. 2) T ensor decomposition models Higher-order tensors can be decomposed into their main components (also known as factors) in a similar way to matrix factorisation methods. Se veral tensor decomposition models ha ve been introduced with T ucker and P ARAF A C be- ing the most prominent ones [42]. In T ucker decomposition, the higher-order tensor is decomposed into a smaller core tensor transformed by a matrix across each mode, where the core tensor determines the interaction between those matri- ces. On the other hand, P ARAF A C could be considered as a restricted case of T ucker with a supra-diagonal core tensor with 1s across its supra-diagonal and 0s elsewhere, which is also known as an “identity tensor". Consequently , the number of components is fixed for all modes for P ARAF A C, unlike T ucker where the numbers of components in the different modes, can dif fer . Thus, the T ucker model has more flexibility in component number unlik e P ARAF A C [43]. The differences between both models are represented in Figure 3. In general, an n th -order tensor X ∈ R i 1 × i 2 × ....i n can be factorised according to the T ucker model as follows: X ≈ G × 1 B (1) × 2 B (2) · · · × n B ( n ) (1) where G ∈ R j 1 × j 2 ···× j n is the core tensor and B ( n ) ∈ R i n × j n are the component matrices transformed across each mode while “ × n " is multiplication across the n th -mode [43]. The core tensor G is flexible to have dif ferent dimensions across each mode as long as it is smaller than the tensor being decomposed, X , so that j n ≤ i n . On the other hand, the P ARAF AC approach f actorises the n th -order tensor X ∈ R i 1 × i 2 × ...i n into its component matrices with fix ed number of components across each mode as X ≈ Λ × 1 A (1) × 2 A (2) · · · × n A ( n ) (2) where Λ ∈ R r × r ···× r n is a super diagonal tensor that hav e same dimension across each mode and the vector λ is across the diagonal of Λ . This limits the interactions in-between VOLUME 4, 2016 3 Ebied et al. : Preparation of Papers f or IEEE TRANSACTIONS and JOURNALS 1 2 3 4 5 0 1 2 3 4 5 0 1 2 3 4 5 0 1 2 3 4 5 0 1 2 3 4 5 0 Normalised Amplitude 1 2 3 4 5 0 1 2 3 4 5 0 1 2 3 4 5 0 1 2 3 4 5 time(s) 0 1 2 3 4 5 time(s) 0 (a) 1-r epetition of ulnar deviation 1 2 3 4 5 0 1 2 3 4 5 0 1 2 3 4 5 0 1 2 3 4 5 0 1 2 3 4 5 0 Normalised Amplitude 1 2 3 4 5 0 1 2 3 4 5 0 1 2 3 4 5 0 1 2 3 4 5 time(s) 0 1 2 3 4 5 time(s) 0 (b) 1-r epetition of radial deviation 1 2 3 4 5 0 1 2 3 4 5 0 1 2 3 4 5 0 1 2 3 4 5 0 1 2 3 4 5 0 Normalised Amplitude 1 2 3 4 5 0 1 2 3 4 5 0 1 2 3 4 5 0 1 2 3 4 5 time(s) 0 1 2 3 4 5 time(s) 0 (c) 10-r epetition of ulnar deviation 1 2 3 4 5 0 1 2 3 4 5 0 1 2 3 4 5 0 1 2 3 4 5 0 1 2 3 4 5 0 Normalised Amplitude 1 2 3 4 5 0 1 2 3 4 5 0 1 2 3 4 5 0 1 2 3 4 5 time(s) 0 1 2 3 4 5 time(s) 0 (d) 10-r epetition of radial deviation Spa ti al Mo de T em por al Mo de (e) 3 rd -or der tensor for radial and ulnar deviations mo vements with modes [ 500-samples (5-seconds) × 10-channels × 20-r epetitions (10 repetitions for each mo vements) ] FIGURE 2: An example for data construction and tensor decomposition. Figures 2a and 2b are 10-channel EMG recordings of Ninapro first database (subject “1"). The recording is for one repetition of ulnar (2a) and radial deviation (2b) movements (DoF1). (Figures 2c and 2d) shows 10 repetitions of each mo vements, which are stacked together to form a 3 rd -order tensor as in 2e for DoF1. components unlike T ucker decomposition. The Core Consis- tency Diagnostic (CORCONDIA) is an index to assess the appropriateness of P ARAF A C decomposition by measuring the the degree of super-diagonally [44]. The core consistency is less than or equal to 100% where consistency close to 100% implies an appropriate multilinear model, whereas a lower core consistency (lower than 50%) would mean a problematic or ev en in v alid model [44]. The P ARAF AC decomposition is unique under very mild conditions [43]. On the other hand, the T ucker decomposition generally does not provide unique solutions [1]. Ho wev er , the uniqueness for the T ucker model can be achiev ed in practice by imposing additional constraints on the modes [45]. 3) Alter nating Least Squares algorithm Both Tuck er and P ARAF AC can be estimated with the Alternating least square (ALS). ALS starts initialising the components (and core tensor in the case of Tuck er decom- position) to be estimated either randomly [46] or by using other methods such as singular v alue decomposition or direct trilinear decomposition [47]. After initialisation, the next step is the iteration phase to minimise the loss function between the original data and its model by breaking down this complex non-conv ex problem into a series of simpler, con vex problems, which are tackled in succession [48]. This is done by fixing all the component matrices to be estimated except for those corresponding to one of the modes and alternate iterativ ely between all the components to solve each con vex problem until con ver gence [49]. In the case of the 3 rd -order EMG tensor X ∈ R i 1 × i 2 × i 3 , The T ucker model equation 1 would be expressed as X ≈ G × 1 B (1) × 2 B (2) × 3 B (3) (3) where B (1) ∈ R i 1 × j 1 is the temporal mode while B (2) ∈ R i 2 × j 2 and B (3) ∈ R i 3 × j 3 are the spatial and repetition modes respecti vely . j 1 , j 2 and j 3 are the number of compo- nents in each mode and the core tensor G dimensions as well. 4 VOLUME 4, 2016 Ebied et al. : Preparation of Papers f or IEEE TRANSACTIONS and JOURNALS ≈ 𝐗 𝐆 B ( 1 ) 𝑖 1 𝑖 2 𝑖 3 𝑖 1 B ( 2 ) 𝑖 2 B ( 3 ) 𝑖 3 𝑗 1 𝑗 1 𝑗 2 𝑗 2 𝑗 3 𝑗 3 (a) . . . . . . . . . (b) FIGURE 3: Illustration of T ucker (3a) and P ARAF A C (3b) decomposition for 3 rd -order tensor X . The least squares loss function for this model would be ar g min B (1) , B (2) , B (3) , G k X ( i 1 × i 2 i 3 ) − B ( 1 ) G ( B ( 3 ) ⊗ B ( 2 ) ) T k 2 (4) where ⊗ is a Kronecker product, a generalisation of the outer product, that can be applied on matrices of arbitrary size [50]. The algorithms in this study are based on adapted P ARAF AC and T ucker functions from the (N-W A Y T oolbox) for Matlab [51]. The ALS algorithm has se veral advantages such as sim- plicity compared to the simultaneous approaches. Howe ver , it is not guaranteed to conv erge to a stationary point as the problem could have sev eral local minima. Therefore, mul- tiple constraints on initialisation and iteration phases would help to improv e the estimation [43]. Moreover , constraining the tensor models has sev eral benefits including: improving the uniqueness of the solution, more interpretable results that do not contradict a priori kno wledge, av oiding degeneracy and numerical problems, and speeding up the algorithm. Although constraints may lead to poorer fit for the data com- pared to the unconstrained model, the adv antages outweigh the decrease in the fit for most cases [1]. The decomposi- tion models are constrained through their ALS algorithm in the initialisation and/or iteration phases. For example, non- negati vity constraint is one of the most commonly used ones due to the illogical meaning for negati ve components in many cases. The non-negati vity constraint is implemented in the updating step by setting the negati ve values of computed components to zero by the end of each iteration to force the algorithm to conv erge into a non-negati ve solution. In the case of muscle synergy extraction, non-ne gativity w ould add more information to the decomposition by taking