Parametric FEM for Shape Optimization applied to Golgi Stack
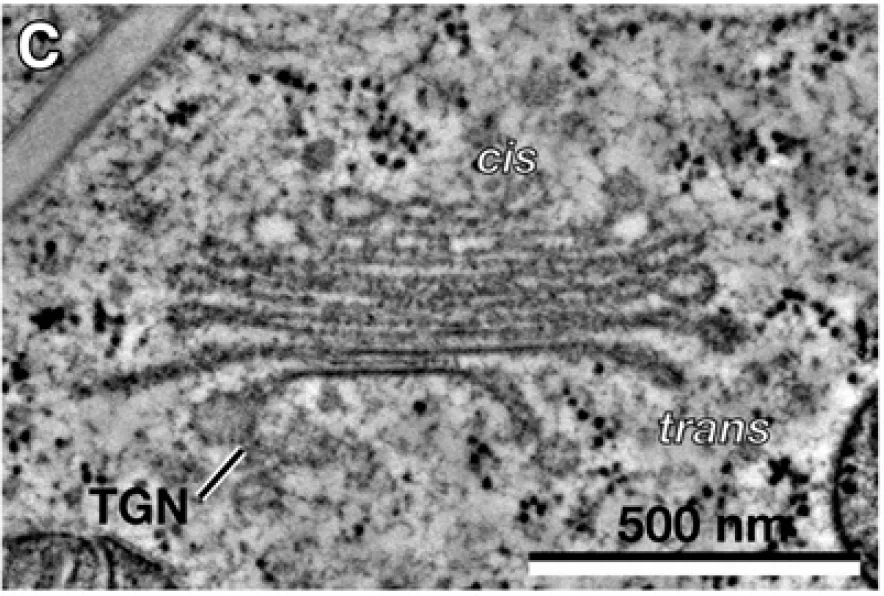
The thesis is about an application of the shape optimization to the morphological evolution of Golgi stack. Golgi stack consists of multiple layers of cisternae. It is an organelle in the biological cells. Inspired by the Helfrich Model \cite{Helfrich}, which is a model for vesicles typically applied to biological cells, a new model specially designed for Golgi stack is developed and then implemented using FEM in this thesis. In the Golgi model, each cisternae of the Golgi stack is viewed as a closed vesicle without topological changes, and our model is adaptable to both single-vesicle case and multiple-vesicle case. The main idea of the math model is to minimize the elastic energy(bending energy) of the vesicles, with some constraints designed regarding the biological properties of Golgi stack. With these constraints attached to the math model, we could extend this model to an obstacle-type problem. Hence, in the thesis, not only the simulations of Golgi stack are shown, but some interesting examples without biological meanings are also demonstrated. Also, as multiple cisternaes are considered as a whole, this is also a model handling multiple objects. A set of numerical examples is shown to compare with the observed shape of Golgi stack, so we can lay down some possible explanations to the morphological performance of trans-Golgi cisternae.
💡 Research Summary
The thesis presents a comprehensive computational framework for modeling the morphological evolution of Golgi stacks, a key organelle composed of multiple stacked cisternae, by formulating and solving a shape‑optimization problem using parametric finite‑element methods (FEM). The core of the work is the construction of an energy functional that captures the dominant physical forces acting on a membrane while incorporating biologically motivated constraints.
Physical model
The authors adopt the Helfrich‑Willmore description of lipid bilayer membranes, where the bending (elastic) energy is expressed as the integral of the squared mean curvature over the surface. Because the Gaussian curvature term reduces to a topological constant for closed surfaces, the functional is simplified to the Willmore energy
(W(\Gamma)=\int_{\Gamma} h^{2},dS).
In addition to this curvature‑driven term, three constraints are introduced:
-
Area constraint – the total membrane area is assumed constant, reflecting a fixed number of lipid molecules in a mature trans‑Golgi cisterna. This is enforced via a Lagrange multiplier.
