NIPS4Bplus: a richly annotated birdsong audio dataset
Recent advances in birdsong detection and classification have approached a limit due to the lack of fully annotated recordings. In this paper, we present NIPS4Bplus, the first richly annotated birdsong audio dataset, that is comprised of recordings c…
Authors: Veronica Morfi, Yves Bas, Hanna Pamu{l}a
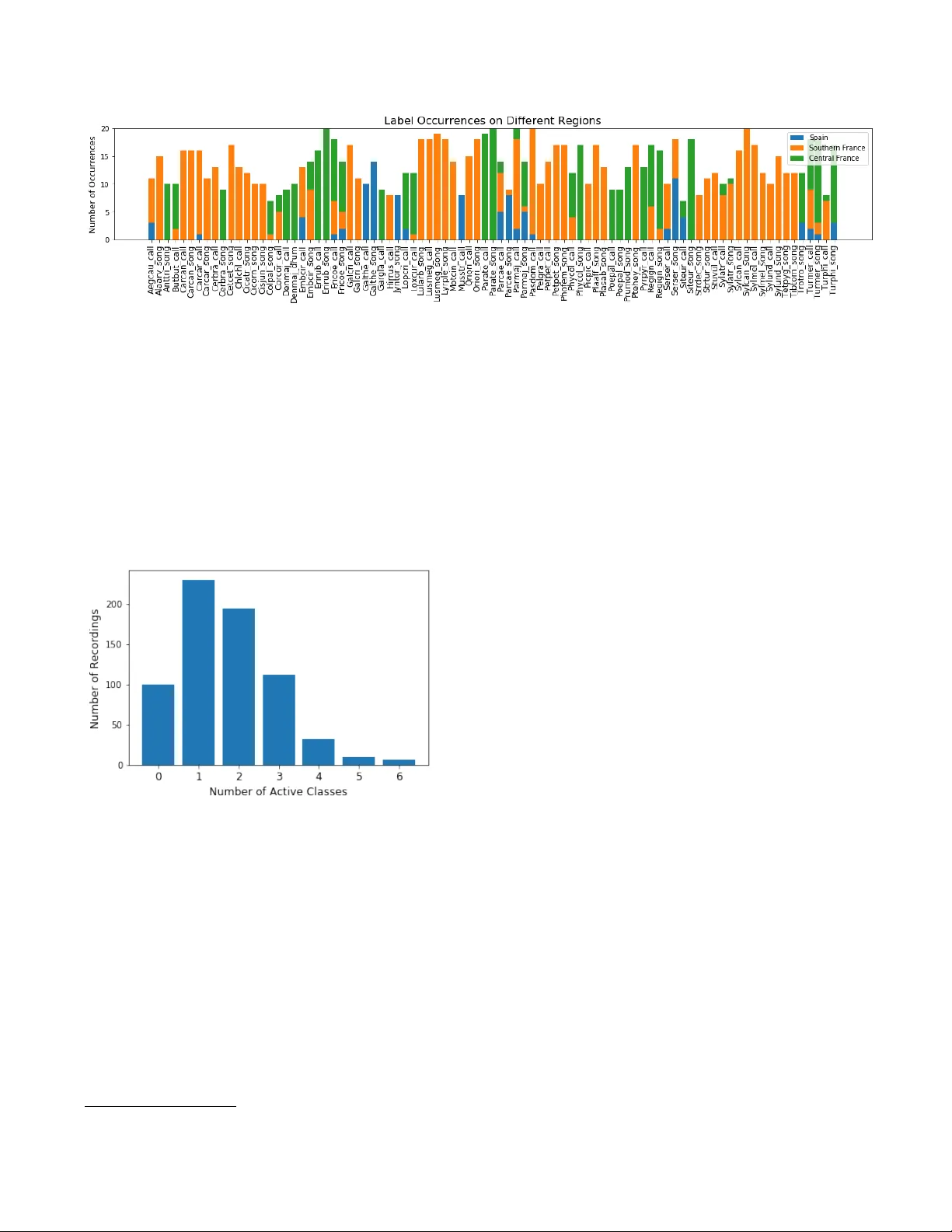
NIPS4BPLUS: A RICHL Y ANNO T A TED BIRDSONG A UDIO D A T ASET V er onica Morfi ? Yves Bas † , ‡ Hanna P amuła †† Herv ´ e Glotin ‡‡ Dan Stowell ? ? Machine Listening Lab, Centre for Digital Music (C4DM), Queen Mary Uni v . of London, UK † Centre d’Ecologie et des Sciences de la Conserv ation (CESCO), Mus ´ eum National d’Histoire Naturelle, CNRS, Sorbonne Uni v ., Paris, France ‡ Centre d’Ecologie Fonctionnelle et Ev olutiv e (CEFE), CNRS, Univ . de Montpellier , Uni v . Paul-V al ´ ery Montpellier , France †† A GH Uni v . of Science and T echnology , Department of Mechanics and V ibroacoustics, Krak ´ ow , Poland ‡‡ Uni v . T oulon, Aix Marseille Univ ., CNRS, LIS, D YNI team, SABIOD, Marseille, France ABSTRA CT Recent adv ances in birdsong detection and classification hav e approached a limit due to the lack of fully annotated record- ings. In this paper , we present NIPS4Bplus, the first richly annotated birdsong audio dataset, that is comprised of record- ings containing bird vocalisations along with their acti ve species tags plus the temporal annotations acquired for them. Statistical information about the recordings, their species spe- cific tags and their temporal annotations are presented along with example uses. NIPS4Bplus could be used in various ecoacoustic tasks, such as training models for bird popula- tion monitoring, species classification, birdsong vocalisation detection and classification. Index T erms — audio dataset, ecosystems, bird vocalisa- tions, rich annotations, ecoacoustics 1. INTR ODUCTION The potential applications of automatic species detection and classification of birds from their sounds are man y (e.g. eco- logical research, biodi versity monitoring, archiv al) [1, 2, 3, 4, 5]. In recent decades, there has been an increasing amount of ecological audio datasets that hav e tags assigned to them to indicate the presence or not of a specific bird species. Utilis- ing these datasets and the provided tags, many authors ha ve proposed methods for bird audio detection [6, 7] and bird species classification, e.g. in the context of LifeCLEF clas- sification challenges [8, 9] and more [10, 11]. Howe ver , these methods do not predict any information about the temporal location of each e vent or the number of its occurrences in a recording. W e thank EADM MaDICS CNRS and SABIOD MI CNRS for support- ing the NIPS4B challenge, Sylvain V igant for pro viding recordings from Central France, and BIOTOPE for making the data public for the NIPS4B 2013 bird classification challenge. Some research has been made into using audio tags in or - der to predict temporal annotations, labels that contain tem- poral information about the audio events. In [12, 13], the au- thors try to exploit audio tags in birdsong detection and bird species classification, in [14], the authors use deep networks to tag the temporal location of active bird vocalisations, while in [15], the authors propose a bioacoustic segmentation based on the Hierarchical Dirichlet Process (HDP-HMM) to infer song units in birdsong recordings. Furthermore, some meth- ods for temporal predictions by using tags