in account the additiv e nature of muscle synergies. B. CONSTRAINED TUCKER DECOMPOSITION In this section, the consTD method is discussed in detail, including number of components and dif ferent constraints imposed to improv e muscle synergy extraction. Imposing constraints on the tensor decomposition model has se veral benefits [1] as discussed in Section III-A3. There- fore, in order to extract consistent and meaningful muscle synergies, a consTD model is proposed for muscle synergy analysis. W e hypothesise that this model would benefit from the flexibility and v ersatility of T ucker model in compari- son with P ARAF AC decomposition while retaining the high explained variance. In addition, the additional constraints would result a unique and consistent synergy e xtraction. This approach was inspired by the shared-synergy concept [30], [31] by including additional components to account for an y shared variability across mo vements, tasks or DoFs. 1) Number of components In this setup, one tempor al component is assigned for each DoF instead of two components as preliminary results showed that when assigning 2 components for one DoF , the temporal activity would be segmented as the following: one component will capture the main acti vity (middle of the segment) will the other will capture the rest period (at the beginning and end of the segments). Therefore, the addi- tional temporal component will not be beneficial for synergy extraction since we are concerned with extracting the main muscle activity to identify shared synergy for each DoF. In addition, we are aiming to reduce the number of elements in the decomposition focusing on the main activity . The number of spatial mode components (syner gies) were chosen according to the functional approach (discussed in Section III-C1). In general, we aim for a parsimonious model that could extract meaningful muscle syner gies with the least number of components and elements. Hence, two compo- nents are assigned to each DoF to estimate a task-specific synergy for each movement for both spatial and r epetition modes. In addition, an additional component (shared) was assigned in these two modes in order to improve the data fit and account for any common variability inspired by the shared synergy concept. 2) Additional constraints Four constraints on the T ucker model were proposed to facilitate the muscle synergy identification. T wo constraints are imposed in the initialisation phase on the core tensor and r epetition mode components, while the other tw o are applied during the iteration phase of ALS algorithm. VOLUME 4, 2016 5 Ebied et al. : Preparation of Papers f or IEEE TRANSACTIONS and JOURNALS The core tensor is initialised to link each of the com- ponents in the tempor al and r epetition modes into their respectiv e spatial components (synergy). The core tensor is initialised and fix ed (does not update with each iteration) into a v alue of 1 between each spatial synergy and its respective components in the other modes and 0 otherwise. This ensures that ev ery spatial synergy is assigned to only one r epetition component and av oids any cross interaction. The v alues of the core tensor are chosen to be 1 to account for all the variability in the mode components. This fixed sparse core tensor setup controls the interactions between components in each mode and links the repetition components to its respectiv e components it tempor al and spatial modes. In addition, since we know that each repetition in the 3 rd -order tensor belongs to a known movement, we use this information in tensor decomposition by constraining the r epetition mode. The components of the r epetition mode are initialised and divided into “task-specific" and “shared" components. The task-specific components are initialised to 1 for a repetition of the considered mov ement or task and 0 otherwise, while the shared component is initialised by a value of 0 . 5 for all repetitions. Unlike the core tensor the update of this mode is not fixed to account for the variability and differences between repetitions of the same mov ement, alternativ ely , a controlled a veraging constraint is used during the iteration phase. The controlled a veraging and repetition mode initialisation works together to incorporate the repeti- tions information into the tensor decomposition and help it to identify the shared and task-specific synergies separately . The other two constraints on updating components in T ucker’ s ALS algorithm are the non-negati vity on tempor al and spatial modes and the controlled averaging on the initialised r epetition mode. The non-neg ativity constraint is imposed in order to ha ve meaningful components (synergies) [7], [29] as discussed in Section III-A3. The controlled av eraging constraint aims to allo w some variability within each r epetition component whether it is shared or task-specific. This approach will hold the structure of r epetition factors that was initialised without fixing it through iterations and take the differences between repeti- tions into account. As for controlled a veraging implemen- tation, it is a simple moving-a veraging filter with windo w length ( k = 3 ) implemented by modifying the iteration phase in the ALS algorithm. It is applied on r epetition mode components at the end of each iteration to account for the variability between each repetition, hence, increase the e xplained v ariance. W e ackno wledge this is a simple implementation but we expect it to be representative of our procedure while achie ving a good performance in muscle synergy identification. C. TENSOR DECOMPOSITION MODELS FOR MUSCLE SYNERGY ANAL YSIS 1) Number of Synergies Selecting the appropriate number of components (including synergies in spatial mode) for higher-order tensor models is instrumental in capturing the underlying structure of the data [49]. Se veral mathematical approaches ha ve been de- ployed to determine the appropriate number of components for higher-order tensor decomposition such as CORCONDIA [44], heuristic and approximating [52] techniques. On the other hand, the number of synergies for 2 nd - order model extracted via matrix factorisation methods have been determined using two main approaches: a functional approach, and a mathematical approach [53]. The functional approach relies on prior knowledge of data structure and myoelectric control requirements to choose the appropriate number of synergies. For instance, one [54] or two [55] syn- ergies are assigned to each wrist’ s DoF for proportional my- oelcetric control. The mathematical approaches are similar to methods traditionally used in higher-order tensor models. It relies mathematical computation such as explained variance or the likelihood criteria [56]. In order to choose the number of components for P ARAF A C and T ucker models, the prior knowledge of the data structure ( i.e., number of movements) had been utilised as in the functional approach of matrix factorisation. In addi- tion, the mathematical criteria (CORCONDIA and e xplained variance) was used to test and compare different number of components. Both the 1-DoF and 2-DoFs tensors were de- composed using P ARAF AC with a set number of components (2, 3 and 4 components) since we aim to estimate at least one synergy for each mov ement and the number of mov ements are 2 and 4 in the 1-DoF and 2-DoFs tensors respectiv ely . This helped to guarantee each mov ement was identified by at least one muscle synergy . Similarly , [2,2,2], [3,3,3] and [4,4,4] T ucker models were used to decompose both tensors. In order to compare both models (Tuck er and P ARAF A C), number of components for the Tuck er model was the same in all modes to match the P ARAF A C model for comprehensive comparison. 