Modularity in biological evolution and evolutionary computation
One of the main properties of biological systems is modularity, which manifests itself at all levels of their organization, starting with the level of molecular genetics, ending with the level of whole organisms and their communities. In a simplified form, these basic principles were transferred from the genetics of populations to the field of evolutionary computations, in order to solve applied optimization problems. Over almost half a century of development in this field of computer science, considerable practical experience has been gained and interesting theoretical results have been obtained. In this survey, the phenomena and patterns associated with modularity in genetics and evolutionary computations are compared. An analysis of similarities and differences in the results obtained in these areas is carried out from the modularity view point. The possibilities for knowledge transfer between the areas are discussed.
💡 Research Summary
The paper presents a comprehensive survey that juxtaposes the concept of modularity as it appears in biological evolution with its adoption in the field of evolutionary computation (EC). Beginning with an overview of modularity in biology, the authors delineate four hierarchical levels: genetic modules (clusters of genes, regulatory motifs), developmental modules (tissues, organs, developmental pathways), ecological/social modules (populations, interaction networks), and community-level modules (species assemblages). They argue that modular organization confers evolutionary advantages by preserving co‑adapted gene complexes, enabling reuse of functional sub‑units, and facilitating rapid phenotypic innovation through recombination of whole modules rather than isolated mutations. The classic “building‑block” hypothesis and the concept of developmental reuse are highlighted as central mechanisms that drive both robustness and evolvability.
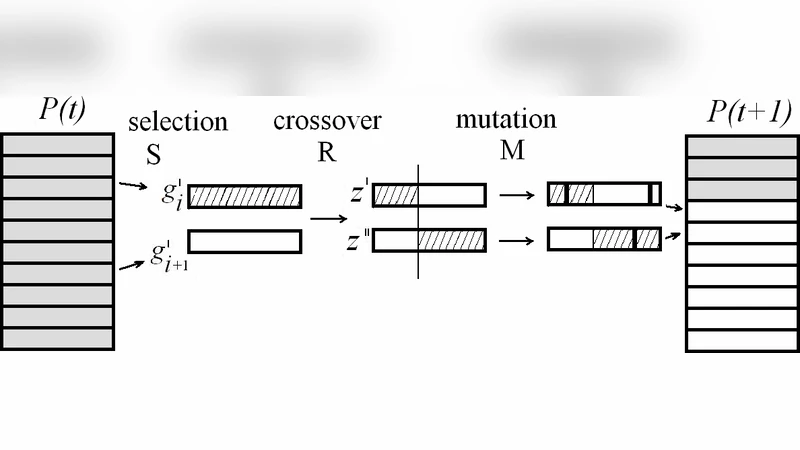
Transitioning to EC, the survey maps these biological principles onto algorithmic design. Traditional genetic algorithms (GAs) treat chromosomes as flat strings, but modular EC approaches identify and preserve substructures—sub‑trees in genetic programming, sub‑graphs in graph‑based encodings, or “building blocks” in binary strings. The paper reviews a variety of techniques for module detection (clustering, community detection, motif mining) and for module‑aware variation operators such as modular crossover, module‑specific mutation, and hierarchical recombination. It also discusses multi‑objective formulations that explicitly reward modularity, as well as diversity‑preserving mechanisms that maintain a repertoire of distinct modules throughout the run.
Empirical evidence is synthesized from a broad set of benchmark problems, including NK‑landscapes, traveling salesman problems, robot controller synthesis, and neural architecture search. Across these domains, modular EC consistently outperforms non‑modular baselines in terms of convergence speed, final fitness, and solution interpretability. The authors attribute these gains to reduced epistatic interactions (the “overlap” problem) and to the effective partitioning of the search space, which mirrors the biological advantage of recombining whole functional units.
On the theoretical side, the survey examines models that quantify the impact of modularity on algorithmic performance. Using Markov chain analyses and “divide‑and‑conquer” frameworks, the authors show that the expected time to reach optimal solutions can drop from polynomial to logarithmic order when modular structures are present and correctly exploited. However, they acknowledge that existing models often assume static modules, whereas biological systems exhibit dynamic re‑configuration in response to environmental change—a gap that remains largely unaddressed in EC theory.
The final sections explore bidirectional knowledge transfer. Biological data—such as gene‑expression correlation networks, protein‑protein interaction maps, and developmental gene regulatory circuits—can be mined to automatically generate modular encodings for EC, providing problem‑specific priors that accelerate search. Conversely, algorithmic innovations like adaptive modular crossover, hierarchical population structures, and modular fitness shaping have potential applications in synthetic biology, where engineered genetic circuits could benefit from modular design principles to improve reliability and evolvability.
The authors conclude with a research agenda that emphasizes (1) dynamic modularity models that capture environmental feedback, (2) explicit modeling of epistatic (non‑linear) interactions between modules, and (3) integrated platforms that combine large‑scale biological network analysis with modular EC frameworks. By highlighting the parallels and divergences between natural evolution and its computational analogues, the paper argues that modularity serves as a powerful conceptual bridge, and that deeper cross‑disciplinary collaboration will likely yield novel algorithms and a richer understanding of evolutionary processes.
Comments & Academic Discussion
Loading comments...
Leave a Comment