Dimensionality Reduction in Deep Learning for Chest X-Ray Analysis of Lung Cancer
Efficiency of some dimensionality reduction techniques, like lung segmentation, bone shadow exclusion, and t-distributed stochastic neighbor embedding (t-SNE) for exclusion of outliers, is estimated for analysis of chest X-ray (CXR) 2D images by deep learning approach to help radiologists identify marks of lung cancer in CXR. Training and validation of the simple convolutional neural network (CNN) was performed on the open JSRT dataset (dataset #01), the JSRT after bone shadow exclusion - BSE-JSRT (dataset #02), JSRT after lung segmentation (dataset #03), BSE-JSRT after lung segmentation (dataset #04), and segmented BSE-JSRT after exclusion of outliers by t-SNE method (dataset #05). The results demonstrate that the pre-processed dataset obtained after lung segmentation, bone shadow exclusion, and filtering out the outliers by t-SNE (dataset #05) demonstrates the highest training rate and best accuracy in comparison to the other pre-processed datasets.
💡 Research Summary
The paper investigates how dimensionality‑reduction techniques can improve deep‑learning‑based detection of lung‑cancer nodules on chest X‑ray (CXR) images. Using the publicly available Japanese Society of Radiological Technology (JSRT) dataset (247 images, 154 with nodules, 93 without), the authors create five distinct training sets: (1) the original JSRT images (dataset #01); (2) bone‑shadow‑eliminated JSRT (BSE‑JSRT, dataset #02); (3) lung‑segmented JSRT (dataset #03); (4) BSE‑JSRT with lung segmentation (dataset #04); and (5) the combination of segmentation, bone‑shadow removal, and outlier filtering via t‑distributed stochastic neighbor embedding (t‑SNE) (dataset #05).
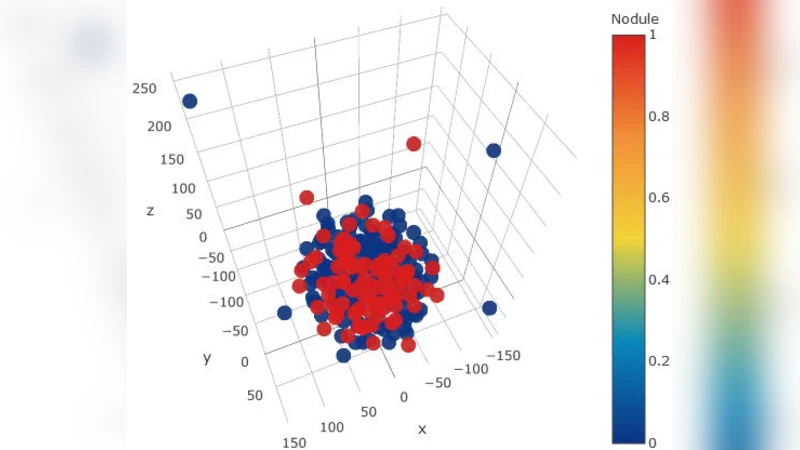
The first two preprocessing steps act as explicit dimensionality reduction. Lung segmentation retains only the lung fields, reducing the pixel‑wise feature space by an average factor of 3.2 ± 0.7 (mask area ≈ 0.32 ± 0.06 of the image). Bone‑shadow removal eliminates rib and clavicle shadows, providing an additional reduction of roughly two‑fold (mask area ≈ 0.15 ± 0.04). The third step uses t‑SNE to embed high‑dimensional lung‑mask images into a 3‑D space, visually separating typical masks from atypical ones. By inspecting the embedding, the authors identify and discard the most dissimilar 5 % of masks, which correspond either to intrinsically small lungs or to manual segmentation errors (e.g., heart tissue left in the lung field).
All five datasets are fed into the same simple convolutional neural network (CNN) architecture previously described in reference
Comments & Academic Discussion
Loading comments...
Leave a Comment