Multiscale edge detection and parametric shape modeling for boundary delineation in optoacoustic images
In this article, we present a novel scheme for segmenting the image boundary (with the background) in optoacoustic small animal in vivo imaging systems. The method utilizes a multiscale edge detection algorithm to generate a binary edge map. A scale dependent morphological operation is employed to clean spurious edges. Thereafter, an ellipse is fitted to the edge map through constrained parametric transformations and iterative goodness of fit calculations. The method delimits the tissue edges through the curve fitting model, which has shown high levels of accuracy. Thus, this method enables segmentation of optoacoutic images with minimal human intervention, by eliminating need of scale selection for multiscale processing and seed point determination for contour mapping.
💡 Research Summary
This paper introduces a fully automated segmentation pipeline tailored for in‑vivo optoacoustic tomography (OAT) of small animals, aiming to delineate tissue boundaries from the acoustic coupling medium with minimal user interaction. The authors first address the limitations of conventional single‑scale edge detectors (e.g., Sobel, Canny) which are highly susceptible to the low intrinsic contrast, limited view artifacts, and noise typical of OAT reconstructions. To overcome these issues, they adapt a multiscale edge‑flow algorithm originally proposed for color images, integrating anisotropic diffusion to preserve edge strength while smoothing homogeneous regions. The process begins at the finest scale (s = 1) and proceeds in sub‑pixel increments (Δs = 0.5) up to a coarser scale (s = 3). At each scale, a gradient vector field is computed; edges are retained only when the directional change of the vector field is consistent across multiple scales, thereby filtering out spurious responses that appear at a single scale.
Following edge detection, a scale‑dependent morphological refinement is applied. For scales ≥ 2, a 1‑pixel disc structuring element erodes the binary edge map to eliminate isolated noisy patches, after which a closing operation with a larger (2‑pixel) disc restores edge continuity. This two‑stage morphological filter adapts its structuring element size to the current scale, ensuring that thin but genuine edges are not lost while noisy artifacts are suppressed.
The refined edge map is then reduced to a set of centroid points representing connected edge clusters. Outlier centroids are removed using a median filter, and the remaining points are used to fit a parametric shape—either a minimum‑bounding ellipse or circle—through an iterative optimization that maximizes a goodness‑of‑fit (GoF) metric. The GoF is quantified by the Dice coefficient (DM), defined as twice the area of overlap between the fitted region (A) and a manually annotated ground truth (M) divided by the sum of their areas. A Dice value ≥ 0.7 is considered acceptable; in practice the proposed method achieves DM values between 0.90 and 0.96, indicating excellent agreement with expert segmentation. The Rand Index (RI) is also reported, consistently exceeding 0.92, further confirming the reliability of the segmentation.
Experimental validation was performed on a dataset of 30 multispectral OAT volumes acquired with an iTheraMedical MSOT256‑TF system. Each mouse was scanned at six positions and eight wavelengths (690–900 nm), with ten‑frame averaging to improve signal‑to‑noise ratio. Reconstruction employed a non‑negative constrained model‑based inversion, yielding 200 × 200 pixel images covering a 20 mm × 20 mm field of view. For quantitative assessment, four volumes from each of three anatomical regions (brain, liver, kidney/spleen) were manually segmented to serve as ground truth.
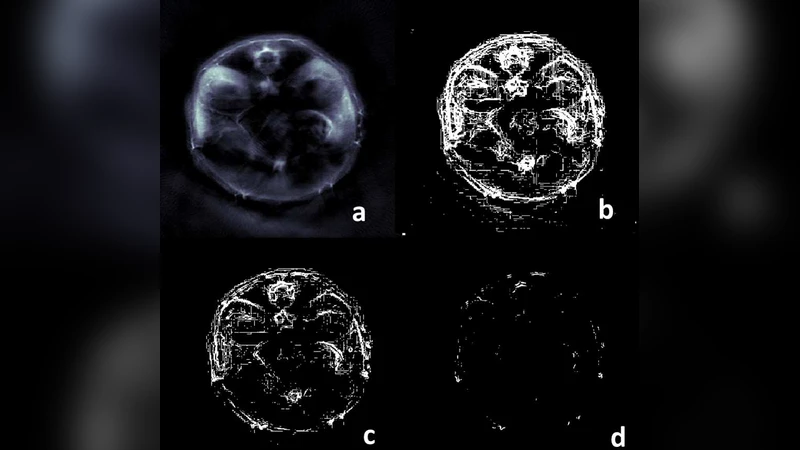
Results demonstrate that the multiscale edge‑flow combined with scale‑dependent morphology markedly outperforms a single‑scale Sobel edge map. The multiscale approach captures fine‑scale edges that are less affected by noise, while the morphological step effectively removes spurious edges beyond the tissue boundary. Visual comparisons (Figures 1–2) show cleaner, more continuous edge maps and well‑fitted ellipses that closely follow the true organ contours. Quantitatively, the Dice coefficient improves from ~0.65 (Sobel) to >0.90 with the proposed method. The authors also observe that most of the performance gain is achieved at scale 2; deeper scales (scale 3) provide marginal benefit but increase computational load, suggesting that a depth of two scales offers an optimal trade‑off for routine OAT studies.
The study’s contributions are threefold: (1) a robust multiscale edge detection framework that leverages vector‑flow consistency across scales; (2) an adaptive morphological cleaning procedure that eliminates noise without sacrificing true edges; and (3) a simple yet effective parametric shape fitting strategy that yields high‑quality segmentations quantified by standard overlap metrics. Limitations include reduced performance in regions with highly irregular geometry (e.g., kidney/spleen) where a single ellipse may not capture complex internal structures, and the need for further acceleration to enable real‑time processing. Future work could explore multi‑ellipse or spline‑based models for intricate anatomies and implement GPU‑accelerated versions of the algorithm. Overall, the paper presents a significant advancement in automated OAT image analysis, facilitating more reliable quantitative studies in preclinical biomedical research.
Comments & Academic Discussion
Loading comments...
Leave a Comment