2) T uck er and P ARAF AC models f or synergy e xtraction In order to examine the use of T ucker decomposition for muscle synergy extraction, both tensors (1-DoF and 2-DoFs) were decomposed with a non-negati ve T ucker decomposi- tion. Three models were applied on the 3 rd -order tensors with different number of components; [2,2,2], [3,3,3] and [4,4,4]. The time for algorithm execution is recorded for ev ery run across the 27 subjects as well as the explained v ariance percentage as a metrics to compare tensor decomposition models. On the other hand, to highlight the differences between T ucker and P ARAF AC models in muscle acti vity analysis, a 2-, 3- and 4-component P ARAF A C models with non- negati vity constraints are applied on the same wrist’ s tensors. The execution time for each run as well as CORCONDIA were recorded to compare between tensor decomposition for synergy analysis. The number of components for both methods were chosen according to the criteria discussed in Section III-C1. 6 VOLUME 4, 2016 Ebied et al. : Preparation of Papers f or IEEE TRANSACTIONS and JOURNALS 3) consTD for synergy e xtraction The proposed consTD method is applied to the 1- and 2- DoFs tensors for muscle synergy analysis. The consTD aims to extract one syner gy for each mov ement (task-specific syn- ergy) in addition to a shared synergy across all movements. The number of components for consTD were [1 , 3 , 3] for 1- DoF tensors and [2 , 5 , 5] for 2-DoFs tensors according to the criteria discussed in Section III-B1. Therefore, two components in the spatial and r epetition modes were assigned to each DoF in addition to an addi- tional “shared" component in these two modes. While one tempor al component is assigned for each DoF since we do not need to segment main temporal activity . Thus, a [1 , 3 , 3] consTD is developed to estimate interpretable components from 1-DoF tensors, while a [2 , 5 , 5] model was used for the 2-DoFs tensors. T ABLE 1: Core tensor intialisation for consTD models. [1 , 3 , 3] [2 , 5 , 5] g 1 ,n,n = 1 n ∈ { 1 , 2 , 3 } g 1 ,n,n = 1 n ∈ { 1 , 2 , 5 } g 2 ,n,n = 1 n ∈ { 3 , 4 , 5 } g = 0 otherw ise g = 0 otherw ise The core tensor is initialised and fixed for both consTD models accordingly as sho wn in T able 1. The repetition mode is initialised as discussed in Section III-B where the task-specific components are initialised by 1 for a repetitions of the considered mo vement and 0 otherwise and the shared component is initialised by a value of 0 . 5 for all repetitions. The repetition mode is constrained in the iteration phase through controlled averaging while the non-negativity con- straint is imposed on the tempor al and spatial modes. 4) Experimental settings In order to compare the proposed constrained T ucker model for muscle synergy analysis with non-negati ve T ucker and P ARAF A C models. The three algorithms were run 10 times in order to examine the uniqueness of solution by testing the ability of algorithms to con ver ge to the same point with similar resulting components. For each run, the time of exe- cution and e xplained v ariance were recorded for both T ucker decomposition models while CORCONDIA and e xecution times were recorded for P ARAF A C. All decomposition mod- els are performed using Matlab 9 with Intel core i7 processor (2.4 GHz, 12 GB RAM). D . SHARED SYNERGY IDENTIFICA TION The NMF approach to identify the shared synergies mainly depends on similarity metrics such as correlation coef ficient or coefficient of determination R 2 . Synergies are estimated from repetitions of single tasks either by taking the average EMG acti vity then applying NMF [23], [33], or by a veraging the synergies extracted from each repetition [35]. The result would be group of syner gies for each task. This is followed by computing correlation coef ficients between synergies of different tasks and identifying shared synergies by matching the synergies of highest correlation coef ficient. On the other hand, the tensor approach stacks the repeti- tions from different tasks together to form a 3 rd -order tensor with modes channels × time × repetitions as shown in Figure 2. Constrained tensor factorisation is applied on this 3 rd -order tensor to e xtract the shared syner gies as well as the task specific ones without the need of any similarity metric, unlike NMF approach. The consTD directly identifies shared synergy and 2 task-specific syner gies in the spatial mode for 3 rd -order tensor with repetitions of 2 tasks (ulnar and radial deviation, for e xample). 1) NMF as benchmark NMF [25] is used in this study as a comparati ve benchmark for shared synergy identification [23], [30], [33], [57]. NMF processes the multi-channel EMG recording as a matrix X with dimensions channel × time . NMF decomposes EMG recordings into two smaller matrices (factors). The first factor holds the temporal information (also known as weighting function) B (1) while the other is the muscle synergy holding the spatial information B (2) as X ≈ B (1) × B (2) T (5) where both B (1) and B (2) are constrained to be non-ne gati ve. For details see [58]. Since the dataset had 10 repetitions for each task, NMF was applied on each of them. The number of synergies was chosen by v ariance accounted for (V AF) as a metric [21]. The first step to identify the shared and task-specific synergies would be finding the reference synergy [23], [33] from the 10 repetitions of that task. This is done by calculating the inter-correlation between the 10 repetitions. Since number of synergies are tw o, 200 correlation processes are needed to identify the reference repetition which is achie ves the highest av erage correlation coefficient between repetitions. The second step is to use this reference to arrange syner- gies within each repetition [33]. Finally , the arranged syn- ergies are a veraged to compute the first and second mean synergies for the task. Then, to identify the shared synergy of one DoF, the mentioned method is applied on the two tasks forming the DoF in question, the correlation coefficients between the resulting four mean syner gies (two for each task) are calculated so that the highly correlated synergies between the two tasks are identified as shared, while the other tw o are considered as task-specific [23], [35], [57]. 2) Comparison between shared synergies identified by consTD and NMF W e compared shared and task-specific synergies identified using the constrained T ucker tensor decomposition method with those identified by using the traditional NMF and cor- relation method. This comparison is held since there is no ground truth about the shared and task-specific synergies. Therefore, for each wrist’ s DoF, three synergies are identified by T ucker (two task-specific and one shared synergy) while VOLUME 4, 2016 7 Ebied et al. : Preparation of Papers f or IEEE TRANSACTIONS and JOURNALS T ABLE 2: Median explained variance and e xecution time for the non-negati ve T ucker decomposition of the 27 Subjects. 