Genomics as a Service: a Joint Computing and Networking Perspective
This paper provides a global picture about the deployment of networked processing services for genomic data sets. Many current research make an extensive use genomic data, which are massive and rapidly increasing over time. They are typically stored in remote databases, accessible by using Internet. For this reason, a significant issue for effectively handling genomic data through data networks consists of the available network services. A first contribution of this paper consists of identifying the still unexploited features of genomic data that could allow optimizing their networked management. The second and main contribution of this survey consists of a methodological classification of computing and networking alternatives which can be used to offer what we call the Genomic-as-a-Service (GaaS) paradigm. In more detail, we analyze the main genomic processing applications, and classify not only the main computing alternatives to run genomics workflows in either a local machine or a distributed cloud environment, but also the main software technologies available to develop genomic processing services. Since an analysis encompassing only the computing aspects would provide only a partial view of the issues for deploying GaaS system, we present also the main networking technologies that are available to efficiently support a GaaS solution. We first focus on existing service platforms, and analyze them in terms of service features, such as scalability, flexibility, and efficiency. Then, we present a taxonomy for both wide area and datacenter network technologies that may fit the GaaS requirements. It emerges that virtualization, both in computing and networking, is the key for a successful large-scale exploitation of genomic data, by pushing ahead the adoption of the GaaS paradigm. Finally, the paper illustrates a short and long-term vision on future research challenges in the field.
💡 Research Summary
The paper presents a comprehensive view of “Genomics‑as‑a‑Service” (GaaS), a paradigm that seeks to deliver large‑scale genomic data processing as a cloud‑based service by jointly considering computing and networking aspects. It begins by outlining the dramatic drop in DNA sequencing costs—now on the order of €1,000 per human genome—and the consequent exponential growth in raw genomic data (≈3.2 GB per genome) and associated metadata (tens to hundreds of gigabytes per analysis). The authors argue that the bottleneck is no longer sequencing capacity but the ability to move, store, and compute on these massive datasets across distributed networks.
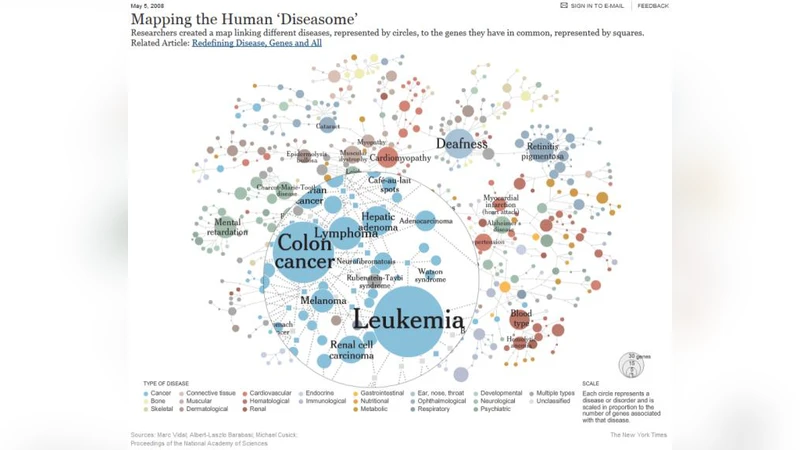
The paper’s first major contribution is the identification of under‑exploited data characteristics that could be leveraged for network‑aware optimization: temporal popularity shifts, inter‑genomic relationships (e.g., disease networks), and the heavy use of auxiliary reference files (GenBank, UniProt, GRCh38, etc.). These features suggest that conventional big‑data management techniques (simple replication, static caching) are insufficient for genomics workloads.
The second contribution is a methodological taxonomy of computing alternatives for GaaS. The authors classify infrastructures ranging from local workstations, high‑performance computing (HPC) clusters, grid systems, to public and private clouds. Within cloud environments they distinguish IaaS, PaaS, and SaaS layers, emphasizing containerization (Docker, Singularity) and workflow engines (Nextflow, Snakemake, Cromwell) to achieve reproducibility and elasticity. They also discuss hardware accelerators (GPU, FPGA, ASIC) and data‑centric programming models such as MapReduce and Spark, showing that these can dramatically speed up alignment, variant calling, and annotation stages. Two real‑world pipeline case studies are used to illustrate the specific compute and network requirements of typical whole‑genome and whole‑exome analyses.
The third contribution focuses on networking technologies. The authors present two complementary taxonomies: (i) wide‑area network (WAN) solutions, including traditional TCP/IP, software‑defined networking (SDN) for traffic engineering, dedicated fiber, satellite links, and emerging 5G mobile backhaul; and (ii) data‑center networking (DCN) solutions, covering classic switch‑based fabrics, overlay virtual networks (VXLAN, NVGRE), and RDMA‑based protocols (RoCE, iWARP). They argue that 5G’s ultra‑low latency and high bandwidth enable “edge‑GaaS” scenarios where raw sequencing data are cached close to the point of care, allowing real‑time clinical diagnostics. Within data centers, network function virtualization (NFV) combined with micro‑service architectures can dynamically allocate bandwidth and compute slices to meet the heterogeneous demands of different pipeline stages.
A central insight is that the convergence of computing virtualization (VMs, containers, Function‑as‑a‑Service) and network virtualization (SDN/NFV) is essential for scalable GaaS. By mapping service slices to both compute and network resources, providers can achieve on‑demand elasticity, cost efficiency, and fine‑grained QoS guarantees. The paper also addresses security and privacy concerns, recommending end‑to‑end encryption, homomorphic encryption, and secure multi‑party computation to satisfy regulatory requirements for medical data.
Finally, the authors outline short‑ and long‑term research challenges: (1) popularity‑aware pre‑fetching and caching strategies tailored to genomic datasets; (2) AI‑driven traffic prediction and auto‑scaling mechanisms; (3) interoperable multi‑cloud and multi‑edge orchestration frameworks; and (4) integration of GaaS into forthcoming 5G/6G service architectures. They conclude that virtualization, both in computing and networking, will be the key enabler for the widespread adoption of GaaS, turning genomics from a niche research activity into a ubiquitous, on‑demand service across medicine, agriculture, forensics, and beyond.
Comments & Academic Discussion
Loading comments...
Leave a Comment