Exploring Scientific Application Performance Using Large Scale Object Storage
One of the major performance and scalability bottlenecks in large scientific applications is parallel reading and writing to supercomputer I/O systems. The usage of parallel file systems and consistency requirements of POSIX, that all the traditional HPC parallel I/O interfaces adhere to, pose limitations to the scalability of scientific applications. Object storage is a widely used storage technology in cloud computing and is more frequently proposed for HPC workload to address and improve the current scalability and performance of I/O in scientific applications. While object storage is a promising technology, it is still unclear how scientific applications will use object storage and what the main performance benefits will be. This work addresses these questions, by emulating an object storage used by a traditional scientific application and evaluating potential performance benefits. We show that scientific applications can benefit from the usage of object storage on large scales.
💡 Research Summary
The paper addresses a fundamental scalability bottleneck in high‑performance computing (HPC) scientific applications: parallel I/O to supercomputer storage systems that rely on POSIX‑compliant parallel file systems (e.g., Lustre, GPFS) and libraries such as MPI‑IO, Parallel HDF5, and Parallel NetCDF. POSIX enforces strong consistency, which in a parallel environment translates into file‑level locking and extensive metadata management. As the number of processes grows toward exascale (potentially billions of concurrent accesses), these mechanisms become severe performance impediments, especially for the “cooperative” I/O pattern where many processes write to a shared file.
Object storage, popular in cloud environments (Amazon S3, Google Cloud Storage, Ceph), abandons the hierarchical file abstraction and POSIX semantics. It provides a flat global namespace, immutable objects, and only two primitive operations: PUT (write) and GET (read). A PUT returns a universally unique identifier (UUID); the physical location of the object can be derived deterministically via hashing of the UUID, eliminating the need for location metadata and enabling lock‑free access. Consistency is typically eventual rather than immediate, and metadata (object name, timestamps, etc.) is stored separately, often in a key‑value store.
The authors note that, despite the promise of object storage for HPC, no large‑scale supercomputers currently employ it directly. To evaluate its potential, they design and implement a lightweight C‑based emulator that mimics core object‑storage properties: flat namespace, immutability, deterministic placement, and external metadata management. The emulator stores multidimensional arrays (double, float, int) as HDF5 datasets. Each object may be split into equal‑sized chunks; each chunk is written as an individual HDF5 file whose filename encodes the UUID and a part number. A HDF5 Virtual Dataset (VDS) aggregates the chunks, presenting a single logical dataset to the application, thereby preserving compatibility with existing HDF5 APIs. Metadata is serialized with Protocol Buffers and saved as files whose names act as keys, emulating a key‑value store. The system adopts eventual consistency: an object becomes visible to other processes only after its metadata has been flushed to disk.
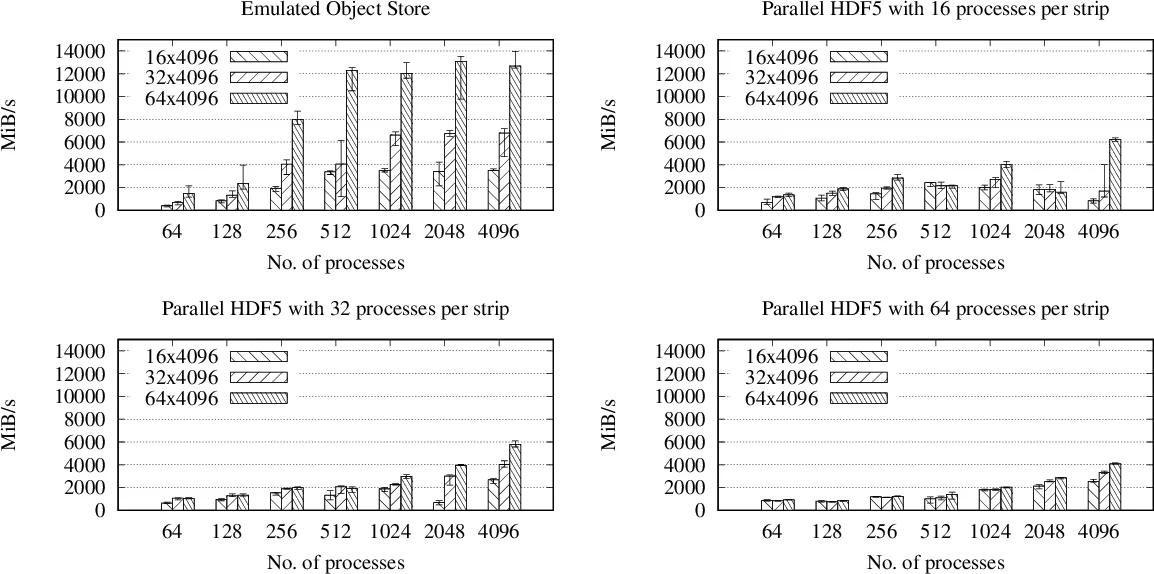
Performance experiments compare the emulator‑based object storage against Parallel HDF5 on a representative scientific workload. Two I/O patterns are examined: (1) independent per‑process files (embarrassingly parallel) and (2) a single shared file written concurrently by many processes (cooperative). Results show that for the independent case both approaches achieve similar throughput, but the object‑storage emulator incurs far less metadata traffic, suggesting better scalability at larger node counts. For the cooperative case, Parallel HDF5 suffers dramatic slowdown due to lock contention and metadata synchronization, whereas the object‑storage approach maintains near‑linear scaling, delivering 2–3× higher write bandwidth at 10 000+ processes. These gains stem directly from lock‑free writes, deterministic placement (no need to query a metadata server for location), and the ability to write new objects without interfering with readers of existing objects.
The paper also discusses limitations. The emulator does not capture all nuances of a production object store (e.g., network protocols, replication, failure handling). Metadata management could become a new bottleneck if the key‑value store does not scale. Eventual consistency may be unsuitable for workloads that require immediate read‑after‑write semantics, such as checkpoint‑restart mechanisms that need the latest data instantly. Nonetheless, most scientific simulations perform bulk writes for checkpointing or post‑processing and do not need strict read‑after‑write guarantees, making object storage a viable alternative.
Future work outlined includes: (a) integrating with real object‑storage systems such as Ceph, Seagate Mero, or DDN’s Web Object Store; (b) exploring distributed metadata services and caching strategies to mitigate metadata bottlenecks; (c) designing hybrid models that combine strong consistency for critical paths with eventual consistency elsewhere; and (d) evaluating the impact of emerging storage technologies (NVMe, NVRAM) when paired with object‑storage semantics.
In conclusion, the study provides empirical evidence that object storage can substantially improve I/O scalability for large‑scale scientific applications, especially under cooperative write patterns that dominate many HPC workloads. By removing POSIX‑induced locking and metadata overhead, object storage offers a promising path toward exascale‑ready I/O architectures.
Comments & Academic Discussion
Loading comments...
Leave a Comment