have been proposed for other types of general audio [16, 17, 18]. Howe ver , in all the abo ve cases some kind of temporal annotations were used in order to ev aluate the performance of the methods. Annotating ecological data with temporal annotations to train sound event detectors and classifiers is a time consum- ing task in volving a lot of manual labour and expert annota- tors. There is a high diversity of animal v ocalisations, both in the types of the basic syllables and in the way they are com- bined [19, 20]. Also, there is noise present in most habitats, and many bird communities contain multiple bird species that can potentially ha ve overlapping vocalizations [21, 22, 23]. These factors mak e detailed annotations laborious to gather, while on the other hand acquiring audio tags takes much less time and ef fort, since the annotator has to only mark the activ e sound ev ent classes in a recording and not their exact bound- aries. This means that man y ecological datasets lack temporal annotations of bird vocalisations ev en though the y are vital to the training of automated methods that predict the temporal annotations which could potentially solve the issue of need- ing a human annotator . Recently , BirdV ox-full-night [24], a dataset contain- ing some temporal and frequency information about flight calls of nocturnally migrating birds, was released. Ho wev er, BirdV ox-full-night only focuses on avian flight calls, a spe- cific type of bird calls, that usually ha ve a v ery short duration in time. The temporal annotations pro vided for them don’t include any onset, offset or information about the duration of the calls, they simply contain a single time marker at which the flight call is activ e. Additionally , there is no distinction between the different bird species, hence no specific species annotations are provided, but only the presence of flight calls through the duration of a recording is denoted. Hence, the dataset can provide data to train models for flight call de- tection but is not ef ficient for models performing both event detection and classification for a variety of bird v ocalisations. In this paper , we introduce NIPS4Bplus, the first ecologi- cal audio dataset that contains bird species tags and temporal annotations [25], and can be used for training supervised au- tomated methods that perform bird v ocalisation detection and classification and can also be used for ev aluating methods that use only audio tags or no annotations for training. The rest of the paper is structured as follo ws: Section 2 describes the pro- cess of collecting and selecting the recordings comprising the dataset, Section 3 presents our approach of acquiring the tags and temporal annotations and provides statistical information about the labels and recordings comprising the dataset fol- lowed by e xample uses of NIPS4Bplus, with the conclusion in Section 4. 2. A UDIO D A T A COLLECTION The recordings that comprise the Neural Information Pro- cessing Scaled for Bioacoustics (NIPS4B) 2013 training and testing dataset were collected by recorders placed in 39 dif- ferent locations, which can be summarised by 7 regions in France and Spain, as depicted in Fig. 1. 