-
Barrier (obstacle) functional – inter‑cisternal elements (e.g., stacking proteins) are modeled as a spatial region (B) that the membrane cannot penetrate. An indicator function (1_{B}) is integrated over the surface, yielding a penalty term that becomes large when the membrane enters the forbidden zone.
-
Distance functional – when several cisternae are considered simultaneously, a repulsive interaction is needed to prevent overlap. The authors define a pairwise distance‑based functional that penalizes small separations, thereby mimicking steric repulsion between adjacent membranes.
Three concrete models are built from these ingredients:
Model 1 includes only the Willmore energy and the area constraint (a classic problem).
Model 2 adds the barrier functional, allowing the study of static or moving obstacles that confine the membrane vertically.
Model 3 incorporates the distance functional to handle multiple vesicles as a coupled system.
Mathematical analysis
For each functional the shape derivative (first variation) is derived using the calculus of variations on manifolds. The Willmore term yields a fourth‑order nonlinear operator involving the surface Laplacian of the mean curvature, while the barrier and distance terms contribute lower‑order, often distributional, contributions localized near the obstacle boundaries or inter‑surface gaps. The area constraint introduces a constant Lagrange multiplier, whose value is updated iteratively to enforce the prescribed area.
Numerical scheme
Time discretization follows an explicit forward‑Euler approach: the normal velocity obtained from the shape derivative is used to update the surface position. Spatial discretization employs a parametric FEM where the evolving surface is represented by a triangulated mesh. Quadratic Lagrange shape functions are used for geometric accuracy, and the surface differential operators (tangential gradient, divergence, Laplace–Beltrami) are assembled on this mesh. Nonlinear terms (especially those arising from the Willmore curvature) are linearized via a Newton–Raphson procedure, leading to a sequence of linear systems solved at each time step. An area‑renormalization step rescales the mesh to keep the total area exactly at the target value, ensuring numerical stability.
Algorithmic flow
- Generate an initial triangulated surface and define obstacle regions and inter‑surface distances.
- Assemble the discrete curvature, barrier, and distance contributions and compute the shape derivative.
- Solve the linearized system for the nodal displacement, including the Lagrange multiplier for area.
- Update node positions, renormalize area, and repeat until convergence or a prescribed final time.
Results
The authors present several simulation scenarios:
- Single cisterna with fixed upper and lower barriers – the membrane expands laterally but is prevented from growing vertically, reproducing the thin, flattened appearance of trans‑Golgi cisternae observed in electron microscopy.
- Moving barriers – the surface adapts smoothly to the shifting obstacle, demonstrating the method’s ability to handle dynamic constraints.
- Multiple cisternae with distance repulsion – the vesicles maintain a uniform spacing without intersecting, mimicking the regular stacking of Golgi layers.
Quantitative diagnostics (energy decay, area conservation) confirm the correctness of the implementation, while qualitative comparison with experimental images suggests that the barrier and distance terms capture essential biological mechanisms: vertical confinement by stacking proteins and steric repulsion between adjacent membranes.
Discussion and limitations
The work successfully integrates curvature‑driven mechanics with biologically realistic constraints, offering a novel computational tool for organelle morphology. However, the model omits volume (pressure) constraints, treats obstacles as binary indicator functions, and does not account for active processes such as protein trafficking or membrane remodeling enzymes. The explicit time integration limits the allowable time step, and higher‑order implicit schemes could improve efficiency. Future extensions could incorporate volumetric terms, heterogeneous material properties, and coupling with reaction‑diffusion models of membrane proteins.
Conclusion
Overall, the thesis delivers a mathematically rigorous and numerically robust framework for simulating Golgi stack morphology. By extending the classical Helfrich‑Willmore model with area, obstacle, and distance constraints, and by implementing the resulting variational problem with parametric FEM, the author provides both a methodological contribution to shape‑optimization on surfaces and biologically relevant insights into the physical determinants of Golgi architecture. This interdisciplinary effort bridges differential geometry, numerical analysis, and cell biology, and sets the stage for more sophisticated multi‑physics models of intracellular organelles.
Comments & Academic Discussion
Loading comments...
Leave a Comment