Nonlinear Dimensionality Reduction for Discriminative Analytics of Multiple Datasets
Principal component analysis (PCA) is widely used for feature extraction and dimensionality reduction, with documented merits in diverse tasks involving high-dimensional data. Standard PCA copes with one dataset at a time, but it is challenged when it comes to analyzing multiple datasets jointly. In certain data science settings however, one is often interested in extracting the most discriminative information from one dataset of particular interest (a.k.a. target data) relative to the other(s) (a.k.a. background data). To this end, this paper puts forth a novel approach, termed discriminative (d) PCA, for such discriminative analytics of multiple datasets. Under certain conditions, dPCA is proved to be least-squares optimal in recovering the component vector unique to the target data relative to background data. To account for nonlinear data correlations, (linear) dPCA models for one or multiple background datasets are generalized through kernel-based learning. Interestingly, all dPCA variants admit an analytical solution obtainable with a single (generalized) eigenvalue decomposition. Finally, corroborating dimensionality reduction tests using both synthetic and real datasets are provided to validate the effectiveness of the proposed methods.
💡 Research Summary
The paper addresses a fundamental limitation of classical Principal Component Analysis (PCA) when multiple related data sets must be analyzed jointly. Standard PCA processes a single data matrix and therefore cannot isolate components that are unique to a “target” data set in the presence of one or more “background” data sets that share common variability. Existing contrastive PCA (cPCA) attempts to solve this by maximizing target variance while penalizing background variance, but it requires a hyper‑parameter α whose selection is non‑trivial and computationally expensive.
To overcome these issues, the authors propose Discriminative PCA (dPCA), a parameter‑free method that directly maximizes the ratio of target to background variance. Formally, dPCA solves
max₍u₎ (uᵀCₓₓ u) / (uᵀCᵧᵧ u) subject to ‖u‖=1,
where Cₓₓ and Cᵧᵧ are the sample covariance matrices of the target and background data, respectively. By Lagrange multiplier theory the problem reduces to a generalized eigenvalue problem Cₓₓ u = λ Cᵧᵧ u. The eigenvector associated with the largest eigenvalue λ₁ provides the first discriminative component; subsequent components are obtained analogously. When Cᵧᵧ = I, dPCA collapses to ordinary PCA, showing that dPCA is a natural extension.
The framework is further generalized to handle multiple background data sets. By aggregating their covariance matrices (e.g., summation) into a single C_bg, the same ratio‑maximization yields Multi‑background dPCA (MdPCA), which simultaneously suppresses all shared background variations.
To capture nonlinear relationships, the authors kernelize the approach. Data are implicitly mapped to a high‑dimensional feature space φ(·) and the covariances are represented by kernel matrices Kₓₓ and Kᵧᵧ. The generalized eigenvalue problem becomes Kₓₓ α = λ Kᵧᵧ α, whose solution α defines the discriminative direction in the feature space. This yields Kernel dPCA (KdPCA) and its multi‑background counterpart (KMdPCA). The computational cost scales with the number of samples rather than the original dimensionality, making the method suitable for high‑dimensional data.
Theoretical contributions include a proof of LS‑optimality under a linear factor‑analysis model: if the background data follow a low‑rank subspace plus isotropic noise and the target data share this subspace plus additional unique components, dPCA exactly recovers the unique subspace. Moreover, the paper shows that cPCA with α equal to the optimal Lagrange multiplier λ* from dPCA yields the same direction, thereby clarifying the relationship between the two methods and highlighting the advantage of a parameter‑free formulation.
Computationally, both dPCA and its kernel versions require a single generalized eigenvalue decomposition, which can be efficiently solved via Cholesky factorization or Lanczos iterations for large problems.
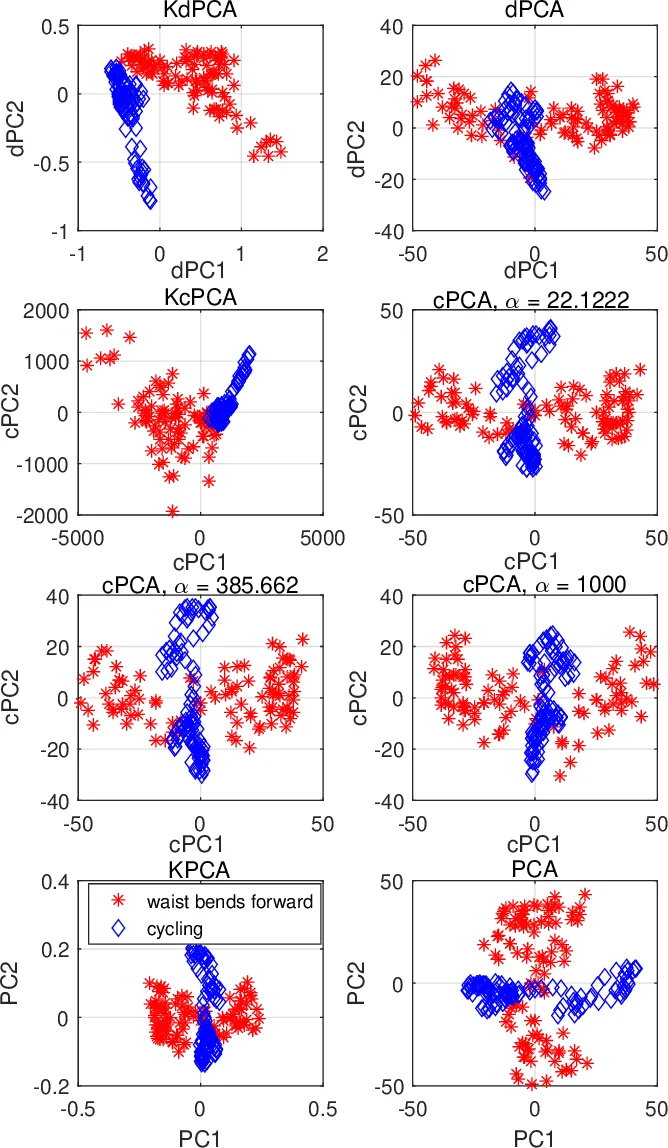
Empirical validation is performed on synthetic and real data. In synthetic two‑cluster data with strong background noise, dPCA/KdPCA cleanly separate the clusters, outperforming PCA, cPCA, and LDA. In a gene‑expression study (cancer patients vs. healthy controls), the methods isolate cancer‑specific subtypes while suppressing demographic variation, achieving clearer clustering than baseline methods. In a face‑image experiment, illumination and expression are treated as background; dPCA extracts identity‑specific features, producing visually distinct representations. Quantitative metrics (variance ratios, clustering scores) corroborate these qualitative observations.
In summary, the paper introduces a unified, parameter‑free discriminative dimensionality reduction framework that (i) isolates target‑specific variance, (ii) extends naturally to multiple background data sets, and (iii) accommodates nonlinear structures via kernelization. Theoretical optimality guarantees and extensive experiments demonstrate its superiority over existing techniques, making it a valuable tool for domains such as bioinformatics, computer vision, and signal processing where distinguishing target signals from shared background is essential.
Comments & Academic Discussion
Loading comments...
Leave a Comment