A Probabilistic U-Net for Segmentation of Ambiguous Images
Many real-world vision problems suffer from inherent ambiguities. In clinical applications for example, it might not be clear from a CT scan alone which particular region is cancer tissue. Therefore a group of graders typically produces a set of diverse but plausible segmentations. We consider the task of learning a distribution over segmentations given an input. To this end we propose a generative segmentation model based on a combination of a U-Net with a conditional variational autoencoder that is capable of efficiently producing an unlimited number of plausible hypotheses. We show on a lung abnormalities segmentation task and on a Cityscapes segmentation task that our model reproduces the possible segmentation variants as well as the frequencies with which they occur, doing so significantly better than published approaches. These models could have a high impact in real-world applications, such as being used as clinical decision-making algorithms accounting for multiple plausible semantic segmentation hypotheses to provide possible diagnoses and recommend further actions to resolve the present ambiguities.
💡 Research Summary
The paper addresses a fundamental limitation of most modern semantic segmentation models: they either produce a single deterministic mask or a pixel‑wise probability map, both of which fail to capture the inherent ambiguity present in many real‑world images. In medical imaging, for example, a CT scan may clearly show a lesion but cannot reveal whether the tissue is malignant; in everyday scenes, a partially occluded animal could be either a cat or a dog. In such cases, multiple plausible segmentations exist, each with its own likelihood, and downstream decision‑making systems would benefit from being aware of this distribution rather than a single guess.
To solve this, the authors propose the “Probabilistic U‑Net”, a novel architecture that merges a conditional variational auto‑encoder (CVAE) with the popular U‑Net segmentation backbone. The key idea is to learn a low‑dimensional latent space (typically six dimensions) that encodes the different segmentation modes for a given input image. A “prior network” conditioned on the input image predicts a Gaussian distribution over this latent space, while a “posterior network” conditioned on both the image and a ground‑truth mask predicts a Gaussian that should match the prior for that specific mode. Training follows the standard evidence‑lower‑bound (ELBO) objective: a cross‑entropy term forces a sampled latent vector to reconstruct the associated ground‑truth mask, and a Kullback‑Leibler (KL) term aligns the posterior with the prior, thereby encouraging the prior to cover all observed modes across the dataset.
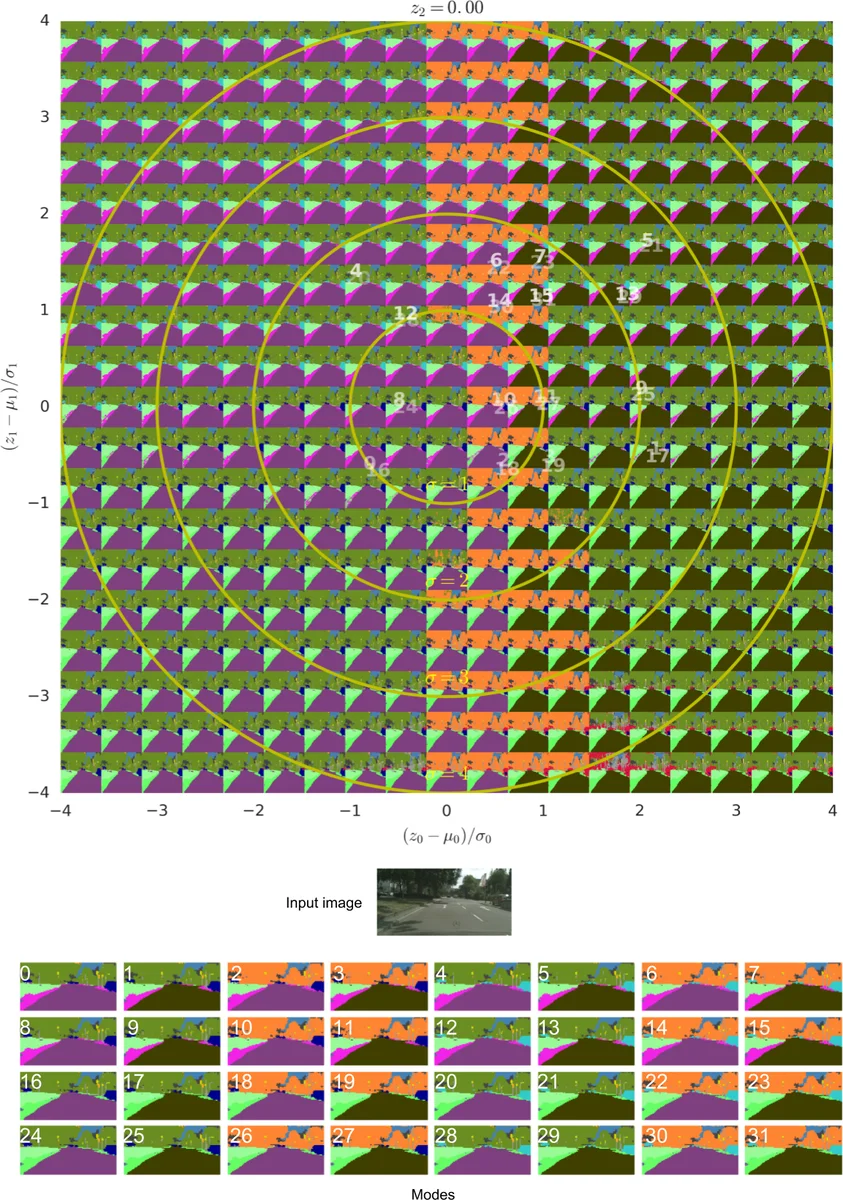
During inference, a latent vector is sampled from the learned prior, broadcast as an N‑channel feature map, concatenated with the final U‑Net feature map, and passed through a small 1×1 convolutional head to produce a full‑resolution segmentation mask. Because the bulk of the U‑Net computation is shared across samples, generating many hypotheses for the same image is computationally cheap.