1-DoF T ensor 2-DoFs T ensor No. of Components Explained V ariance time(s) Explained V ariance time(s) [2,2,2] 87.8% 12.5 77.5% 24.9 [3,3,3] 92.2% 25.7 86.4% 59.7 [4,4,4] 94.3% 73 89.8% 75.2 four mean synergies (tw o for each task in the DoF) are estimated using NMF. The correlation coef ficient between T ucker and NMF synergies are calculated and averaged across all 27 subjects. The comparison is held between the main three wrist’ s DoFs: ulnar and radial deviation (DoF1); wrist extension/flexion (DoF2); and wrist supina- tion/pronation (DoF3). 3) V alidation with randomised repetitions In order to provide further validation to the approach of shared synergy identification using consTD, the r epetition mode in the 3 rd -order tensor of each DoF was randomly shuffled to destroy any task-repetition information. The same consTD algorithm is applied on the tensor to identify the shared syner gy between the two tasks. The 2 task-specific synergies will be corrupted since information about the tasks are missing. Howe ver , this experiment tests the ability of constrained Tuck er method to identify the shared syner gies without any data arrangement which cannot be achiev ed us- ing the traditional NMF and correlation method. The shared synergies identified from the shuf fled 3 rd -order tensors is compared against the shared synergies estimated from un- corrupted ones by calculating the correlation coeffi cients between them. The comparison is done using 15 shuffled tensors for each DoF of the main 3 wrist’ s DoF and the av erage correlated is computed. IV . RESUL TS A. TENSOR DECOMPOSITION FOR MUSCLE SYNERGY ANAL YSIS 1) T uck er and P ARAF AC models f or synergy e xtraction Three non-ne gati ve T ucker decomposition models with [2 , 2 , 2] , [3 , 3 , 3] and [4 , 4 , 4] components were applied on 1- and 2-DoFs tensors for muscle synergy extraction. An example of the 10 runs of the [3 , 3 , 3] T ucker decomposition for 1-DoF tensor is shown in Figure 4a. The explained variance and the algorithm execution time were recorded for each decomposition and the median v alues across the 27 subjects are summarised in T able 2. The P ARAF AC decomposition model was applied on both 1- and 2-DoFs tensors of wrist horizontal and vertical DoFs. The number of components explored were 2, 3 and 4 where a non-negati vity constraint was applied on all components. An example of 3-component P ARAF AC decomposition on T ABLE 3: The median core consistency and time of ex ecu- tion for the P ARAF A C decomposition across the 27 subjects. 1-DoF T ensor 2-DoFs T ensor No. of Components Core consistency time(s) Core consistency time(s) 2 95.2% 0.39 91.1% 0.58 3 30% 0.60 64.6% 0.72 4 6% 0.91 29.3% 1.13 T ABLE 4: The median explained variance and time for ex ecution for the consTD across the 27 Subjects. 1-DoF T ensor 2-DoF T ensor No. of Components [1,3,3] [2,5,5] Explained V ariance 78.28% 73.21% time(s) 0.26 0.65 2-DoFs tensor is shown in Figure 4b. The time of execution for P ARAF A C algorithm as well as CORCONDIA were recorded across the 27 subjects are summarised in T able 3. 2) Constrained T uc ker decomposition The consTD models were applied on 1- and 2-DoFs tensors for muscle syner gy estimation for 10 runs across the 27 subjects. The 1-DoF tensor was decomposed using [1 , 3 , 3] constrained T ucker method while the 2-DoFs tensor was decomposed using [2 , 5 , 5] constrained T ucker model. An example of [1 , 3 , 3] constrained T ucker method for T ensor shown in Figure 2e is illustrated in Figure 5. Explained variance and ex ecution time were recorded and the median values are sho wn in T able 4. B. SHARED SYNERGY IDENTIFICA TION 1) NMF synergies A number of wrist tasks were selected and 10-channel EMG recording was decomposed using NMF to extract two syn- ergies for each task. Our analysis found that two synergies could account for over 90% of the variability in data for all repetitions. For each task, NMF was applied on each of the 10 repetitions and the estimated syner gies were rearranged using mutual correlation coefficients then av eraged across repetitions to result two muscle synergies for each movement. An example of the av eraged syner gies are shown in Figure 6 for the ulnar and radial deviation mo vements (DoF1) of subject “1". Shared synergies are determined through correlation. As shown in Figure 6, the second synergy of ulnar de viation (Fig. 6a) is highly correlated with the first component of radial deviation (Fig. 6b) with r = 0 . 91 . Therefore, accord- ing to the standard NMF approach the average of these two synergies is considered as a shared syner gy between the ulnar and radial de viation tasks while the remaining synergies are task-specific. 8 VOLUME 4, 2016 Ebied et al. : Preparation of Papers f or IEEE TRANSACTIONS and JOURNALS 1 2 3 4 5 Seconds -0.5 0 0.5 Normalised Amplitude Temporal Mode Component 1 Component 2 Component 3 -0.5 0 0.5 Spatial Mode Ch 1 Ch 2 Ch 3 Ch 4 Ch 5 Ch 6 Ch 7 Ch 8 Ch 9 Ch 10 2 4 6 8 10 12 14 16 18 20 Repetitions -0.5 0 0.5 Repetition Mode (a) The averag e and standar d deviation for 10 runs of non-ne gative [3 , 3 , 3] T uck er decomposition for 1-DoF tensor . 1 2 3 4 5 Seconds -0.5 0 0.5 Normalised Amplitude Temporal Mode Component 1 Component 2 Component 3 -0.5 0 0.5 Spatial Mode Ch 1 Ch 2 Ch 3 Ch 4 Ch 5 Ch 6 Ch 7 Ch 8 Ch 9 Ch 10 2 4 6 8 10 12 14 16 18 20 Repetitions -0.5 0 0.5 Repetition Mode (b) The averag e and standar d deviation for 10 runs of non-ne gative 3 -components P ARAF AC decomposition for 1-DoF tensor . FIGURE 4: The a verage (solid line) and standard deviations (shaded areas) for 10 runs of non-negativ e T ucker (4a) and P ARAF A C (4b) applied on the 3 rd -order tensor . Because of the uniqueness of P ARAF A C solution, its standard deviation is zero as shown in P anel 4b. While only one component (blue) in T ucker seems to be unique in the tempor al and repetition mode as shown in P anel 4a. 2) Shared synergies comparison Synergies extracted from consTD are compared against NMF synergies to test the ability of this method to identify shared and task-specific synergies. The correlation coef ficients be- tween Tuck er and NMF synergies are used as a metric. This was done for 3 pairs of tasks (DoFs) for the wrist mov ements across the 27 subjects in the dataset, where the 3 synergies from Tuck er decomposition are compared against each a veraged NMF syner gies of each task. F or example, the correlation coef ficients between the estimated consTD syner - gies and NMF synergies of ulnar and radial deviation (DoF) for the 27 subjects are represented in Figure 7. The a verage correlations for all 3 DoFs are summarised in T able 5. The third (shared) synergy is highly correlated with both tasks as the av erage correlation coefficients for ulnar and radial deviation are 0 . 819 and 0 . 857 , respecti vely . Each of the other 2 task-specific synergies are correlated with its respectiv e task. F or ulnar de viation, the first synergy has cor - relation coefficient of 0 . 778 compared to 0 . 575 for the second synergy , while for the radial de viation the second syner gy has an av erage correlation coef ficient of 0 . 887 compared to 0 . 