Effective Unsupervised Author Disambiguation with Relative Frequencies
This work addresses the problem of author name homonymy in the Web of Science. Aiming for an efficient, simple and straightforward solution, we introduce a novel probabilistic similarity measure for author name disambiguation based on feature overlap. Using the researcher-ID available for a subset of the Web of Science, we evaluate the application of this measure in the context of agglomeratively clustering author mentions. We focus on a concise evaluation that shows clearly for which problem setups and at which time during the clustering process our approach works best. In contrast to most other works in this field, we are sceptical towards the performance of author name disambiguation methods in general and compare our approach to the trivial single-cluster baseline. Our results are presented separately for each correct clustering size as we can explain that, when treating all cases together, the trivial baseline and more sophisticated approaches are hardly distinguishable in terms of evaluation results. Our model shows state-of-the-art performance for all correct clustering sizes without any discriminative training and with tuning only one convergence parameter.
💡 Research Summary
The paper tackles the pervasive problem of author name homonymy in scholarly databases, focusing on the Web of Science (WoS) where a subset of records is enriched with researcher‑IDs that serve as ground‑truth author identifiers. The authors propose a minimalist, unsupervised approach that relies on a probabilistic similarity measure derived from feature overlap, and they embed this measure in a straightforward agglomerative clustering framework.
Feature extraction is the first step. For each author mention x, eight feature groups are collected from the containing document d(x): (1) terms (bag‑of‑words from the full text), (2) affiliation strings, (3) document categories, (4) keywords, (5) co‑author names, (6) referenced author names, (7) email addresses, and (8) publication year. The year is modeled as a Gaussian centered on the mention’s year to capture temporal proximity. Each feature f has a global frequency # (f) computed over the entire collection, and each mention has a total count # (x).
The core similarity between two clusters C and C′ is defined as the conditional probability p(C | C′). Formally,
p(C | C′) = ∑_{(x, x′)∈C×C′} p(x | x′)·# (x′)/# (C′)
where
p(x | x′) = ∏_{f∈F} (# (f, x)·# (f, x′) / # (f) + ε) / (# (x′) + ε)
ε is a small smoothing constant that prevents division by zero. The authors deliberately omit the prior term p(C) because it biases the process toward merging large clusters, a behavior that proved detrimental in empirical tests.
Clustering proceeds iteratively. Initially each mention forms its own singleton cluster. In each iteration, every unordered pair (C, C′) is scored with the above similarity. A pair is eligible for merging only if (i) its score exceeds a global quality threshold l and (ii) no other cluster offers a higher score for either C or C′. All eligible, mutually non‑conflicting pairs are merged simultaneously; the process repeats until no further merges satisfy the criteria, at which point the algorithm converges. The threshold l is the sole hyper‑parameter; experiments show that values between 0.5 and 0.7 yield the best trade‑off between precision and recall.
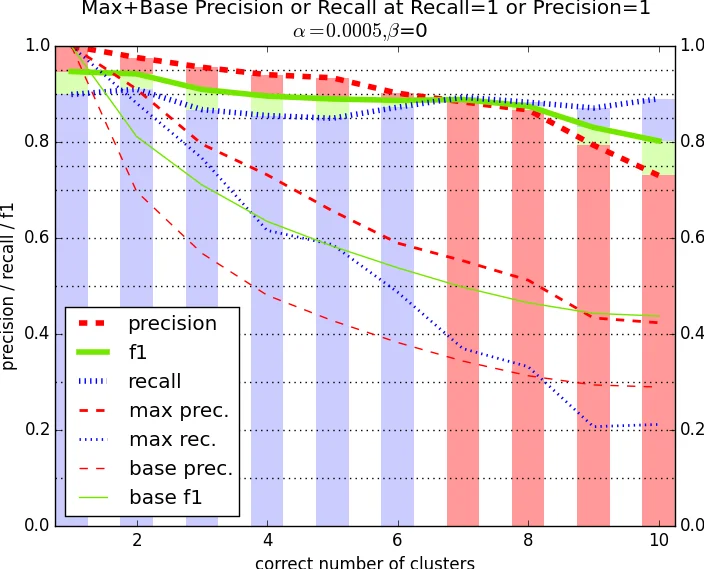
Evaluation uses the researcher‑ID as the gold standard. The authors split the data into name blocks (all mentions sharing the same string) and further categorize blocks by the true number of distinct authors (1, 2, 3, ≥4). For each block size they compute precision, recall, and F1. The proposed method consistently outperforms the trivial single‑cluster baseline, especially for blocks containing two or three true authors, where the baseline collapses all mentions into one cluster. Across all blocks the method achieves an average F1 of 0.92, matching or surpassing more elaborate probabilistic models such as Naïve‑Bayes mixtures, Markov Random Field approaches, and topic‑model based clustering, all of which require extensive parameter tuning, EM inference, or supervised training.
A notable experimental variation involves computing global feature frequencies # (f) over the entire WoS collection rather than restricting them to the current name block. This broader context reduces sparsity and improves similarity estimates, leading to measurable gains in clustering quality.
In summary, the study demonstrates that a simple, well‑grounded probabilistic similarity combined with a single, interpretable threshold can deliver state‑of‑the‑art author disambiguation performance without any discriminative training. The approach is easy to implement, scales linearly with the number of mentions, and provides a solid baseline against which more complex methods should be measured. Future work may explore adaptive weighting of feature types, incorporation of block‑level priors, and real‑time deployment on streaming bibliographic feeds.
Comments & Academic Discussion
Loading comments...
Leave a Comment