Reduce Noise in Computed Tomography Image using Adaptive Gaussian Filter
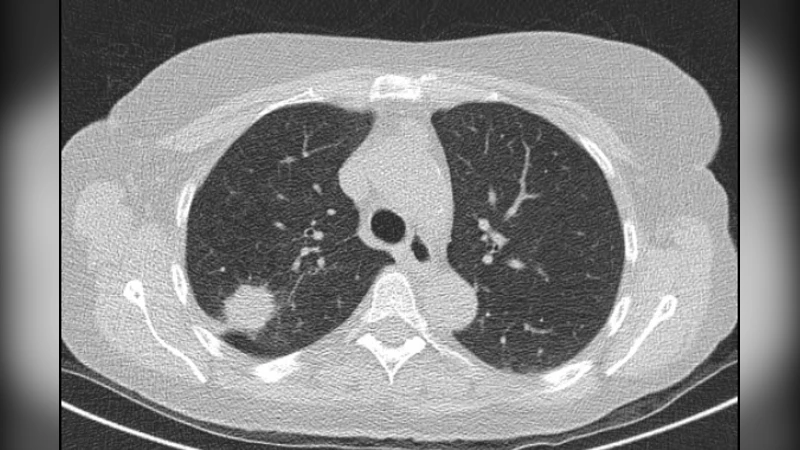
One image processing application that is very helpful for humans is to improve image quality, poor image quality makes the image more difficult to interpret because the information conveyed by the image is reduced. In the process of the acquisition of medical images, the resulting image has decreased quality (degraded) due to external factors and medical equipment used. For this reason, it is necessary to have an image processing process to improve the quality of medical images, so that later it is expected to help facilitate medical personnel in analyzing and translating medical images, which will lead to an improvement in the quality of diagnosis. In this study, an analysis will be carried out to improve the quality of medical images with noise reduction with the Gaussian Filter Method. Next, it is carried out, and tested against medical images, in this case, the lung photo image. The test image is given noise in the form of impulse salt & pepper and adaptive Gaussian then analyzed its performance qualitatively by comparing the output filter image, noise image, and the original image by naked eye.
💡 Research Summary
The paper addresses the pervasive problem of noise degradation in computed tomography (CT) images, focusing specifically on lung scans. Recognizing that noisy images hinder accurate diagnosis, the authors propose an “adaptive Gaussian filter” as a preprocessing step to improve visual quality. The core idea is to retain the smoothing benefits of a conventional Gaussian blur while dynamically adjusting the filter’s standard deviation (σ) based on local image statistics. In regions where noise density is high, a larger σ is applied to achieve stronger smoothing; in areas containing critical anatomical edges, a smaller σ preserves detail.
Methodologically, the study follows a straightforward pipeline. First, a single lung CT slice is selected as the test image. Synthetic impulse noise—commonly known as salt‑and‑pepper noise—is then added to simulate a worst‑case degradation scenario. This type of noise flips pixel values to either the minimum or maximum intensity, creating stark outliers that are easy to detect visually. Next, the adaptive Gaussian filter is applied. For each pixel, a fixed‑size neighborhood window is examined; the local mean and variance are computed, and these statistics are used to determine an appropriate σ for that window. The resulting filter is thus spatially variant: high‑variance windows receive stronger blurring, while low‑variance windows are treated more gently.
The experimental results are presented solely as side‑by‑side visual comparisons among three images: the original clean CT slice, the noise‑contaminated version, and the filtered output. The authors claim that the adaptive filter substantially reduces the salt‑and‑pepper speckles while preserving the overall lung anatomy. They argue that this balance between noise suppression and edge retention is superior to a conventional Gaussian filter with a fixed σ, which either leaves residual noise or overly smooths important structures.
Despite these qualitative observations, the paper lacks several critical components that are standard in modern image‑processing research. First, there is no quantitative evaluation. Metrics such as Peak Signal‑to‑Noise Ratio (PSNR), Structural Similarity Index (SSIM), or Mean Squared Error (MSE) are absent, making it impossible to objectively assess the improvement. Second, the study uses only a single 2‑D CT slice, which raises concerns about the generalizability of the findings to full 3‑D volumes, different anatomical regions, or other noise models (e.g., quantum noise, reconstruction artifacts). Third, the authors do not compare their method against established alternatives such as median filtering, bilateral filtering, non‑local means, or wavelet‑based denoising, leaving the claimed superiority unsubstantiated.
Implementation details are also sparse. The paper does not disclose the exact formula used to map local variance to σ, the size of the sliding window, how border pixels are handled, or the computational cost of the adaptive process. These omissions hinder reproducibility and prevent an assessment of whether the method can be deployed in real‑time clinical workflows, where processing speed is a crucial factor.
The discussion acknowledges the visual success of the adaptive filter but fails to address its limitations. For instance, adaptive smoothing can inadvertently introduce artifacts near high‑contrast edges if the variance estimate is unstable. Moreover, the lack of a statistical analysis of the results means that the observed improvements could be due to chance or specific to the chosen test image.
In conclusion, the paper introduces an intuitive concept—locally varying the Gaussian kernel’s σ—to tackle impulse noise in CT images. While the idea is sound and the visual examples are encouraging, the work falls short of the rigorous standards required for clinical translation. Future research should (1) provide a detailed algorithmic description, (2) evaluate performance on a diverse dataset of CT volumes with realistic noise, (3) benchmark against a suite of state‑of‑the‑art denoising techniques using objective quality metrics, (4) involve radiologists in a blind reading study to measure diagnostic impact, and (5) analyze computational complexity to ensure feasibility in a hospital setting. Only with such comprehensive validation can the adaptive Gaussian filter be considered a viable tool for improving CT image quality in practice.
Comments & Academic Discussion
Loading comments...
Leave a Comment