232 with the first synergy . Similar results are found with other movements such as wrist e xtension/flexion and wrist supination/pronation) as shown in T able 5. VOLUME 4, 2016 9 Ebied et al. : Preparation of Papers f or IEEE TRANSACTIONS and JOURNALS 1 2 3 4 5 Seconds -1 0 1 Normalised Amplitude Temporal Mode Component 1 Component 2 Component 3 -0.5 0 0.5 Spatial Mode Ch1 Ch2 Ch3 Ch4 Ch5 Ch6 Ch7 Ch8 Ch9 Ch10 2 4 6 8 10 12 14 16 18 20 Repetitions -0.5 0 0.5 Repetition Mode FIGURE 5: Constrained [1 , 3 , 3] Tuck er decomposition for the 3 rd -order tensor in Fig. (2e). The spatial mode had 3 components, the first 2 components are muscle syner gies specified for the ulnar and radial de viation movements while the third synergy represents the shared syner gy between them. T ABLE 5: A verage correlation coefficients between T ucker and NMF synergies for the 3 Main DoFs of wrist. Synergy 1 Synergy 2 Synergy 3 DOF 1 Ulnar deviation 0.778 0.575 0.819 Radial deviation 0.232 0.887 0.857 Wrist extension 0.729 0.337 0.868 DOF 2 Wrist flexion 0.408 0.776 0.880 DOF 3 Wrist supination 0.911 0.481 0.879 Wrist pronation 0.104 0.920 0.792 3) V alidation with randomised repetitions In order to validate the approach of shared synergy identifica- tion and to show that it is robust to an y repetitions disarrange- ment, the 3 rd -order tensor in Figure 2e was randomly shuf- fled across the repetition mode to destroy the task-repetition information. The consTD was applied on the randomly shuf- fled tensor to identify shared synergy as shown in Figure 8. In comparison with the normal tensor decomposition (Fig. 5), we noticed that the task-specific components were dif ferent as e xpected since the information was destroyed. On the other hand, the shared synergy in the spatial mode were very similar . The av erage correlation coefficients between shared synergies identified from 15 shuffled tensors and from arranged ones were found to be 0 . 89 . This sho ws the ability of the algorithm to identify the shared syner gy despite the corruption in the task-repetition information during the tensor construction. V . DISCUSSION W e proposed the use of higher-order tensors and their de- compositions as a frame work for muscle syner gy analysis. Although it may not fully agree with the classical definition of muscle synergies, it is inspired by it to provide more synergistic information from the data. This is moti vated by the fact that most of EMG datasets are naturally in multi-way form with different repetitions from tasks and/or subjects. In addition, some synergy analysis techniques [9], [10], [59] extracted spatial and temporal components of muscle acti vity using a space-by-time decomposition model that resembles T ucker2 tensor decomposition model. Surprisingly , a fully dev eloped tensor factorisation has barely been widely used in EMG analysis [7]. The main objectiv e was to explore the use of tensor fac- torisation models in muscle syner gy analysis. W e proposed a consTD model inspired by the shared synergy concept and compare it with the most prominent methods (P ARAF A C and T ucker decomposition models). This was approached by applying these methods with dif ferent number of components on 3 rd − order tensors consisting of 20 and 40 multichannel EMG repetitions for one and two wrist’ s DoFs respectively (10 repetitions each). The three tensor decomposition meth- ods were assessed according to the algorithm e xecution time, explained v ariance and CORCONDIA, as appropriate. How- ev er , we ackno wledge that ex ecution time is for compari- son reasons only , not for assessing the whole performance. The constraint T ucker decomposition model was the best approach to e xtract muscle synergies from 3 rd -order tensor as it was capable to estimate unique synergies in a short ex ecution time with acceptable explained v ariance. 10 VOLUME 4, 2016 Ebied et al. : Preparation of Papers f or IEEE TRANSACTIONS and JOURNALS Synergy 1 Synergy 2 0 1 Normalised Amplitude Ch1 Ch2 Ch3 Ch4 Ch5 Ch6 Ch7 Ch8 Ch9 Ch10 (a) Synergy 1 Synergy 2 0 1 Normalised Amplitude Ch1 Ch2 Ch3 Ch4 Ch5 Ch6 Ch7 Ch8 Ch9 Ch10 (b) FIGURE 6: two-component NMF for the ulnar (6a) and radial (6b) deviations mov ements of subject “1" averaged across 10 repetitions for each task. The second component of ulnar de viation is highly correlated with the first compo- nent of radial de viation suggesting that these are the shared synergies between those tasks. The secondary objectiv e was to illustrate the potential use of tensor decomposition for shared and task-specific syner- gies identification. The proposed consTD method was used to identify shared synergies between each pair of tasks that forms a main DoF of wrist movements. The resulting syner- gies were compared against the standard NMF factorisation approach for the 3 wrist DoFs across 27 subjects. The results showed that shared and task-specific synergies estimated via the consTD method were highly correlated with those iden- tified through traditional NMF approach. In addition, tensor shared synergies were compared to randomly shuf fled tensor without an y task-repetition information for further v alidation and the proposed algorithm was able to estimate nearly the same shared syner gy as the ordered tensor , thus sho wing robustness to disarrangement. A. TENSOR DECOMPOSITION MODELS FOR MUSCLE SYNERGY ANAL YSIS The comparison between Tuck er , P ARAF A C and consTD models for muscle synergy analysis showed that T ucker decomposition can provide a good fit for the data as shown by the high explained variance percentage in T able 2. How- ev er , the estimated synergies via non-ne gativ e T ucker de- composition were inconsistent as shown in Figure 4a. This inconsistency arises from the f act that T ucker decomposition generally does not provide unique solutions [1], and unique- Synergy 1 Synergy 2 Shared Synergy -1 -0.5 0 0.5 1 Correlation Coefficient 0.778 0.575 0.819 (a) Synergy 1 Synergy 2 Shared Synergy -1 -0.5 0 0.5 1 Correlation Coefficient 0.232 0.857 0.887 (b) FIGURE 7: V isualisation of six histograms for the correla- tion coefficients between syner gies extracted by consTD and NMF synergies across the 27 subjects for ulnar (Panel 7a) and radial (Panel 7b) deviation (DoF1). Each line represents frequency of occurrence for the histogram, where darker shades refer to higher frequency of occurrence. The red crosses each histogram are its mean v alue. The full mean value comparison across all DoFs is represented in T able 5. ness can be achiev ed in practice by imposing additional constraints [45]. Since the decomposition is unconstrained, the initialisation (which is set randomly) changes in each run. This would result in different components in each run since there are no constraints to achie ve uniqueness of the solution and the model con verges to dif ferent local minima. This also explain the longer ex ecution time since the unconstrained T ucker algorithm took more iterations to conv erge takes more time to con verge as represented in T able 2. Despite the increase of explained v ariance percentage with the additional number of components, the e xecution time increased as well, since the algorithm could not con ver ge easily . On the other hand, P ARAF A C was significantly faster as seen in T able 3, since it con ver ged to the same local minima most of the time due to its extreme constrained nature. P ARAF A C with non-neg ativity constraints was capable of estimating estimate muscle synergies from the 1-DoF tensor as sho wn in Figure 4b. Ho wever , P ARAF A C could not deal with 2-DoFs tensors or higher number of synergies as the de- composition deviates from the trilinear model and P ARAF A C could not hold. This is illustrated with low CORCONDIA as shown in T able 3. In addition, synergy estimation is affected by inflexibility of the P ARAF A C model as the number of components are fixed across modes. Therefore, the informa- VOLUME 4, 2016 11 Ebied et al. : Preparation of Papers f or IEEE TRANSACTIONS and JOURNALS 