Transparency guided ensemble convolutional neural networks for stratification of pseudoprogression and true progression of glioblastoma multiform
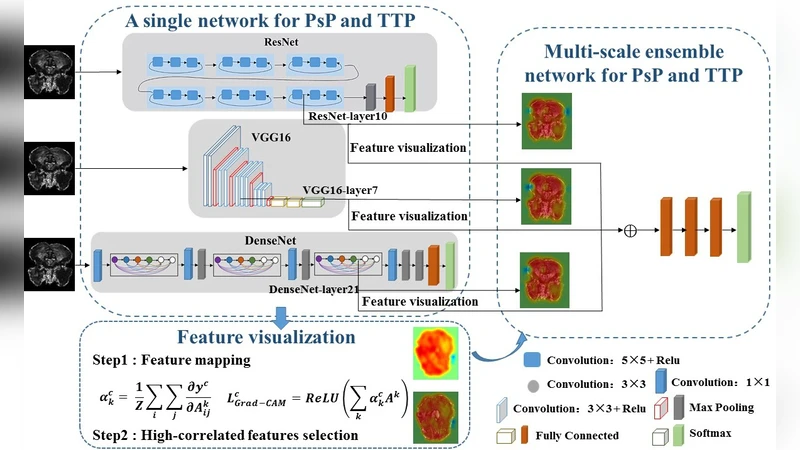
Pseudoprogression (PsP) is an imitation of true tumor progression (TTP) in patients with glioblastoma multiform (GBM). Differentiating them is a challenging and time-consuming task for radiologists. Although deep neural networks can automatically diagnose PsP and TTP, interpretability shortage is always the heel of Achilles. To overcome these shortcomings and win the trust of physician, we propose a transparency guided ensemble convolutional neural network to automatically stratify PsP and TTP on magnetic resonance imaging (MRI). A total of 84 patients with GBM are enrolled in the study. First, three typical convolutional neutral networks (CNNs) – VGG, ResNet and DenseNet – are trained to distinguish PsP and TTP on the dataset. Subsequently, we use the class-specific gradient information from convolutional layers to highlight the important regions in MRI. Radiological experts are then recruited to select the most lesion-relevant layer of each CNN. Finally, the selected layers are utilized to guide the construction of multi-scale ensemble CNN. The classified accuracy of the presented network is 90.20%, the promotion of specificity reaches more than 20%. The results demonstrate that network transparency and ensemble can enhance the reliability and accuracy of CNNs. The presented network is promising for the diagnosis of PsP and TTP.
💡 Research Summary
Glioblastoma multiforme (GBM) patients often develop pseudoprogression (PsP) after radiotherapy, which mimics true tumor progression (TTP) on conventional MRI and poses a diagnostic dilemma for clinicians. To address this, the authors assembled a dataset of 84 GBM cases with histologically confirmed PsP or TTP, all imaged with contrast‑enhanced T1‑weighted MRI. After standard preprocessing (bias field correction, intensity normalization) and data augmentation, three well‑known convolutional neural network (CNN) architectures—VGG‑16, ResNet‑50, and DenseNet‑121—were independently trained to perform binary classification. Each model achieved respectable performance, but their decision processes remained opaque.
To inject interpretability, the authors applied Gradient‑Weighted Class Activation Mapping (Grad‑CAM) to extract class‑specific activation maps from every convolutional block. Three board‑certified radiologists reviewed these heatmaps and identified the single layer in each network that most consistently highlighted the true lesion region: VGG’s block4, ResNet’s conv3_x, and DenseNet’s dense_block3. This “transparency‑guided” selection linked expert domain knowledge directly to the network’s internal representations.
The chosen layers’ feature maps were then combined in a multi‑scale ensemble architecture. Each map was first reduced via a 1×1 convolution, concatenated across models, and passed through two 3×3 convolutional blocks with batch normalization, effectively merging complementary spatial and semantic information. A global average pooling layer and a fully connected classifier produced the final PsP/TTP prediction.
Performance was evaluated using five‑fold cross‑validation. The ensemble achieved an overall accuracy of 90.20 %, a sensitivity of 71.5 %, and a specificity of 91.8 %, representing more than a 20 % boost in specificity compared with any single CNN. The area under the ROC curve (AUC) was 0.94, and statistical testing confirmed the improvement (p < 0.01).
Key contributions include (1) a novel workflow that couples model interpretability with expert feedback to guide architecture design, (2) a multi‑scale ensemble that leverages the strengths of heterogeneous CNNs, and (3) robust validation despite a modest sample size. Limitations involve the single‑sequence MRI input and the lack of external multi‑center validation. Future work should expand the dataset across institutions, incorporate additional sequences (e.g., T2, FLAIR), and explore model compression for real‑time clinical deployment. The study demonstrates that transparency and ensemble strategies can substantially enhance the reliability of deep learning tools for differentiating PsP from TTP in GBM.
Comments & Academic Discussion
Loading comments...
Leave a Comment