20% of the record- ings were collected from the Haute-Loire region in Central France, 65% of them were collected from the Pyr ´ en ´ ees- Orientales, Aude and H ´ erault regions in south-central France along the Mediterranean cost and the remaining 15% of the recordings originated from the Granada, Ja ´ en and Almeria regions in eastern Andalusia, Spain. The Haute-Loire area is a more hilly and cold region, while the rest of the re gions are mostly along the Mediterranean coast and hav e a more Mediterranean climate. The recorders used to acquire the recordings were the SM2B A T using SMX-US microphones. They were originally put in the field for bat echolocation call sampling, b ut they were also set to record for 3 hours single channel at 44.1 kHz sampling rate starting 30 minutes after sunrise, right after bat sampling. The recorders were set to a 6 dB Signal-to- Noise Ratio (SNR) trigger with a window of 2 seconds, and acquired recordings only when the trigger was acti vated. Approximately 30 hours of field recordings were col- lected. Any recording longer than 5 seconds was split into multiple 5 second files. SonoChiro, a chirp detection tool used for bat vocalisation detection, was used on each file to identify recordings with bird v ocalisations. 1 A stratified random sampling was then applied to all acquired recordings, 1 http://www.leclub- biotope.com/fr/72- sonochiro based on locations and clustering of features, to maximise the div ersity in the labelled dataset, resulting in nearly 5000 files being chosen. Follo wing the first stage of selection, manual annotations were produced for the classes activ e in these 5000 files and any recordings that contained unidentified species’ vocalisations were discarded. Furthermore, the training set and testing set recordings were allocated so that the same species were active in both. Finally , for training purposes, only species that could be cov ered by at least 7 recordings in the training set were included in the final dataset, the rest were considered rare species’ occurrences that would make it hard to train any classifier , hence were discarded. The final training and testing set consist of 687 files of total duration of less than an hour , and 1000 files of total duration of nearly two hours, respecti vely . Fig. 1 . Re gions where the dataset recordings were collected from. Green indicates Central France region Haute-Loire. Or- ange indicates Southern France regions Pyr ´ en ´ ees-Orientales, Aude and H ´ erault. Blue indicates Southern Spain re gions Granada, Ja ´ en and Almeria. 3. ANNO T A TIONS 3.1. T ags The labels for the species acti ve in each recording of the train- ing set were initially created for the NIPS4B 2013 bird song classification challenge [26]. There is a total of 61 different bird species active in the dataset. For some species we dis- criminate the song from the call and from the drum. W e also include some species li ving with these birds: 7 insects and an amphibian. This tagging process resulted in 87 classes. A de- tailed list of the class names and their corresponding species English and scientific names can be found in [25]. These tags only pro vide information about the species activ e in a record- ing and do not include any temporal information. In addition to the recordings containing bird v ocalisations, some training files only contain background noise acquired from the same regions and hav e no bird song in them, these files can be used to tune a model during training. Fig. 2 depicts the number of occurrences per class for recordings collected in each of the 3 Fig. 2 . Number of occurrences of each sound type in recordings collected from Spain, Southern France and Central France. different general regions of Spain, South France and Central France. Each tag is represented by at least 7 up to a maximum of 20 recordings. Each recording that contains bird vocalisations includes 1 to 6 individual labels. These files may contain different vocalisations from the same species and also may contain a variety of other species that vocalise along with this species. Fig. 3 depicts the distribution of the number of active classes in the dataset. Fig. 3 . Distribution of number of active classes in dataset recordings. 