Relevant Word Order Vectorization for Improved Natural Language Processing in Electronic Healthcare Records
Objective: Electronic health records (EHR) represent a rich resource for conducting observational studies, supporting clinical trials, and more. However, much of the relevant information is stored in an unstructured format that makes it difficult to use. Natural language processing approaches that attempt to automatically classify the data depend on vectorization algorithms that impose structure on the text, but these algorithms were not designed for the unique characteristics of EHR. Here, we propose a new algorithm for structuring so-called free-text that may help researchers make better use of EHR. We call this method Relevant Word Order Vectorization (RWOV). Materials and Methods: As a proof-of-concept, we attempted to classify the hormone receptor status of breast cancer patients treated at the University of Kansas Medical Center during a recent year, from the unstructured text of pathology reports. Our approach attempts to account for the semi-structured way that healthcare providers often enter information. We compared this approach to the ngrams and word2vec methods. Results: Our approach resulted in the most consistently high accuracy, as measured by F1 score and area under the receiver operating characteristic curve (AUC). Discussion: Our results suggest that methods of structuring free text that take into account its context may show better performance, and that our approach is promising. Conclusion: By using a method that accounts for the fact that healthcare providers tend to use certain key words repetitively and that the order of these key words is important, we showed improved performance over methods that do not.
💡 Research Summary
Electronic health records (EHR) contain a wealth of clinical information, but most of it resides in free‑text notes that are difficult to mine automatically. Conventional natural‑language‑processing pipelines typically rely on generic vectorization techniques such as n‑grams, TF‑IDF, or word2vec, which do not exploit the semi‑structured, repetitive nature of medical documentation. In this study the authors introduce Relevant Word Order Vectorization (RWOV), a novel approach that builds a document‑by‑word matrix centered on a “term of interest” (TOI) – for example, the hormone‑receptor label in a pathology report. First, the most frequent words that co‑occur with the TOI across the whole corpus are identified (the “top words”). For each patient document the algorithm locates the sentence containing the TOI, then computes, for every top word, the number of other top words that appear between it and the TOI. The cell value is the signed inverse of this distance (sign indicates whether the top word precedes or follows the TOI, and a value of 0 is assigned when the top word is absent). Thus each row encodes not only which important words are present but also how close they are to the TOI, capturing order information that is often crucial in medical narratives (e.g., “ER positive, PR negative”).
The method has a single hyper‑parameter – the number of top words – which the authors tuned on an independent dataset and then fixed for all experiments. To evaluate RWOV they used pathology reports from the University of Kansas Medical Center for women with breast cancer, labeling each report for estrogen receptor (ER), progesterone receptor (PR), and HER2 status. The dataset comprised 293 ER, 274 PR, and 94 HER2 reports, with a marked class imbalance (e.g., only 14 HER2‑positive cases). For each biomarker a binary classifier was trained using either a support‑vector machine (SVM) or a multilayer perceptron (MLP). RWOV’s performance was compared against two widely used vectorizers: (1) n‑grams (various ranges from unigrams to trigrams) with TF‑IDF weighting, and (2) word2vec skip‑gram embeddings (size 200, window 6, negative 5, min‑count 3). Both baseline methods were paired with the same SVM and MLP classifiers, and all models were evaluated with three‑fold cross‑validation. The primary metrics were F1‑score (to balance precision and recall, especially important under class imbalance) and area under the ROC curve (AUC).
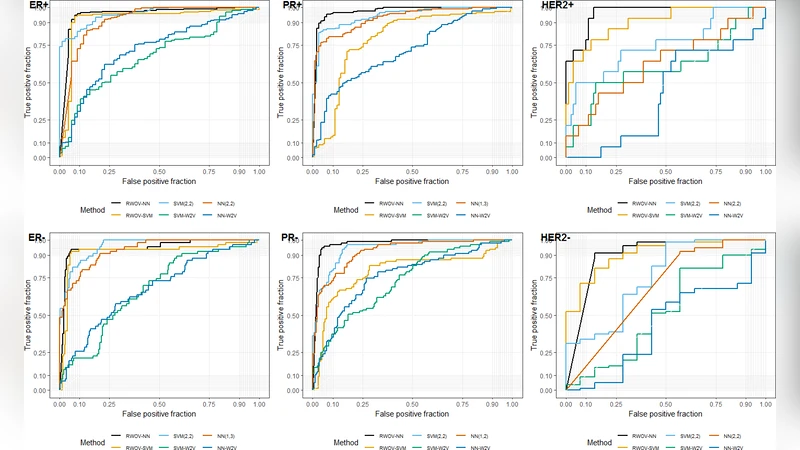
Across all three biomarkers and both classifiers, RWOV achieved the highest F1 and AUC values. For the minority HER2‑positive class, RWOV‑NN reached an F1 of 0.71 and an AUC of 0.96, far surpassing the best n‑gram (F1 ≈ 0.35, AUC ≈ 0.75) and word2vec (F1 ≈ 0.16, AUC ≈ 0.60) results. Even for the majority classes (ER+, PR+), RWOV maintained F1 scores above 0.90, while baseline methods showed modest declines when the class was under‑represented. Bootstrap confidence intervals confirmed that the superiority of RWOV was statistically significant in every scenario.
The authors attribute RWOV’s advantage to its explicit modeling of word order and proximity to the TOI, which aligns with the way clinicians document findings (e.g., “positive for ER, negative for PR”). By focusing on a small, high‑frequency vocabulary and weighting words by their distance from the target, RWOV amplifies the signal from sparse but informative cues, especially beneficial for rare classes. Moreover, the approach is computationally simple, requiring only tokenization, stemming, and a single matrix construction step.
Nevertheless, the study has limitations. The data come from a single institution and a single year, raising concerns about external validity. The top‑word selection is purely frequency‑based, so synonyms, abbreviations, or domain‑specific jargon that appear less often may be missed. The distance‑based weighting does not explicitly handle negation or more complex linguistic constructs (e.g., “no evidence of HER2 overexpression”). Finally, the method was evaluated only on binary classification of hormone‑receptor status; its applicability to multi‑label or more nuanced extraction tasks remains to be demonstrated.
Future work suggested includes testing RWOV on multi‑institutional, multilingual corpora, integrating dynamic top‑word selection (e.g., TF‑IDF or embedding‑driven), and extending the weighting scheme to capture negation and other contextual cues. Combining RWOV with richer contextual embeddings (e.g., BERT) could further boost performance while preserving the interpretability of the distance‑based features.
In conclusion, Relevant Word Order Vectorization offers a tailored, low‑complexity solution for structuring free‑text EHR data. By leveraging the repetitive vocabulary and the positional relationship of key terms, RWOV consistently outperforms standard n‑gram and word2vec pipelines, especially in scenarios with severe class imbalance. This work underscores the value of domain‑specific vectorization strategies for unlocking the clinical insights hidden in unstructured health records.
Comments & Academic Discussion
Loading comments...
Leave a Comment