Leabra7: a Python package for modeling recurrent, biologically-realistic neural networks
Emergent is a software package that uses the AdEx neural dynamics model and LEABRA learning algorithm to simulate and train arbitrary recurrent neural network architectures in a biologically-realistic manner. We present Leabra7, a complementary Python library that implements these same algorithms. Leabra7 is developed and distributed using modern software development principles, and integrates tightly with Python’s scientific stack. We demonstrate recurrent Leabra7 networks using traditional pattern-association tasks and a standard machine learning task, classifying the IRIS dataset.
💡 Research Summary
Leabra7 is a Python library that re‑implements the core components of the Emergent cognitive modeling system—namely the Adaptive‑Exponential (AdEx) neuronal dynamics and the LEABRA learning algorithm—within a modern software engineering framework. The authors begin by motivating the need for biologically plausible recurrent neural network (RNN) models, noting that virtually all cortical areas contain feedback connections and that current deep learning approaches rely heavily on specialized architectures such as LSTMs to circumvent vanishing gradients. By combining a biophysically realistic spiking model (AdEx) with a local, error‑driven learning rule (LEABRA), Leabra7 aims to provide a platform capable of training arbitrary recurrent topologies without back‑propagation.
The library is built around three primary objects: Net, Layer, and Projection. A Net instance orchestrates the entire simulation, while layers contain units (neurons) and projections bundle the connections between layers. Parameter configuration is handled through “Spec” objects (e.g., LayerSpec, ProjnSpec), which expose the same descriptive names used in Emergent, allowing users to override defaults such as inhibition type (feed‑forward, feedback, or k‑winner‑take‑all) or logging behavior. The API supports three cycle‑level operations—simple stepping (net.cycle()), minus‑phase cycling, and plus‑phase cycling—mirroring the phases required by LEABRA. After each phase, net.learn() computes weight changes based on the difference between minus and plus activations.
Observation and logging are tightly integrated with the scientific Python stack. Net.observe() returns instantaneous snapshots of any attribute as a pandas DataFrame, facilitating immediate analysis or visualization. Persistent logging is configured at network creation via Spec parameters; logged data are also stored as DataFrames, with a “time” column indicating the simulation cycle. Because logging incurs memory and CPU overhead, the library provides pause_logging() and resume_logging() methods to disable logging during intensive training phases.
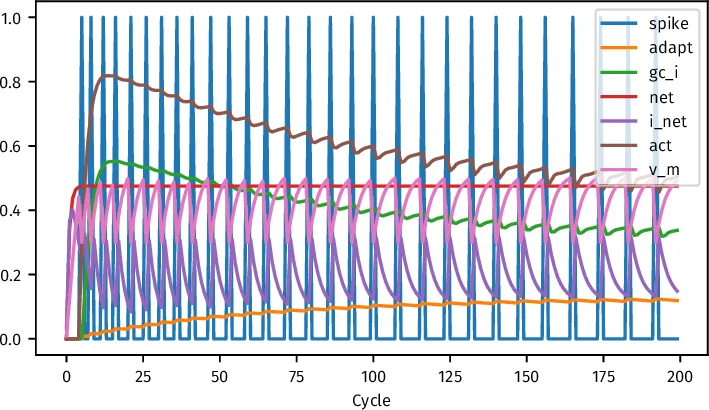
Four example networks illustrate the library’s capabilities. The first, a minimal two‑neuron system, demonstrates basic spiking dynamics: input clamping drives the output neuron’s membrane potential, spiking, adaptation current, and inhibitory feedback. The second example implements a simple feed‑forward pattern‑association task (2‑input → 4‑output) and shows a monotonic decline in a thresholded mean‑squared error over 500 epochs. The third example introduces a hidden layer and a recurrent feedback projection (output → hidden) to solve a non‑linearly separable problem; the loss curve exhibits oscillations due to the sensitivity of recurrent connections, yet learning converges. The fourth example applies Leabra7 to the classic IRIS classification benchmark. After quantile‑based binning of the four continuous features (10 bins per feature, yielding 36 input units) and a hidden layer of 23 units, the network attains 95.83 % training accuracy and 90.00 % test accuracy after 500 epochs, with loss curves plotted for both training and test sets.
The authors conclude that Leabra7 successfully brings biologically realistic recurrent modeling into the Python ecosystem, offering ease of modification, reproducibility via conda distribution, and seamless integration with Jupyter notebooks and containerized workflows. However, they acknowledge two primary limitations for scaling to large cognitive models: (1) learning performance—specifically the lack of adaptive learning‑rate schedules and annealing, which currently cause late‑stage loss oscillations; and (2) computational speed, as the pure‑Python implementation lacks the efficiency of compiled engines. Future work will explore adding adaptive learning‑rate mechanisms and possibly a C++ backend to accelerate large‑scale simulations, while preserving the library’s openness and modifiability. With these enhancements, Leabra7 could be employed for sophisticated models such as hippocampal simulations and other high‑dimensional cognitive architectures.
Comments & Academic Discussion
Loading comments...
Leave a Comment