3.2. T emporal Annotations T emporal annotations for each recording in the training set of the NIPS4B dataset were produced manually using Sonic V isualiser . 2 The temporal annotations were made by a single annotator , Hanna Pamuła, and can be found in [25]. T able 1 presents the temporal annotation format as is provided in NIPS4Bplus and Fig. 4 depicts the visual representation of the temporal annotations. In concern to the temporal annotations for the dataset, we should mention the following: 2 https://www.sonicvisualiser.org/ • The original tags were used for guidance, ho wev er some files were judged to ha ve a dif ferent set of species than the ones giv en in the original metadata. • In a fe w rare occurrences, despite the tags suggesting a bird species acti ve in a recording, the annotator was not able to detect any bird v ocalisation. • An e xtra ‘Unknown’ tag was added to the dataset for vocalisations that could not be classified to a class. • An e xtra ‘Human’ tag was added to a few recordings that have very obvious human sounds, such as speech, present in them. • Out of the 687 recordings of the training set 100 record- ings contain only background noise, hence no temporal annotations were needed for them. • Of the remaining 587 recordings that contain vocali- sations, 6 could not be unambiguously labelled due to hard to identify v ocalisations, thus no temporal annota- tion files were produced for them. • An annotation file for any recording containing mul- tiple insects does not differentiate between the insect species and the ‘Unkno wn’ label was given to all insect species present. • In the rare case where no birds were active along with the insects no annotation file was pro vided. Hence, 7 recordings containing only insects were left unlabelled. • In total, 13 recordings ha ve no temporal annotation files. These can be used when training a model that does not use temporal annotations. • On some occasions, the different syllables of a song were separated in time into different e vents while in other occasions they were summarised into a larger ev ent, according to the judgement of the expert anno- tator . This v ariety could help train an unbiased model regarding separating ev ents or grouping them together as one continuous time ev ent. As mentioned above, each recording may contain multiple species vocalising at the same time. This can often occur in wildlife recordings and is important to be taken into account Fig. 4 . Mel-band spectrogram of a recording in NIPS4Bplus and the visual representation of the corresponding temporal annotations as noted in T able 1. T able 1 . NIPS4Bplus temporal annotations of the recording depicted in Fig. 4 Starting Time (sec) Duration (sec) T ag 0.00 0.37 Serser call 0.00 2.62 Ptehey song 1.77 0.06 Carcar call 1.86 0.07 Carcar call 2.02 0.41 Serser call 3.87 1.09 Ptehey song when training a model. Fig. 5 presents the fraction of the total duration containing overlapping vocalisations as well as the number of simultaneously occurring classes. A few examples of the NIPS4Bplus dataset and temporal