Learning Individualized Cardiovascular Responses from Large-scale Wearable Sensors Data
We consider the problem of modeling cardiovascular responses to physical activity and sleep changes captured by wearable sensors in free living conditions. We use an attentional convolutional neural network to learn parsimonious signatures of individual cardiovascular response from data recorded at the minute level resolution over several months on a cohort of 80k people. We demonstrate internal validity by showing that signatures generated on an individual’s 2017 data generalize to predict minute-level heart rate from physical activity and sleep for the same individual in 2018, outperforming several time-series forecasting baselines. We also show external validity demonstrating that signatures outperform plain resting heart rate (RHR) in predicting variables associated with cardiovascular functions, such as age and Body Mass Index (BMI). We believe that the computed cardiovascular signatures have utility in monitoring cardiovascular health over time, including detecting abnormalities and quantifying recovery from acute events.
💡 Research Summary
The paper tackles the problem of modeling how an individual’s heart rate responds to everyday physical activity and sleep, using massive wearable‑sensor data collected at one‑minute resolution. The authors assembled a cohort of 80,137 users of commercial activity trackers (Fitbit, Apple Watch, etc.) who consented to share step counts, sleep stage flags, and average heart‑rate streams for the full months of January 2017 and January 2018. After basic preprocessing—per‑person z‑scoring of heart rate, log‑transform and scaling of step counts, binary encoding of two sleep states, and simple mean‑imputation for missing minutes—the data were split into training (≈44 k users), tuning (≈11 k) and validation (≈25 k) sets.
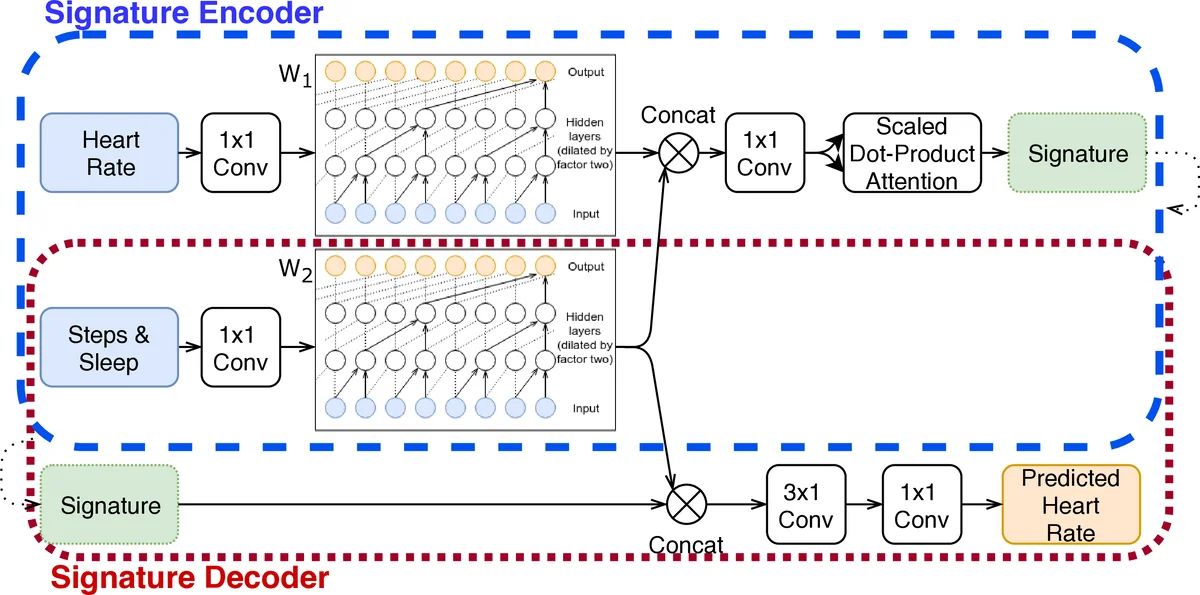
The core contribution is a “cardiovascular signature network,” an auto‑encoder that learns a compact, fixed‑size vector (the signature) summarizing an individual’s cardio‑vascular transfer function. The encoder consists of two parallel WaveNet‑style dilated causal convolution blocks (W₁ for activity, W₂ for heart rate), each with seven layers, residual connections, and 32/16 filters respectively, giving a receptive field of up to 128 minutes. After concatenating the two block outputs, a scaled‑dot‑product attention layer (query/key dimension = 8, value dimension = signature size) aggregates temporal information into the signature. The decoder re‑uses the same W₂ weights (weight‑tying), concatenates the learned signature with the activity representation at each time step, and passes the result through two additional 1‑D convolutions to predict the minute‑wise heart rate. Training minimizes the mean L₂ error using Adam (α = 0.001) on mini‑batches of 16 for up to 30 epochs with early stopping.
Baseline comparisons include (1) a naïve mean‑heart‑rate predictor split by wake/sleep, (2) a per‑person XGBoost model that uses the previous 120 minutes of activity to predict heart rate, and (3) a population‑level XGBoost model trained on all users. The proposed model consistently outperforms these baselines: validation RMSE drops from ~0.38–0.40 (baselines) to ~0.28–0.30 for the signature model. A sensitivity analysis shows that signature dimensionality beyond 16 yields diminishing returns, and that using the full training set improves performance by roughly 14 % compared with using only 1 % of the data.
Internal validity (test‑retest reliability) is demonstrated by training signatures on each individual’s 2017 data and applying them to predict 2018 heart rates. A person’s own signature yields a mean‑square error 60 % lower than a randomly chosen other person’s signature (Wilcoxon signed‑rank, p < 10⁻¹⁶). External validity is shown by using the learned signatures as features in an XGBoost classifier to predict whether a user is above or below the cohort median age (31 years) and whether they are obese (BMI ≥ 30). The signatures achieve AUROC ≈ 70 % for both tasks, substantially higher than using only resting heart rate (AUROC ≈ 60 % for age, 54 % for obesity). This indicates that the signatures capture richer information about the dynamic relationship between activity and heart rate than static measures.
The authors discuss failure modes: short observation windows, high missingness, erratic behavior, or unmeasured confounders (stress, medication, meals) can degrade signature quality. They illustrate a case where heart‑rate spikes occur without corresponding activity, likely reflecting anxiety. Future work will explore the motifs highlighted by the attention mechanism, relate them to health outcomes, and experiment with variational auto‑encoders to regularize the latent space. Additional hyper‑parameter tuning and architectural refinements are also planned.
In summary, the study demonstrates that large‑scale, real‑world wearable data can be leveraged to learn individualized, parsimonious representations of cardiovascular response. These “cardiovascular signatures” not only improve minute‑level heart‑rate forecasting over strong baselines but also serve as informative biomarkers for demographic and health‑related traits. The approach opens avenues for unobtrusive, continuous cardiovascular monitoring, early detection of abnormalities, and quantitative assessment of recovery after acute events.
Comments & Academic Discussion
Loading comments...
Leave a Comment