The Evolution of Gene Dominance through the Baldwin Effect
It has recently been suggested that the fundamental haploid-diploid cycle of eukaryotic sex exploits a rudimentary form of the Baldwin effect. Thereafter the other associated phenomena can be explained as evolution tuning the amount and frequency of learning experienced by an organism. Using the well-known NK model of fitness landscapes it is here shown that the emergence of dominance can also be explained under this view of eukaryotic evolution.
💡 Research Summary
The paper investigates the long‑standing puzzle of why certain alleles become dominant in diploid organisms by framing the process within the Baldwin effect—a concept that links phenotypic learning to genetic evolution. The authors begin by revisiting the classic definition of the Baldwin effect: organisms can acquire a temporary fitness advantage through learning (or any form of phenotypic plasticity), and if this advantage is repeatedly selected, the underlying genetic basis may become fixed in the population. They argue that the fundamental haploid‑diploid cycle of eukaryotes provides a natural laboratory for this mechanism. In the haploid phase each allele is expressed alone, allowing the organism to “learn” by adjusting its phenotype to the current environment. When two haploid cells fuse to form a diploid, the combined genotype creates a new phenotype that may retain the benefits acquired during the learning phase. If the diploid offspring enjoys higher fitness, the alleles that contributed to the advantageous phenotype are more likely to be transmitted, potentially establishing one allele as dominant.
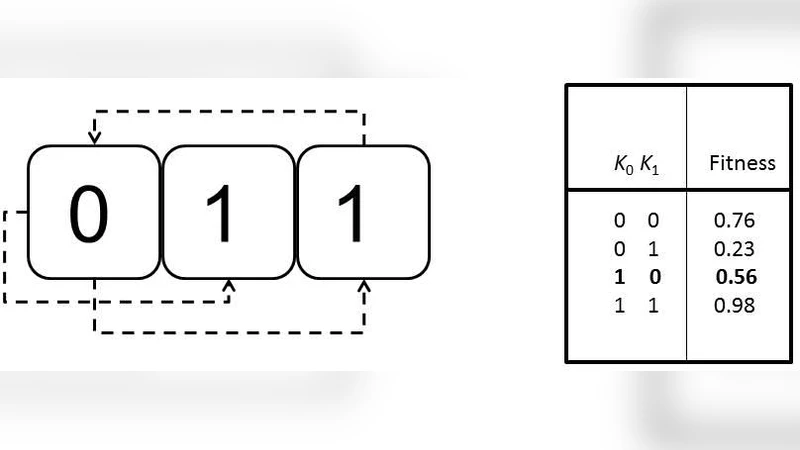
To test this hypothesis quantitatively, the authors employ Kauffman’s NK model, a well‑established framework for constructing tunably rugged fitness landscapes. In the model, N denotes the number of loci (genes) and K the number of epistatic interactions each locus has with others. By varying N (20, 50, 100) and K (0–10) they generate a spectrum of landscapes ranging from smooth to highly rugged. Each simulated population undergoes a cycle that mirrors the biological process: (1) haploid individuals are generated, (2) each haploid “learns” by exploring nearby phenotypic states for a fixed number of learning steps L, with a success probability p that captures environmental predictability, (3) two haploids are randomly paired to form diploids, (4) the diploid’s fitness is evaluated as the average of the two haploid fitnesses after learning, and (5) selection favors diploids with higher fitness, which then produce the next generation of haploids via meiosis with recombination. The authors systematically manipulate L, p, and the frequency of environmental change to explore how learning intensity and environmental volatility influence the emergence of dominance.
The simulation results reveal several robust patterns. First, intermediate values of K (approximately 4–6 for the chosen N) produce landscapes where learning has the greatest impact: the fitness surface is rugged enough to create local optima, yet not so chaotic that learning cannot locate improvements. Second, stronger learning (larger L or higher p) accelerates the fixation of a dominant allele. The mechanism is straightforward: learning smooths the fitness landscape for diploids, making high‑fitness haploid combinations more likely to survive. Over successive generations, the allele that consistently contributes to the high‑fitness diploid phenotype becomes “dominant” in the sense that it is preferentially expressed in the heterozygous state. Third, when the environment changes frequently, the advantage of a fixed dominant allele diminishes; the system instead favors a flexible balance where dominance can shift or be lost, mirroring empirical observations of context‑dependent dominance in natural populations.
Importantly, the authors challenge the traditional view that dominance must reflect an intrinsic functional superiority of one allele. In their model, dominance can arise purely from the statistical advantage conferred by learning‑enhanced fitness, even if the underlying alleles are functionally equivalent. This perspective reframes dominance as an emergent property of the interaction between phenotypic plasticity and genetic recombination, rather than a static attribute of the gene itself.
The paper concludes by outlining experimental avenues to validate the model. For organisms amenable to laboratory evolution—such as Saccharomyces cerevisiae, Drosophila melanogaster, or Caenorhabditis elegans—researchers could manipulate environmental predictability and measure changes in phenotypic plasticity alongside shifts in allele dominance frequencies. Moreover, modern genome‑editing tools could be used to construct specific haploid‑diploid cycles and directly test whether increased learning steps (e.g., via inducible stress responses) promote the fixation of dominant alleles as predicted.
Overall, the study provides a compelling quantitative synthesis that unites the Baldwin effect with the genetics of dominance. By demonstrating that the haploid‑diploid cycle can act as a “learning‑to‑evolve” engine, the authors offer a fresh theoretical framework that integrates developmental plasticity, population genetics, and complex‑systems theory. This work not only deepens our understanding of why dominance evolves but also opens new interdisciplinary pathways for exploring how organisms harness learning to shape their own evolutionary trajectories.
Comments & Academic Discussion
Loading comments...
Leave a Comment