annotations being used can be found in [27] and [28]. First, in [27], we use NIPS4Bplus to carry out the training and ev alu- ation of a newly proposed multi-instance learning (MIL) loss function for audio ev ent detection. And in [28], we com- bine the proposed method of [27] and a network trained on the NIPS4Bplus tags that performs audio tagging in a multi- task learning (MTL) setting. Additional applications using NIPS4Bplus could include training models for bird species audio ev ent detection and classification, ev aluating how gen- eralisable of method trained on a different set of data is, and many more. Fig. 5 . Distribution of simultaneous number of activ e classes on the total duration of the recordings. 4. CONCLUSION In this paper, we present NIPS4Bplus, the first richly anno- tated birdsong audio dataset. NIPS4Bplus is comprised of the NIPS4B dataset and tags used for the 2013 bird song clas- sification challenge plus the ne wly acquired temporal annota- tions. W e provide statistical information about the recordings, their species specific tags and their temporal annotations. 5. REFERENCES [1] D. K. Dawson and M. G. Efford, “Bird population den- sity estimated from acoustic signals, ” Journal of Applied Ecology , v ol. 46, no. 6, pp. 1201–1209, 2009. [2] K. T . A. Lambert and P . G. McDonald, “ A low-cost, yet simple and highly repeatable system for acoustically surve ying cryptic species, ” A ustral Ecology , v ol. 39, no. 7, pp. 779–785, 2014. [3] K. L. Drake, M. Frey , D. Hogan, and R. Hedle y , “Us- ing digital recordings and sonogram analysis to obtain counts of yellow rails, ” W ildlife Society Bulletin , v ol. 40, no. 2, pp. 346–354, 2016. [4] S. G. Sovern, E. D. F orsman, G. S. Olson, B. L. Biswell, M. T aylor, and R. G. Anthony , “Barred owls and land- scape attributes influence territory occupancy of north- ern spotted owls, ” The Journal of W ildlife Manag ement , vol. 78, no. 8, pp. 1436–1443, 2014. [5] T iago A. Marques, Len Thomas, Stephen W . Martin, David K. Mellinger , Jessica A. W ard, David J. Moretti, Danielle Harris, and Peter L. T yack, “Estimating animal population density using passiv e acoustics, ” Biolo gical Revie ws , vol. 88, no. 2, pp. 287–309, 2013. [6] S. Adav anne, K. Drossos, E. C ¸ akir , and T . V irtanen, “Stacked con volutional and recurrent neural networks for bird audio detection, ” in 2017 25th Eur opean Sig- nal Pr ocessing Confer ence (EUSIPCO) , Aug 2017, pp. 1729–1733. [7] T . Pellegrini, “Densely connected cnns for bird audio detection, ” in 2017 25th Eur opean Signal Pr ocessing Confer ence (EUSIPCO) , Aug 2017, pp. 1734–1738. [8] H. Go ¨ eau, H. Glotin, W .-P . V ellinga, R. Planqu ´ e, and A. Joly , “LifeCLEF Bird Identification T ask 2016: The arriv al of Deep Learning, ” 2016. [9] H. Go ¨ eau, H. Glotin, W .-P . V ellinga, R. Planqu ´ e, and A. Joly , “LifeCLEF Bird Identification T ask 2017, ” 2017. [10] J. Salamon and J. P . Bello, “Deep con volutional neu- ral networks and data augmentation for en vironmental sound classification, ” IEEE Signal Pr ocessing Letter s , vol. 24, pp. 279–283, 2017. [11] E. Knight, K. C. Hannah, G. J. Foley , C. D. Scott, R. Brigham, and E. Bayne, “Recommendations for acoustic recognizer performance assessment with appli- cation to fiv e common automated signal recognition pro- grams, ” A vian Conservation and Ecology , vol. 12, 11 2017. [12] F . Briggs, B. Lakshminarayanan, L. Neal, X. Fern, R. Raich, S. J. K. Hadle y , A. S. Hadley , and M. G. Betts, “ Acoustic classification of multiple simultaneous bird species: A multi-instance multi-label approach, ” Jour- nal of the Acoustic Society of America , v ol. 131, pp. 4640–4650, 2014. [13] J. F . Ruiz-Mu ˜ noz, Mauricio Orozco-Alzate, and G. Castellanos-Dominguez, “Multiple instance learning-based birdsong classification using unsuper- vised recording segmentation, ” in Pr oceedings of the 24th International Confer ence on Artificial Intelligence . 