1 2 3 4 5 Seconds -1 0 1 Normalised Amplitude Temporal Mode Component 1 Component 2 Component 3 -0.5 0 0.5 Spatial Mode Ch1 Ch2 Ch3 Ch4 Ch5 Ch6 Ch7 Ch8 Ch9 Ch10 2 4 6 8 10 12 14 16 18 20 Repetitions -0.5 0 0.5 Repetition Mode FIGURE 8: consTD for the DoF1 tensor in Figure (2e) but with shuffled r epetition mode. The algorithm was able to identify the same shared synergy (third spatial component) as the decomposition of the regular tensor (Fig. 5) ev en without any task- repetition information. tion of each task in the tempor al mode is segmented between components. T wo consTD models ( [1 , 3 , 3] and [2 , 5 , 5] ) were proposed to decompose the 1-DoF and 2-DoFs tensors respectively . They were able to achiev e over 70% explained v ariance and decrease the e xecution time by about 10-fold compared to the non-negati ve T ucker model as shown in T able 4. Moreov er , the resulting synergies were consistent ov er the runs as shown in Figure 5 unlik e T ucker model. The consTD approach allocates one synergy for each mo vement and addi- tional “‘shared" synergy to account for v ariability inspired by the shared synergy concept. This additional shared synergy improv ed the explained variance compared to [1 , 2 , 2] con- strained T ucker model where the median explained variance was 59 . 3% in the preliminary results. Moreov er , the total number of components in consTD may be greater than or equal to the number of components in traditional T ucker decomposition. Ho wever , the total number of elements for consTD that need to be estimated is sig- nificantly less than T ucker model. For example, the 1-DoF tensor ( 500 sample × 10 channel s × 20 r epetitions ) with number of elements= 100 , 000 is decomposed via [2 , 2 , 2] T ucker decomposition into 1060 elements in addition to 8 elements in the core tensor . On the other hand, a [1 , 3 , 3] consTD can decompose the same tensor into 590 elements in addition to 9 elements in its sparse core tensor . Hence, the proposed consTD for synergy e xtraction is more ef ficient in comparison to unconstrained T ucker model. T wo important variants for the T ucker decomposition models are worth noting, T ucker1 and T ucker2, which can be seen as a special case of Tuck er model where only one and two modes are estimated respecti vely . In both models, the additional modes are set to be identity matrices and absorbed into the core tensor [60]. As a result, T ucker1 model is equiv alent to the ordinary two-dimensional PCA, while T ucker2 is a model of intermediate comple xity as compared with the T ucker1 and the standard T ucker model [43]. In T ucker2, the third mode is absorbed to the core tensor and the model explains the v ariability of the first two modes only . Hence, Delis el al. [9] approach focused on spatial and tempor al components and their interactions, while the consTD method adds r epetition mode to spatial and temporal modes to incorporate different movements and the shared synergies between them to provide a 3 rd -order EMG tensor decomposition for muscle synergy analysis. Hence, we conclude that the proposed consTD method is the best solution to obtain unique and interpretable syner - gies from 3 rd -order tensor decomposition. It was capable to achiev e this with smallest number of synergies and elements because of the utilisation of shared syner gy concept. The non-negati vity constraint is essential because of the additiv e nature of syner gies. Moreover , the fixed core tensor is piv otal, as we can directly relate synergies (in the spatial mode) to other specific components in tempor al and r epetition modes. This is in contrary with the unconstrained core tensor in Tuck er model, which allows for interactions between all components in each mode. Due to these interactions, it becomes difficult to achie ve a unique of solution for T ucker decomposition. This increases the computational time dra- matically . 12 VOLUME 4, 2016 Ebied et al. : Preparation of Papers f or IEEE TRANSACTIONS and JOURNALS B. SHARED MUSCLE SYNERGY IDENTIFICA TION W e sho wed that higher-order tensor decomposition models can achieve direct identification of shared synergies without relying on any similarity metric such as correlation coef- ficient. This is in contrast with the current approaches for shared synergy estimation [23], [24], [33], [34], [57] which apply NMF repetitively on multi-channel EMG recordings of different repetitions and then rely on maximising the correlation coef ficients between the estimated syner gies with regard to a reference one. Then, shared and task-specific synergies are identified through the correlation coefficient threshold. This was illustrated in Figure 5 of consTD model, where component 1 and 2 in the spatial mode are task- specific for the two task forming the tensor while component 3 is the shared synergy between them. Synergies identified via consTD were compared against synergies e xtracted using NMF for the 27 subjects. In spite of the potential drawbacks of NMF shared synergies, we used NMF as benchmark since there is no ground truth for shared synergies to compare both methods against. In addition, the wrist movement included in the study are limited since shared synergies are easier to identify . Only two NMF synergies could e xplain ov er 90% of variance. Hence, the errors of disarrangement is minimised. This was done for 3 pairs of tasks (DoFs) for the wrist mov ements (T able 5). The shared synergies identified by T ucker (third synergy in Figure 5) were highly correlated with both tasks while each of the other two tasks correlated with one task as a task-specific synergy . This highlights the ability of consTD to identify task-specific and shared synergies directly from the multi-way datasets. Further v alidation were held by applying the consTD on randomly shuffled tensor without any task- repetition information. The proposed algorithm was able to estimate nearly the same shared synergy as the ordered tensor (Fig. 8), which indicated robustness of the method. In addition, the standard NMF approaches for shared and task-specific synergies identification are vulnerable to errors and biases since they depend on the particular arrangement of the data, the choice of the reference synergy , and the correlation coefficient threshold. This is not the case for consTD approach where it w as able to identify the shared synergy ev en with a shuffled tensor as shown in Figure 8. In addition, it is a more direct and f aster alternative since there is no need to apply repetitiv e NMF and correlation. C. APPLICA TIONS, LIMIT A TIONS AND FUTURE W ORK Usually , the shared and task-specific synergies are identified in a study of complex multi-joint mov ements such as gait and posture analysis [23], [24], [33]. Howe ver , we choose simple wrist mov ements for few reasons. Firstly , this is a first study to sho w how higher-order tensors could be beneficial for muscle synergy analysis. Secondly , wrist movements are simple tasks that could be described by two synergies as mentioned before. Therefore, shared syner gies can be iden- tified easily with minimum disarrangement errors for good comparison and validation for our proposed tensor approach. Finally , we are interested in upper -limb myoelectric control and looking to the shared synergy concept as an inspiration for proportional myoelectric control based on muscle syner- gies in the future. Moreov er , the main aim for this is study is to highlight the potential of higher -order tensor model for muscle acti v- ity analysis especially extracting muscle synergies. Hence, consTD could be extended to various applications by conv ert- ing the information we ha ve into the right set of constraints. For e xample, this approach could be extended to estimate the shared synergies across subjects to explore the subject- specific syner gies [35]. In addition, and in relation to the point abo ve, dif ferent set of constraints could help to dev elop a myoelectric control based on muscle synergies as in [61]. VI. CONCLUSION In conclusion, we introduced tensor decomposition models (P ARAF A C and T ucker) for muscle synergy extraction and compared their use in EMG analysis to e xtract meaningful muscle synergies with a proposed consTD model. The de- veloped method was the best approach for muscle synergy estimation by pro viding unique and interpretable synergies with high explained v ariance and short execution time. The proposed consTD model can be used to identify shared and task-specific syner gies. The results were compared against the standard NMF approach using data from the publicly av ailable Ninapro dataset. The consTD method was more suitable to the multi-way nature of the datasets without relying on symmetry metrics or synergies arrangements. Fur - thermore, it pro vided more direct and data-driv en estimations of the syner gies in comparison with NMF-based approaches, making our approach more rob ust to disarrangement of rep- etitions and the loss of task-repetition information. Thus, we expect that this study will pave the way for the development of muscle activity processing and analysis methods based on higher-order techniques. REFERENCES [1] A. Cichocki, D. Mandic, A. H. Phan, C. Caiafa, G. Zhou, Q. Zhao, and L. De Lathauwer , “T ensor Decompositions for Signal Processing Applications From T wo-way to Multiway Component Analysis, ” IEEE Signal Processing Magazine, vol. 32, pp. 1–23, Mar . 2014. [2] L. R. T ucker , “Some mathematical notes on three-mode factor analysis, ” Psychometrika, vol. 31, pp. 279–311, Sept. 1966. [3] R. a. Harshman, “Foundations of the P ARAF A C procedure: Models and conditions for an “explanatory” multimodal factor analysis, ” UCLA W ork- ing Papers in Phonetics, v ol. 16, no. 10, pp. 1–84, 1970. [4] F . Cong, Q.-H. Lin, L.-D. K uang, X.-F . Gong, P . Astikainen, and T . Ris- taniemi, “T ensor decomposition of EEG signals: A brief review , ” Journal of Neuroscience Methods, vol. 248, pp. 59–69, June 2015. [5] L. Sp yrou, S. Kouchaki, and S. Sanei, “Multiview Classification and Dimensionality Reduction of Scalp and Intracranial EEG Data through T ensor Factorisation, ” Aug. 2016. [6] J. Escudero, E. Acar, A. Fernández, and R. Bro, “Multiscale entropy analysis of resting-state magnetoencephalogram with tensor f actorisations in Alzheimer’ s disease, ” Brain Research Bulletin, vol. 119, pp. 136–144, 2015. [7] A. Ebied, L. Spyrou, E. Kinne y-Lang, and J. Escudero, “On the use of higher-order tensors to model muscle synergies, ” in 2017 39 th Annual In- ternational Conference of the IEEE Engineering in Medicine and Biology Society (EMBC), pp. 1792–1795, IEEE, July 2017. VOLUME 4, 2016 13 Ebied et al. : Preparation of Papers f or IEEE TRANSACTIONS and JOURNALS [8] P . Xie and Y . Song, “Multi-domain feature extraction from surface EMG signals using nonnegativ e tensor factorization, ” in 2013 IEEE International Conference on Bioinformatics and Biomedicine, pp. 322–325, IEEE, Dec. 2013. [9] I. Delis, S. Panzeri, T . Pozzo, and B. Berret, “A unifying model of concurrent spatial and temporal modularity in muscle activity , ” Journal of Neurophysiology , vol. 111, no. 3, pp. 675–693, 2014. [10] I. Delis, S. Panzeri, T . Pozzo, and B. Berret, “T ask-discriminativ e space- by-time f actorization of muscle acti vity , ” Frontiers in Human Neuro- science, vol. 9, p. 399, July 2015. [11] P . M. Hilt, I. Delis, T . Pozzo, and B. Berret, “Space-by-Time Modular Decomposition Effectiv ely Describes Whole-Body Muscle Activity Dur- ing Upright Reaching in V arious Directions, ” Frontiers in Computational Neuroscience, vol. 12, p. 20, Apr . 2018. [12] M. C. T resch, P . Saltiel, and E. Bizzi, “The construction of movement by the spinal cord., ” Nature neuroscience, vol. 2, pp. 162–7, Feb . 1999. [13] P . Saltiel, K. W yler-Duda, A. D’A vella, M. C. T resch, and E. Bizzi, “Muscle synergies encoded within the spinal cord: evidence from focal intraspinal NMDA iontophoresis in the frog., ” Journal of neuroph ysiology , vol. 85, pp. 605–619, Feb . 2001. [14] A. d’A vella, P . Saltiel, and E. Bizzi, “Combinations of muscle syner gies in the construction of a natural motor behavior ., ” Nature neuroscience, v ol. 6, pp. 300–308, Mar . 2003. [15] A. d’A vella, M. Giese, Y . P . Ivanenko, T . Schack, and T . Flash, “Editorial: Modularity in motor control: from muscle syner gies to cognitive action representation., ” Frontiers in computational neuroscience, vol. 9, p. 126, Jan. 2015. [16] V . C.-K. K. Cheung, A. Turolla, M. Agostini, S. Silv oni, C. Bennis, P . Kasi, S. Paganoni, P . Bonato, and E. Bizzi, “Muscle synergy patterns as physiological markers of motor cortical damage, ” Proceedings of the National Academy of Sciences, vol. 109, pp. 14652–14656, Sept. 2012. [17] J. J. Kutch and F . J. V alero-Cuev as, “Challenges and ne w approaches to proving the existence of muscle synergies of neural origin., ” PLoS computational biology , vol. 8, p. e1002434, Jan. 2012. [18] E. Bizzi and V . C.-K. K. Cheung, “The neural origin of muscle synergies., ” Frontiers in computational neuroscience, vol. 7, p. 51, Feb . 2013. [19] M. C. Tresch and A. Jarc, “The case for and against muscle synergies, ” Current Opinion in Neurobiology, vol. 19, pp. 601–607, Dec. 2009. [20] D. T orricelli, F . Barroso, M. Coscia, C. Alessandro, F . Lunardini, E. Bravo Esteban, and A. D’A vella, “Muscle Synergies in Clinical Prac- tice: Theoretical and Practical Implications, ” in Emer ging Therapies in Neurorehabilitation II (J. L. Pons, R. Raya, and J. González, eds.), vol. 10 of Biosystems & Biorobotics, pp. 251–272, Cham: Springer International Publishing, 2016. [21] G. Rasool, K. Iqbal, N. Bouaynaya, and G. White, “Real-T ime T ask Discrimination for Myoelectric Control Employing T ask-Specific Muscle Synergies., ” IEEE transactions on neural systems and rehabilitation engi- neering, vol. 24, pp. 98–108, Jan. 2016. [22] M. Ison and P . Artemiadis, “Proportional Myoelectric Control of Robots: Muscle Syner gy Dev elopment Dri ves Performance Enhancement, Re- tainment, and Generalization, ” IEEE T ransactions on Robotics, v ol. 31, pp. 259–268, Apr . 2015. [23] M. M. Nazifi, H. U. Y oon, K. Beschorner , and P . Hur , “Shared and T ask-Specific Muscle Synergies during Normal W alking and Slipping, ” Frontiers in Human Neuroscience, vol. 11, pp. 1–14, Feb . 2017. [24] G. Martino, Y . P . Ivanenko, A. D’A vella, M. Serrao, A. Ranav olo, F . Draicchio, G. Cappellini, C. Casali, and F . Lacquaniti, “Neuromuscular adjustments of gait associated with unstable conditions, ” Journal of Neu- rophysiology , vol. 114, p. jn.00029.2015, Sept. 2015. [25] D. D. Lee and H. S. Seung, “Learning the parts of objects by non-ne gativ e matrix factorization., ” Nature, vol. 401, pp. 788–91, Oct. 1999. [26] J. E. Jackson, A User’s Guide to Principal Components. W iley Series in Probability and Statistics, Hoboken, NJ, USA: John Wiley & Sons, Inc., Mar . 