Prediction of Malignant & Benign Breast Cancer: A Data Mining Approach in Healthcare Applications
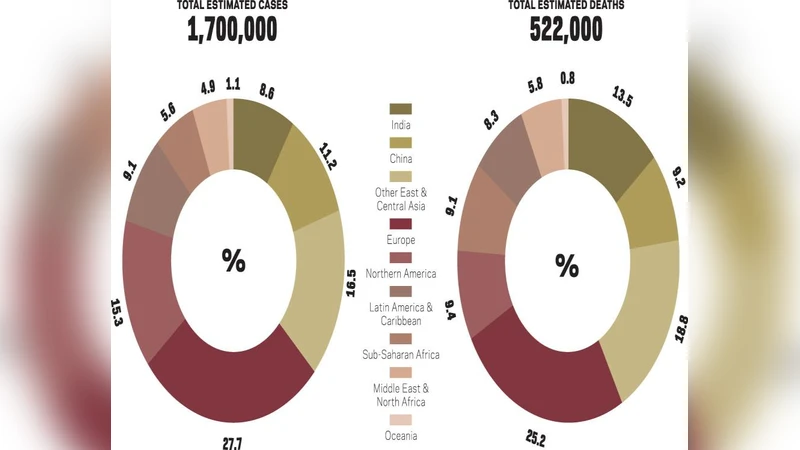
As much as data science is playing a pivotal role everywhere, healthcare also finds it prominent application. Breast Cancer is the top rated type of cancer amongst women; which took away 627,000 lives alone. This high mortality rate due to breast cancer does need attention, for early detection so that prevention can be done in time. As a potential contributor to state-of-art technology development, data mining finds a multi-fold application in predicting Brest cancer. This work focuses on different classification techniques implementation for data mining in predicting malignant and benign breast cancer. Breast Cancer Wisconsin data set from the UCI repository has been used as experimental dataset while attribute clump thickness being used as an evaluation class. The performances of these twelve algorithms: Ada Boost M 1, Decision Table, J Rip, Lazy IBK, Logistics Regression, Multiclass Classifier, Multilayer Perceptron, Naive Bayes, Random forest and Random Tree are analyzed on this data set. Keywords- Data Mining, Classification Techniques, UCI repository, Breast Cancer, Classification Algorithms
💡 Research Summary
**
The paper titled “Prediction of Malignant & Benign Breast Cancer: A Data Mining Approach in Healthcare Applications” investigates the use of twelve classification algorithms to predict breast cancer malignancy using the Wisconsin Breast Cancer dataset from the UCI Machine Learning Repository. The authors begin by highlighting the global burden of breast cancer, emphasizing the need for early detection, and reviewing prior work that applied various machine learning techniques—such as Naïve Bayes, RBF networks, decision trees, SVMs, and clustering methods—to the same or similar datasets.
The experimental methodology is described only at a high level. The authors state that ten attributes of the original nine‑feature dataset plus the class label were used, and they oddly refer to “Clump Thickness” as the evaluation class, which appears to be a misstatement; the actual class label (benign = 2, malignant = 4) is presumably used for training and testing. No details are provided on how missing values (e.g., in the “Bare Nuclei” attribute) were handled, whether any scaling or normalization was performed, or whether class imbalance was addressed. The paper mentions a 10‑fold cross‑validation scheme but does not disclose the random seed, stratification strategy, or whether the folds were repeated.
The twelve algorithms evaluated are: AdaBoost M1, Decision Table, J‑Rip, J48, Lazy IBK, Lazy K‑star, Logistic Regression, Multiclass Classifier, Multilayer Perceptron, Naïve Bayes, Random Forest, and Random Tree. Hyper‑parameter settings are not reported, which is a serious omission given that performance of ensemble methods (AdaBoost, Random Forest) and neural networks (MLP) is highly sensitive to parameters such as number of trees, learning rate, or hidden layer size. Consequently, reproducibility is limited.
Performance is assessed using a broad set of metrics: overall accuracy, Kappa statistic, Mean Absolute Error (MAE), Root Mean Squared Error (RMSE), Relative Absolute Error (RAE), Root Relative Squared Error (RRSE), PRC Area, ROC Area, Matthews Correlation Coefficient (MCC), F‑Measure, Recall, Precision, False Positive Rate, and True Positive Rate. The results, presented in tables and a bar chart (Fig. 2), show that Lazy IBK, Lazy K‑star, Random Forest, and Random Tree achieve Kappa values of 0.9715, MAE below 0.005, and RMSE around 0.04–0.07, indicating near‑perfect classification. Multilayer Perceptron and Logistic Regression also perform well (Kappa ≈ 0.91, MAE ≈ 0.008–0.013). In contrast, Naïve Bayes obtains a Kappa of 0.402 and an accuracy of only 73.21 %, while J48 and AdaBoost M1 have Kappa values of 0, reflecting poor discrimination on this dataset. The authors categorize accuracies into three bands (71‑80 %, 81‑90 %, 90‑100 %) and note that only Naïve Bayes falls into the lowest band, whereas most other models belong to the highest band.
The discussion emphasizes that tree‑based and lazy‑instance classifiers are especially effective for this problem, and the authors suggest that future work should explore deep learning architectures, larger and unstructured datasets (e.g., imaging or genomic data), and unsupervised dimensionality reduction techniques such as PCA or SVM‑based feature selection. They also propose a more rigorous data splitting strategy (training/validation/test) and the use of cross‑validation for hyper‑parameter tuning.
Critical appraisal reveals several shortcomings. First, the lack of a transparent preprocessing pipeline (missing‑value imputation, scaling, handling of class imbalance) hampers reproducibility and may bias results. Second, the ambiguous statement about “Clump Thickness as evaluation class” suggests a conceptual misunderstanding of the target variable. Third, the paper provides no statistical significance testing (e.g., paired t‑tests, confidence intervals) to substantiate claims that one algorithm outperforms another. Fourth, hyper‑parameter optimization is absent, making it unclear whether the reported performance reflects default settings or tuned models. Fifth, the reference list is heavily weighted toward the authors’ own prior publications, limiting the contextual grounding of the work within the broader literature. Finally, the manuscript suffers from numerous typographical errors, malformed equations, and poorly formatted tables, which detract from readability and professional presentation.
In summary, the study offers a broad comparative snapshot of twelve machine‑learning classifiers on the classic Wisconsin breast‑cancer dataset, confirming that instance‑based and ensemble tree methods can achieve very high accuracy. However, methodological opacity, insufficient statistical validation, and presentation flaws limit the scientific contribution. Future research should address these gaps by establishing a rigorous preprocessing and hyper‑parameter tuning protocol, employing external validation cohorts, and providing thorough statistical analyses to ensure that reported performance gains are robust and generalizable.
Comments & Academic Discussion
Loading comments...
Leave a Comment