2015, IJCAI’15, pp. 2632–2638, AAAI Press. [14] L. F anioudakis and I. Potamitis, “Deep networks tag the location of bird vocalisations on audio spectrograms, ” 2017, . [15] V . Roger , M. Bartcus, F . Chamroukhi, and H. Glotin, “Unsupervised bioacoustic segmentation by hierarchical dirichlet process hidden markov model, ” in Multimedia T ools and Applications for En vir onmental & Biodiver- sity Informatics , pp. 113–130. Springer , 2018. [16] J. Schl ¨ uter , “Learning to Pinpoint Singing V oice from W eakly Labeled Examples, ” in Proceedings of the 17th International Society for Music Information Retrie val Confer ence (ISMIR 2016) , New Y ork, USA, 2016. [17] S. Adav anne and T . V irtanen, “Sound event detec- tion using weakly labeled dataset with stacked con volu- tional and recurrent neural network, ” in Proceedings of the Detection and Classification of Acoustic Scenes and Events 2017 W orkshop (DCASE2017) , No vember 2017, pp. 12–16. [18] A. Kumar and B. Raj, “ Audio ev ent detection using weakly labeled data, ” in Pr oceedings of the 2016 ACM on Multimedia Confer ence , New Y ork, NY , USA, 2016, MM ’16, pp. 1038–1047, A CM. [19] T . S. Brandes, “ Automated sound recording and analy- sis techniques for bird surve ys and conservation, ” Bird Conservation International - BIRD CONSER V INT , vol. 18, pp. S163–S173, 09 2008. [20] D.E. Kroodsma, The Singing Life of Bir ds: Audio CD , The Singing Life of Birds: The Art and Sci- ence of Listening to Birdsong. Houghton Mifflin, 2005, isbn:9780618405688 . [21] D. Luther , “Signaller: Receiv er coordination and the timing of communication in amazonian birds, ” Biology Letters , v ol. 4, pp. 651–654, 2008. [22] D. Luther and R. W iley , “Production and perception of communicatory signals in a noisy environment, ” Biol- ogy Letters , v ol. 5, pp. 183–187, 2009. [23] K. Pacifici, T . R. Simons, and K. H. Pollock, “Effects of ve getation and background noise on the detection pro- cess in auditory avian point-count surveys, ” The Auk , vol. 125, no. 4, pp. 998–998, 2008. [24] V . Lostanlen, J. Salamon, A. Farnsworth, S. Kelling, and J. P . Bello, “Birdvox-full-night: a dataset and bench- mark for a vian flight call detection, ” in Pr oc. IEEE ICASSP , April 2018. [25] V . Morfi, D. Stowell, and H. Pamuła, “Transcrip- tions of NIPS4B 2013 Bird Challenge training dataset, ” 2018, doi: https://doi.org/10.6084/m9. figshare.6798548 , Accessed: 29-Oct-2018. [26] Arti ` eres T . Mallat S. Tchernichovski O. Halkias X. Glotin H., LeCun Y ., “Proc. neural information process- ing scaled for bioacoustics, from neurons to big data, ” USA, 2013, NIPS Int. Conf. [27] V . Morfi and D. Stowell, “Data-efficient weakly su- pervised learning for low-resource audio ev ent detec- tion using deep learning, ” 2018, arXiv pr eprint arXiv:1807.06972 . [28] V . Morfi and D. Stowell, “Deep learning for audio e vent detection and tagging on lo w-resource datasets, ” Ap- plied Sciences , vol. 8, no. 8, pp. 1397, Aug 2018.
Original Paper
Loading high-quality paper...
Comments & Academic Discussion
Loading comments...
Leave a Comment