1991. [27] A. Hyvärinen and E. Oja, “Independent component analysis: algorithms and applications, ” Neural Networks, vol. 13, pp. 411–430, June 2000. [28] M. C. T resch, V . C.-K. K. Cheung, and A. D’A vella, “Matrix factorization algorithms for the identification of muscle synergies: ev aluation on sim- ulated and experimental data sets., ” Journal of neurophysiology , vol. 95, pp. 2199–2212, Apr . 2006. [29] C. Choi and J. Kim, “Synergy matrices to estimate fluid wrist movements by surface electromyography, ” Medical Engineering and Physics, vol. 33, pp. 916–923, Oct. 2011. [30] A. d’A vella and E. Bizzi, “Shared and specific muscle syner gies in natural motor behaviors., ” Proceedings of the National Academy of Sciences of the United States of America, vol. 102, pp. 3076–81, Feb . 2005. [31] V . C.-K. K. Cheung, “Central and Sensory Contrib utions to the Activ ation and Organization of Muscle Synergies during Natural Motor Behaviors, ” Journal of Neuroscience, vol. 25, pp. 6419–6434, July 2005. [32] G. T orres-Oviedo, J. M. Macpherson, and L. H. Ting, “Muscle synergy organization is robust across a variety of postural perturbations., ” Journal of neurophysiology, vol. 96, pp. 1530–1546, Jan. 2006. [33] F . O. Barroso, D. T orricelli, J. C. Moreno, J. T aylor , J. Gomez-Soriano, E. Bravo-Esteban, S. Piazza, C. Santos, and J. L. Pons, “Shared muscle synergies in human walking and cycling, ” Journal of Neurophysiology, vol. 112, no. 8, pp. 1984–1998, 2014. [34] S. A. Chvatal, G. T orres-Oviedo, S. A. Safavynia, and L. H. T ing, “Com- mon muscle synergies for control of center of mass and force in non- stepping and stepping postural behaviors, ” Journal of Neurophysiology , vol. 106, pp. 999–1015, Aug. 2011. [35] G. T orres-Oviedo and L. H. Ting, “Subject-Specific Muscle Synergies in Human Balance Control Are Consistent Across Different Biomechanical Contexts, ” Journal of Neurophysiology , vol. 103, pp. 3084–3098, June 2010. [36] M. Atzori, A. Gijsberts, C. Castellini, B. Caputo, A.-G. M. Hager, S. Elsig, G. Giatsidis, F . Bassetto, and H. Müller, “Electromyography data for non- in vasi ve naturally-controlled robotic hand prostheses., ” Scientific data, vol. 1, p. 140053, Jan. 2014. [37] M. Atzori, A. Gijsberts, I. Kuzborskij, S. Elsig, A.-G. Mittaz Hager , O. Deriaz, C. Castellini, H. Muller, and B. Caputo, “Characterization of a Benchmark Database for Myoelectric Movement Classification, ” IEEE T ransactions on Neural Systems and Rehabilitation Engineering, vol. 23, pp. 73–83, Jan. 2015. [38] N. Jiang, H. Rehbaum, I. V ujaklija, B. Graimann, and D. Farina, “Intu- itiv e, online, simultaneous, and proportional myoelectric control over two degrees-of-freedom in upper limb amputees., ” IEEE transactions on neural systems and rehabilitation engineering, vol. 22, no. 3, pp. 501–10, 2014. [39] J. Ma, N. V . Thak or, and F . Matsuno, “Hand and Wrist Mo vement Control of Myoelectric Prosthesis Based on Synergy , ” IEEE Transactions on Human-Machine Systems, vol. 45, pp. 74–83, Feb . 2015. [40] C. Lin, B. W ang, N. Jiang, and D. Farina, “Robust extraction of basis functions for simultaneous and proportional myoelectric control via sparse non-negati ve matrix factorization, ” Journal of Neural Engineering, v ol. 15, p. 026017, Apr . 2018. [41] O. Debals and L. De Lathauwer , “Stochastic and Deterministic T ensoriza- tion for Blind Signal Separation, ” in Latent V ariable Analysis and Signal Separation, pp. 3–13, Springer , Cham, 2015. [42] P . Comon, “T ensors : A brief introduction, ” IEEE Signal Processing Magazine, vol. 31, pp. 44–53, May 2014. [43] T . G. Kolda and B. W . Bader, “T ensor Decompositions and Applications, ” SIAM Revie w, vol. 51, pp. 455–500, Aug. 2008. [44] R. Bro and H. A. L. Kiers, “A new efficient method for determining the number of components in P ARAF AC models, ” Journal of Chemometrics, vol. 17, pp. 274–286, June 2003. [45] G. Zhou and A. Cichocki, “F ast and unique Tucker decompositions via multiway blind source separation, ” Bulletin of the Polish Academy of Sciences: T echnical Sciences, vol. 60, pp. 389–405, Jan. 2012. [46] R. A. Harshman and M. E. Lundy , “P ARAF AC: Parallel factor analysis, ” Computational Statistics & Data Analysis, vol. 18, pp. 39–72, Aug. 1994. [47] R. Sands and F . W . Y oung, “Component models for three-way data: An alternating least squares algorithm with optimal scaling features, ” Psychometrika, vol. 45, pp. 39–67, Mar . 1980. [48] P . Comon, X. Luciani, and A. L. F . de Almeida, “T ensor decomposi- tions, alternating least squares and other tales, ” Journal of Chemometrics, vol. 23, pp. 393–405, July 2009. [49] A. Cichocki, R. Zdunek, A. H. Phan, and S.-I. Amari, Nonnegativ e Matrix and T ensor Factorizations: Applications to Exploratory Multi-W ay Data Analysis and Blind Source Separation, vol. 1. John Wile y & Sons, 2009. [50] S. Liu and G. Trenkler , “Hadamard, Khatri-Rao, Kronecker and other matrix products, ” Int. J. Inf. Syst. Sci, vol. 4, no. 1, pp. 160–177, 2008. [51] C. A. Andersson and R. Bro, “The N-way T oolbox for MA TLAB, ” Chemometrics and Intelligent Laboratory Systems, vol. 52, pp. 1–4, Aug. 2000. [52] F . Nie, S. Xiang, Y . Song, and C. Zhang, “Extracting the optimal di- mensionality for local tensor discriminant analysis, ” Pattern Recognition, vol. 42, no. 1, pp. 105–114, 2009. 14 VOLUME 4, 2016 Ebied et al. : Preparation of Papers f or IEEE TRANSACTIONS and JOURNALS [53] A. Ebied, E. Kinney-Lang, L. Spyrou, and J. Escudero, “Ev aluation of matrix factorisation approaches for muscle synergy extraction, ” Medical Engineering & Physics, vol. 57, pp. 51–60, July 2018. [54] S. Muceli and D. Farina, “Simultaneous and proportional estimation of hand kinematics from EMG during mirrored mov ements at multiple degrees-of-freedom, ” IEEE T ransactions on Neural Systems and Rehabil- itation Engineering, vol. 20, pp. 371–378, May 2012. [55] N. Jiang, K. B. Englehart, and P . a. Parker , “Extracting simultaneous and proportional neural control information for multiple-dof prostheses from the surface electromyographic signal, ” IEEE Transactions on Biomedical Engineering, vol. 56, pp. 1070–1080, Apr . 2009. [56] S. Ikeda and K. T oyama, “Independent component analysis for noisy data - MEG data analysis, ” Neural Networks, vol. 13, no. 10, pp. 1063–1074, 2000. [57] S. A. Chvatal and L. H. Ting, “Common muscle synergies for balance and walking., ” Frontiers in computational neuroscience, vol. 7, p. 48, Jan. 2013. [58] K. Devarajan, “Nonneg ative matrix factorization: an analytical and in- terpretiv e tool in computational biology ., ” PLoS computational biology , vol. 4, p. e1000029, Jan. 2008. [59] M. Semprini, A. V . Cuppone, I. Delis, V . Squeri, S. Panzeri, and J. Kon- czak, “Biofeedback Signals for Robotic Rehabilitation: Assessment of Wrist Muscle Acti vation Patterns in Healthy Humans, ” IEEE T ransactions on Neural Systems and Rehabilitation Engineering, vol. 25, pp. 883–892, July 2017. [60] M. Mørup, “ Applications of tensor (multiway array) factorizations and decompositions in data mining, ” W iley Interdisciplinary Reviews: Data Mining and Knowledge Disco very , vol. 1, pp. 24–40, Jan. 2011. [61] N. Jiang, T . Lorrain, and D. Farina, “A state-based, proportional myoelec- tric control method: online validation and comparison with the clinical state-of-the-art., ” Journal of neuroengineering and rehabilitation, vol. 11, p. 110, July 2014. VOLUME 4, 2016 15
Original Paper
Loading high-quality paper...
Comments & Academic Discussion
Loading comments...
Leave a Comment