The authors evaluate the model on two datasets that provide multiple annotations per image: the LIDC‑IDRI lung CT dataset (four radiologists per scan) and a modified version of Cityscapes where certain classes are artificially swapped to create label ambiguity. They introduce a rigorous quantitative metric, the Generalized Energy Distance (GED), which measures the distance between two distributions of segmentations using an IoU‑based ground metric. GED is well‑suited for assessing mode coverage and calibration, unlike simple pixel‑wise metrics.
Compared against several strong baselines—Dropout U‑Net (Monte‑Carlo dropout), an ensemble of independent U‑Nets, an M‑head U‑Net (multiple heads sharing a backbone), and an Image‑to‑Image VAE (fixed prior, no conditioning)—the Probabilistic U‑Net consistently achieves lower GED scores, indicating superior modeling of the true distribution of segmentations. On LIDC‑IDRI, the model not only reproduces the four expert masks but also captures the relative frequencies with which each expert’s annotation occurs, even for rare modes that appear in only a small fraction of cases. On Cityscapes, the model accurately reflects the artificially imposed class‑swap probabilities, demonstrating that it learns calibrated probabilities rather than merely generating diverse masks.
Beyond quantitative results, the paper provides qualitative examples showing that each sampled mask is globally coherent (i.e., respects object shape and context) unlike pixel‑wise dropout samples that can be noisy and inconsistent. The authors also analyze calibration by correlating the empirical frequency of sampled modes with their ground‑truth frequencies, reporting high correlation coefficients (R² ≈ 0.92), confirming that the latent space’s density reflects true mode likelihoods.
The proposed approach has several notable strengths: (1) it produces full‑image, jointly consistent segmentation hypotheses; (2) it can model arbitrarily complex, multimodal output distributions, including low‑probability events; (3) sampling is computationally efficient, enabling real‑time use; (4) it allows rigorous, distribution‑level evaluation via GED, moving beyond pixel‑wise uncertainty estimates. Limitations include the fixed latent dimensionality (six dimensions may be insufficient for extremely complex ambiguities) and the reliance on fully supervised data; extending the method to semi‑supervised or weakly supervised settings remains an open question.
In conclusion, the Probabilistic U‑Net represents a significant step toward uncertainty‑aware semantic segmentation. By learning a conditional distribution over plausible masks, it equips downstream systems—particularly in high‑stakes domains like medical diagnosis or autonomous driving—with richer information about what the model “believes” could be true, enabling better risk assessment, targeted follow‑up actions, and ultimately more trustworthy AI. Future work may explore adaptive latent dimensionality, integration with active learning for label acquisition, and deployment in clinical pipelines where multiple plausible segmentations can be presented to radiologists for review.
Comments & Academic Discussion
Loading comments...
Leave a Comment