A New Approach to Privacy-Preserving Clinical Decision Support Systems
Background: Clinical decision support systems (CDSS) are a category of health information technologies that can assist clinicians to choose optimal treatments. These support systems are based on clinical trials and expert knowledge; however, the amount of data available to these systems is limited. For this reason, CDSSs could be significantly improved by using the knowledge obtained by treating patients. This knowledge is mainly contained in patient records, whose usage is restricted due to privacy and confidentiality constraints. Methods: A treatment effectiveness measure, containing valuable information for treatment prescription, was defined and a method to extract this measure from patient records was developed. This method uses an advanced cryptographic technology, known as secure Multiparty Computation (henceforth referred to as MPC), to preserve the privacy of the patient records and the confidentiality of the clinicians’ decisions. Results: Our solution enables to compute the effectiveness measure of a treatment based on patient records, while preserving privacy. Moreover, clinicians are not burdened with the computational and communication costs introduced by the privacy-preserving techniques that are used. Our system is able to compute the effectiveness of 100 treatments for a specific patient in less than 24 minutes, querying a database containing 20,000 patient records. Conclusion: This paper presents a novel and efficient clinical decision support system, that harnesses the potential and insights acquired from treatment data, while preserving the privacy of patient records and the confidentiality of clinician decisions.
💡 Research Summary
The paper addresses a fundamental limitation of clinical decision support systems (CDSS): the scarcity of high‑quality, patient‑derived data due to strict privacy regulations. Focusing on HIV‑1 treatment, the authors define a treatment‑effectiveness metric—time‑to‑treatment‑failure (TTF)—which measures the number of days from therapy initiation to a failure event (switch, discontinuation, or death). To make this metric useful for a new patient, the system must retrieve the average TTF of past patients whose viral genotypes are “similar” to the current case.
Similarity is operationalized by encoding each patient’s viral genotype as a fixed‑length bit‑string (≈100–200 bits) where each bit indicates the presence of a known resistance‑associated mutation. The Hamming distance between two bit‑strings serves as a simple similarity measure; patients whose distance is below a pre‑defined threshold θ are considered similar. Although this representation is a simplification of the sophisticated sequence‑based or neural‑network distance metrics used in practice, it suffices as a proof‑of‑concept and can be replaced by richer models without altering the overall architecture.
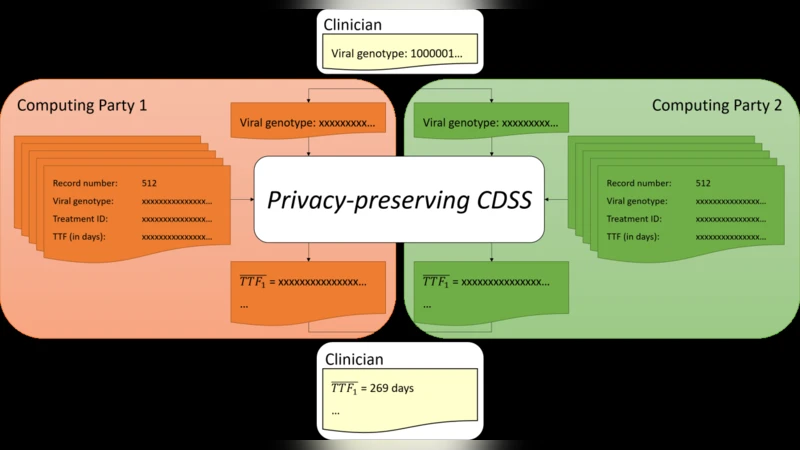
The core technical contribution is the integration of secure multiparty computation (MPC) to compute the average TTF over the selected similar patients while keeping all raw data (genotypes, treatments, outcomes) confidential. The authors adopt the SPDZ protocol, a state‑of‑the‑art MPC framework that separates an offline phase (generation of authenticated MACs) from an online phase (fast arithmetic on secret‑shared values). Inputs are secret‑shared among a set of computing parties (e.g., hospitals, research institutes). During the online phase, the parties jointly evaluate the function “average TTF of patients with Hamming distance < θ” using only the shares; the final result is reconstructed and revealed to the querying clinician.
Key design choices include:
- Security model – The primary analysis assumes semi‑honest participants (they follow the protocol but may try to infer others’ data). The protocol can be hardened to a fully malicious model by adding consistency checks, at the cost of extra communication.
- Client‑side simplicity – Clinicians submit only their query (genotype, treatment option) and receive the result; they do not perform any cryptographic computation, making the solution practical for real‑world clinical workflows.
- Scalability – In experiments, a database of 20 000 patient records was processed to evaluate 100 treatment options in under 24 minutes. This performance is enabled by the efficiency of SPDZ’s online phase and the lightweight Hamming‑distance computation on secret‑shared bits.
The paper situates its contribution within a broader landscape of privacy‑preserving health analytics, citing prior work on MPC for genomic studies, secure data mining, and federated health statistics. It argues that many existing CDSSs cannot exploit the full richness of patient data because viral genotypes are quasi‑identifiers; even a single genotype can uniquely identify a patient when combined with external information. By keeping the genotype and outcome data secret‑shared, the proposed system eliminates this risk while still delivering actionable insights.
Limitations are acknowledged. The simplified genotype representation may miss subtle resistance patterns captured by more advanced bioinformatics tools. The semi‑honest assumption may be insufficient for adversarial settings where a party could deviate from the protocol. Moreover, the current implementation focuses on a single effectiveness metric; extending to multi‑objective evaluation (e.g., side‑effects, cost) would require additional secure computations.
Future work outlined includes: (1) integrating richer similarity metrics (sequence alignment, machine‑learning embeddings); (2) scaling to larger, multi‑institution consortia with potentially hundreds of parties; (3) transitioning to a fully malicious security model; (4) embedding the system into clinical workflows and evaluating usability with clinicians.
In conclusion, the study demonstrates that secure MPC, specifically the SPDZ framework, can be practically applied to CDSSs to unlock the value of sensitive patient data without compromising privacy. The prototype for HIV‑1 treatment recommendation shows that clinically relevant queries can be answered within minutes, paving the way for broader adoption of privacy‑preserving analytics in personalized medicine.
Comments & Academic Discussion
Loading comments...
Leave a Comment